Abstract
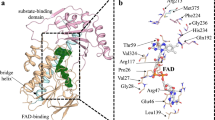
Although tetracyclines are an important class of antibiotics for use in agriculture and the clinic, their efficacy is threatened by increasing resistance. Resistance to tetracyclines can occur through efflux, ribosomal protection, or enzymatic inactivation. Surprisingly, tetracycline enzymatic inactivation has remained largely unexplored, despite providing the distinct advantage of antibiotic clearance. The tetracycline destructases are a recently discovered family of tetracycline-inactivating flavoenzymes from pathogens and soil metagenomes that have a high potential for broad dissemination. Here, we show that tetracycline destructases accommodate tetracycline-class antibiotics in diverse and novel orientations for catalysis, and antibiotic binding drives unprecedented structural dynamics facilitating tetracycline inactivation. We identify a key inhibitor binding mode that locks the flavin adenine dinucleotide cofactor in an inactive state, functionally rescuing tetracycline activity. Our results reveal the potential of a new tetracycline and tetracycline destructase inhibitor combination therapy strategy to overcome resistance by enzymatic inactivation and restore the use of an important class of antibiotics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Knapp, C.W., Dolfing, J., Ehlert, P.A. & Graham, D.W. Evidence of increasing antibiotic resistance gene abundances in archived soils since 1940. Environ. Sci. Technol. 44, 580–587 (2010).
Kinch, M.S., Patridge, E., Plummer, M. & Hoyer, D. An analysis of FDA-approved drugs for infectious disease: antibacterial agents. Drug Discov. Today 19, 1283–1287 (2014).
Davies, J. Inactivation of antibiotics and the dissemination of resistance genes. Science 264, 375–382 (1994).
Allen, H.K. et al. Call of the wild: antibiotic resistance genes in natural environments. Nat. Rev. Microbiol. 8, 251–259 (2010).
Berendonk, T.U. et al. Tackling antibiotic resistance: the environmental framework. Nat. Rev. Microbiol. 13, 310–317 (2015).
Benveniste, R. & Davies, J. Aminoglycoside antibiotic-inactivating enzymes in actinomycetes similar to those present in clinical isolates of antibiotic-resistant bacteria. Proc. Natl. Acad. Sci. USA 70, 2276–2280 (1973).
D'Costa, V.M. et al. Antibiotic resistance is ancient. Nature 477, 457–461 (2011).
Forsberg, K.J. et al. The shared antibiotic resistome of soil bacteria and human pathogens. Science 337, 1107–1111 (2012).
Forsberg, K.J. et al. Bacterial phylogeny structures soil resistomes across habitats. Nature 509, 612–616 (2014).
Yong, D. et al. Characterization of a new metallo-β-lactamase gene, bla(NDM-1), and a novel erythromycin esterase gene carried on a unique genetic structure in Klebsiella pneumoniae sequence type 14 from India. Antimicrob. Agents Chemother. 53, 5046–5054 (2009).
Liu, Y.Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect. Dis. 16, 161–168 (2016).
Poirel, L., Rodriguez-Martinez, J.M., Mammeri, H., Liard, A. & Nordmann, P. Origin of plasmid-mediated quinolone resistance determinant QnrA. Antimicrob. Agents Chemother. 49, 3523–3525 (2005).
Thaker, M., Spanogiannopoulos, P. & Wright, G.D. The tetracycline resistome. Cell. Mol. Life Sci. 67, 419–431 (2010).
State of the World's Antibiotics. 2015. (Center for Disease Dynamics, Economics & Policy, Washington, DC, USA, 2015).
Kasbekar, N. Tigecycline: a new glycylcycline antimicrobial agent. Am. J. Health Syst. Pharm. 63, 1235–1243 (2006).
Sutcliffe, J.A., O'Brien, W., Fyfe, C. & Grossman, T.H. Antibacterial activity of eravacycline (TP-434), a novel fluorocycline, against hospital and community pathogens. Antimicrob. Agents Chemother. 57, 5548–5558 (2013).
Macone, A.B. et al. In vitro and in vivo antibacterial activities of omadacycline, a novel aminomethylcycline. Antimicrob. Agents Chemother. 58, 1127–1135 (2014).
Chopra, I. & Roberts, M. Tetracycline antibiotics: mode of action, applications, molecular biology, and epidemiology of bacterial resistance. Microbiol. Mol. Biol. Rev. 65, 232–260 (2001).
Park, B.H. & Levy, S.B. The cryptic tetracycline resistance determinant on Tn4400 mediates tetracycline degradation as well as tetracycline efflux. Antimicrob. Agents Chemother. 32, 1797–1800 (1988).
Speer, B.S. & Salyers, A.A. Characterization of a novel tetracycline resistance that functions only in aerobically grown Escherichia coli. J. Bacteriol. 170, 1423–1429 (1988).
Whittle, G., Hund, B.D., Shoemaker, N.B. & Salyers, A.A. Characterization of the 13-kilobase ermF region of the Bacteroides conjugative transposon CTnDOT. Appl. Environ. Microbiol. 67, 3488–3495 (2001).
Nonaka, L. & Suzuki, S. New Mg2+-dependent oxytetracycline resistance determinant tet 34 in Vibrio isolates from marine fish intestinal contents. Antimicrob. Agents Chemother. 46, 1550–1552 (2002).
Diaz-Torres, M.L. et al. Novel tetracycline resistance determinant from the oral metagenome. Antimicrob. Agents Chemother. 47, 1430–1432 (2003).
Ghosh, S., Sadowsky, M.J., Roberts, M.C., Gralnick, J.A. & LaPara, T.M. Sphingobacterium sp. strain PM2-P1-29 harbours a functional tet(X) gene encoding for the degradation of tetracycline. J. Appl. Microbiol. 106, 1336–1342 (2009).
Yang, W. et al. TetX is a flavin-dependent monooxygenase conferring resistance to tetracycline antibiotics. J. Biol. Chem. 279, 52346–52352 (2004).
Moore, I.F., Hughes, D.W. & Wright, G.D. Tigecycline is modified by the flavin-dependent monooxygenase TetX. Biochemistry 44, 11829–11835 (2005).
Volkers, G., Palm, G.J., Weiss, M.S., Wright, G.D. & Hinrichs, W. Structural basis for a new tetracycline resistance mechanism relying on the TetX monooxygenase. FEBS Lett. 585, 1061–1066 (2011).
Forsberg, K.J., Patel, S., Wencewicz, T.A. & Dantas, G. The tetracycline destructases: a novel family of tetracycline-inactivating enzymes. Chem. Biol. 22, 888–897 (2015).
Drawz, S.M., Papp-Wallace, K.M. & Bonomo, R.A. New β-lactamase inhibitors: a therapeutic renaissance in an MDR world. Antimicrob. Agents Chemother. 58, 1835–1846 (2014).
Hornak, V., Okur, A., Rizzo, R.C. & Simmerling, C. HIV-1 protease flaps spontaneously close to the correct structure in simulations following manual placement of an inhibitor into the open state. J. Am. Chem. Soc. 128, 2812–2813 (2006).
Pesnot, T., Jørgensen, R., Palcic, M.M. & Wagner, G.K. Structural and mechanistic basis for a new mode of glycosyltransferase inhibition. Nat. Chem. Biol. 6, 321–323 (2010).
Cazalet, C. et al. Analysis of the Legionella longbeachae genome and transcriptome uncovers unique strategies to cause Legionnaires' disease. PLoS Genet. 6, e1000851 (2010).
Whiley, H. & Bentham, R. Legionella longbeachae and legionellosis. Emerg. Infect. Dis. 17, 579–583 (2011).
Ballou, D.P., Entsch, B. & Cole, L.J. Dynamics involved in catalysis by single-component and two-component flavin-dependent aromatic hydroxylases. Biochem. Biophys. Res. Commun. 338, 590–598 (2005).
van Berkel, W.J., Kamerbeek, N.M. & Fraaije, M.W. Flavoprotein monooxygenases, a diverse class of oxidative biocatalysts. J. Biotechnol. 124, 670–689 (2006).
Gatti, D.L. et al. The mobile flavin of 4-OH benzoate hydroxylase. Science 266, 110–114 (1994).
Massey, V. Activation of molecular oxygen by flavins and flavoproteins. J. Biol. Chem. 269, 22459–22462 (1994).
Volkers, G. et al. Putative dioxygen-binding sites and recognition of tigecycline and minocycline in the tetracycline-degrading monooxygenase TetX. Acta Crystallogr. D Biol. Crystallogr. 69, 1758–1767 (2013).
Liu, L.K. et al. The structure of the antibiotic deactivating, N-hydroxylating rifampicin monooxygenase. J. Biol. Chem. 291, 21553–21562 (2016).
Gibson, M., Nur-e-alam, M., Lipata, F., Oliveira, M.A. & Rohr, J. Characterization of kinetics and products of the Baeyer–Villiger oxygenase MtmOIV, the key enzyme of the biosynthetic pathway toward the natural product anticancer drug mithramycin from Streptomyces argillaceus. J. Am. Chem. Soc. 127, 17594–17595 (2005).
Wang, P., Bashiri, G., Gao, X., Sawaya, M.R. & Tang, Y. Uncovering the enzymes that catalyze the final steps in oxytetracycline biosynthesis. J. Am. Chem. Soc. 135, 7138–7141 (2013).
Yuen, P.H. & Sokoloski, T.D. Kinetics of concomitant degradation of tetracycline to epitetracycline, anhydrotetracycline, and epianhydrotetracycline in acid phosphate solution. J. Pharm. Sci. 66, 1648–1650 (1977).
Palmer, A.C., Angelino, E. & Kishony, R. Chemical decay of an antibiotic inverts selection for resistance. Nat. Chem. Biol. 6, 105–107 (2010).
Liu, F. & Myers, A.G. Development of a platform for the discovery and practical synthesis of new tetracycline antibiotics. Curr. Opin. Chem. Biol. 32, 48–57 (2016).
Lienhart, W.D., Gudipati, V. & Macheroux, P. The human flavoproteome. Arch. Biochem. Biophys. 535, 150–162 (2013).
Leski, T.A. et al. Multidrug-resistant tet(X)-containing hospital isolates in Sierra Leone. Int. J. Antimicrob. Agents 42, 83–86 (2013).
Deng, M. et al. Molecular epidemiology and mechanisms of tigecycline resistance in clinical isolates of Acinetobacter baumannii from a Chinese university hospital. Antimicrob. Agents Chemother. 58, 297–303 (2014).
Boucher, H.W. et al. Bad bugs, no drugs: no ESKAPE! An update from the Infectious Diseases Society of America. Clin. Infect. Dis. 48, 1–12 (2009).
Walsh, C.T. & Wencewicz, T.A. Flavoenzymes: versatile catalysts in biosynthetic pathways. Nat. Prod. Rep. 30, 175–200 (2013).
Merriam, J.J., Mathur, R., Maxfield-Boumil, R. & Isberg, R.R. Analysis of the Legionella pneumophila fliI gene: intracellular growth of a defined mutant defective for flagellum biosynthesis. Infect. Immun. 65, 2497–2501 (1997).
Feeley, J.C. et al. Charcoal-yeast extract agar: primary isolation medium for Legionella pneumophila. J. Clin. Microbiol. 10, 437–441 (1979).
Berger, K.H. & Isberg, R.R. Two distinct defects in intracellular growth complemented by a single genetic locus in Legionella pneumophila. Mol. Microbiol. 7, 7–19 (1993).
Sexton, J.A. et al. The Legionella pneumophila PilT homologue DotB exhibits ATPase activity that is critical for intracellular growth. J. Bacteriol. 186, 1658–1666 (2004).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
CLSI. Methods for Dilution Antimicrobial Susceptibility Testing for Bacteria That Grow Aerobically; Approved Standard–Tenth Edition. Vol. M07–A10 (Clinical and Laboratory Standards Institute, 2015).
Berenbaum, M.C. A method for testing for synergy with any number of agents. J. Infect. Dis. 137, 122–130 (1978).
Acknowledgements
We thank J. Nix and ALS beamline 4.2.2 (contract DE-AC02-05CH11231) for assistance with X-ray data collection and A. Durairaj of the Proteomics & Mass Spectrometry Facility at the Danforth Plant Science Center for assistance with HR–MS/MS experiments. This work was supported by an award to N.H.T., G.D., and T.A.W. from the National Institute of Allergy and Infectious Diseases of the National Institutes of Health (R01 AI123394). A.J.G. is supported by the National Institute of General Medical Sciences Cell and Molecular Biology Training Grant (T32 GM007067). The content is solely the responsibility of the authors and does not necessarily represent the official views of the funding agencies.
Author information
Authors and Affiliations
Contributions
J.P. designed and performed crystallographic experiments and X-ray structure determination, analyzed data, and wrote the paper; A.J.G. designed and performed in vitro and microbiological experiments, analyzed data, and wrote the paper; M.R.R. and C.T.S. performed in vitro experiments; J.L.E. performed crystallographic experiments; J.P.V. performed Legionella experiments; T.A.W., G.D., and N.H.T. designed experiments, analyzed data, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–5 and Supplementary Figures 1–10 (PDF 2707 kb)
Rights and permissions
About this article
Cite this article
Park, J., Gasparrini, A., Reck, M. et al. Plasticity, dynamics, and inhibition of emerging tetracycline resistance enzymes. Nat Chem Biol 13, 730–736 (2017). https://doi.org/10.1038/nchembio.2376
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2376
This article is cited by
-
Sequence-structure-function characterization of the emerging tetracycline destructase family of antibiotic resistance enzymes
Communications Biology (2024)
-
Oxidation of chlortetracycline and its isomers by Botrytis aclada laccase in the absence of mediators: pH dependence and identification of transformation products by LC–MS
Biodegradation (2024)
-
Structure of anhydrotetracycline-bound Tet(X6) reveals the mechanism for inhibition of type 1 tetracycline destructases
Communications Biology (2023)
-
Synthesis of Mo-Based/Carbon Nanocomposistes for Water Decontamination via Percarbonate Activation
Catalysis Letters (2023)
-
Biology and applications of co-produced, synergistic antimicrobials from environmental bacteria
Nature Microbiology (2021)