Abstract
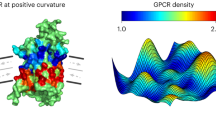
The targeted spatial organization (sorting) of Gprotein-coupled receptors (GPCRs) is essential for their biological function and often takes place in highly curved membrane compartments such as filopodia, endocytic pits, trafficking vesicles or endosome tubules. However, the influence of geometrical membrane curvature on GPCR sorting remains unknown. Here we used fluorescence imaging to establish a quantitative correlation between membrane curvature and sorting of three prototypic class A GPCRs (the neuropeptide Y receptor Y2, the β1 adrenergic receptor and the β2 adrenergic receptor) in living cells. Fitting of a thermodynamic model to the data enabled us to quantify how sorting is mediated by an energetic drive to match receptor shape and membrane curvature. Curvature-dependent sorting was regulated by ligands in a specific manner. We anticipate that this curvature-dependent biomechanical coupling mechanism contributes to the sorting, trafficking and function of transmembrane proteins in general.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Allen, J.A., Halverson-Tamboli, R.A. & Rasenick, M.M. Lipid raft microdomains and neurotransmitter signalling. Nat. Rev. Neurosci. 8, 128–140 (2007).
Irannejad, R. et al. Conformational biosensors reveal GPCR signalling from endosomes. Nature 495, 534–538 (2013).
Ritter, S.L. & Hall, R.A. Fine-tuning of GPCR activity by receptor-interacting proteins. Nat. Rev. Mol. Cell Biol. 10, 819–830 (2009).
Soubias, O., Teague, W.E. Jr., Hines, K.G. & Gawrisch, K. Rhodopsin/lipid hydrophobic matching-rhodopsin oligomerization and function. Biophys. J. 108, 1125–1132 (2015).
Kimura, T. et al. Recombinant cannabinoid type 2 receptor in liposome model activates G protein in response to anionic lipid constituents. J. Biol. Chem. 287, 4076–4087 (2012).
Mondal, S. et al. Membrane driven spatial organization of GPCRs. Sci. Rep. 3, 2909 (2013).
Koldsø, H. & Sansom, M.S. Organization and Dynamics of Receptor Proteins in a Plasma Membrane. J. Am. Chem. Soc. 137, 14694–14704 (2015).
Mattila, P.K. & Lappalainen, P. Filopodia: molecular architecture and cellular functions. Nat. Rev. Mol. Cell Biol. 9, 446–454 (2008).
Zimmerberg, J. & Kozlov, M.M. How proteins produce cellular membrane curvature. Nat. Rev. Mol. Cell Biol. 7, 9–19 (2006).
Temkin, P. et al. SNX27 mediates retromer tubule entry and endosome-to-plasma membrane trafficking of signalling receptors. Nat. Cell Biol. 13, 715–721 (2011).
Iversen, L., Mathiasen, S., Larsen, J.B. & Stamou, D. Membrane curvature bends the laws of physics and chemistry. Nat. Chem. Biol. 11, 822–825 (2015).
Shim, S.H. et al. Super-resolution fluorescence imaging of organelles in live cells with photoswitchable membrane probes. Proc. Natl. Acad. Sci. USA 109, 13978–13983 (2012).
Veshaguri, S. et al. Direct observation of proton pumping by a eukaryotic P-type ATPase. Science 351, 1469–1473 (2016).
Sorre, B. et al. Nature of curvature coupling of amphiphysin with membranes depends on its bound density. Proc. Natl. Acad. Sci. USA 109, 173–178 (2012).
Larsen, J.B. et al. Membrane curvature enables N-Ras lipid anchor sorting to liquid-ordered membrane phases. Nat. Chem. Biol. 11, 192–194 (2015).
Kunding, A.H., Mortensen, M.W., Christensen, S.M. & Stamou, D. A fluorescence-based technique to construct size distributions from single-object measurements: application to the extrusion of lipid vesicles. Biophys. J. 95, 1176–1188 (2008).
Kuo, L.E. et al. Neuropeptide Y acts directly in the periphery on fat tissue and mediates stress-induced obesity and metabolic syndrome. Nat. Med. 13, 803–811 (2007).
Movafagh, S., Hobson, J.P., Spiegel, S., Kleinman, H.K. & Zukowska, Z. Neuropeptide Y induces migration, proliferation, and tube formation of endothelial cells bimodally via Y1, Y2, and Y5 receptors. FASEB J. 20, 1924–1926 (2006).
Gerhardt, H. et al. VEGF guides angiogenic sprouting utilizing endothelial tip cell filopodia. J. Cell Biol. 161, 1163–1177 (2003).
Stanić, D. et al. Characterization of neuropeptide Y2 receptor protein expression in the mouse brain. I. Distribution in cell bodies and nerve terminals. J. Comp. Neurol. 499, 357–390 (2006).
Rustom, A., Saffrich, R., Markovic, I., Walther, P. & Gerdes, H.H. Nanotubular highways for intercellular organelle transport. Science 303, 1007–1010 (2004).
Tian, A. & Baumgart, T. Sorting of lipids and proteins in membrane curvature gradients. Biophys. J. 96, 2676–2688 (2009).
Bornschlögl, T. et al. Filopodial retraction force is generated by cortical actin dynamics and controlled by reversible tethering at the tip. Proc. Natl. Acad. Sci. USA 110, 18928–18933 (2013).
Romero, S. et al. Filopodium retraction is controlled by adhesion to its tip. J. Cell Sci. 125, 4999–5004 (2012).
Revenu, C., Athman, R., Robine, S. & Louvard, D. The co-workers of actin filaments: from cell structures to signals. Nat. Rev. Mol. Cell Biol. 5, 635–646 (2004).
Keppler, A. et al. A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat. Biotechnol. 21, 86–89 (2003).
Aimon, S. et al. Membrane shape modulates transmembrane protein distribution. Dev. Cell 28, 212–218 (2014).
Brownell, W.E., Qian, F. & Anvari, B. Cell membrane tethers generate mechanical force in response to electrical stimulation. Biophys. J. 99, 845–852 (2010).
Leijnse, N., Oddershede, L.B. & Bendix, P.M. Helical buckling of actin inside filopodia generates traction. Proc. Natl. Acad. Sci. USA 112, 136–141 (2015).
Markin, V.S. Lateral organization of membranes and cell shapes. Biophys. J. 36, 1–19 (1981).
Ramaswamy, S., Toner, J. & Prost, J. Nonequilibrium fluctuations, traveling waves, and instabilities in active membranes. Phys. Rev. Lett. 84, 3494–3497 (2000).
Netz, R.R. & Pincus, P. Inhomogeneous fluid membranes: Segregation, ordering, and effective rigidity. Phys. Rev. E Stat. Phys. Plasmas Fluids Relat. Interdiscip. Topics 52, 4114–4128 (1995).
Božič, B., Kralj-Iglič, V. & Svetina, S. Coupling between vesicle shape and lateral distribution of mobile membrane inclusions. Phys. Rev. E 73, 041915 (2006).
Keire, D.A. et al. Primary structures of PYY, [Pro34]PYY, and PYY-(3-36) confer different conformations and receptor selectivity. Am. J. Physiol. Gastrointest. Liver Physiol. 279, G126–G131 (2000).
Baker, J.G. The selectivity of β-adrenoceptor agonists at human β1-, β2- and β3-adrenoceptors. Br. J. Pharmacol. 160, 1048–1061 (2010).
Lippincott-Schwartz, J. & Phair, R.D. Lipids and cholesterol as regulators of traffic in the endomembrane system. Annu. Rev. Biophys. 39, 559–578 (2010).
Ramamurthi, K.S., Lecuyer, S., Stone, H.A. & Losick, R. Geometric cue for protein localization in a bacterium. Science 323, 1354–1357 (2009).
Galic, M. et al. External push and internal pull forces recruit curvature-sensing N-BAR domain proteins to the plasma membrane. Nat. Cell Biol. 14, 874–881 (2012).
Hägerstrand, H. et al. Curvature-dependent lateral distribution of raft markers in the human erythrocyte membrane. Mol. Membr. Biol. 23, 277–288 (2006).
Palczewski, K. et al. Crystal structure of rhodopsin: a Gprotein-coupled receptor. Science 289, 739–745 (2000).
Rasmussen, S.G.F. et al. Structure of a nanobody-stabilized active state of the β2 adrenoceptor. Nature 469, 175–180 (2011).
Costanzi, S. On the applicability of GPCR homology models to computer-aided drug discovery: a comparison between in silico and crystal structures of the β2-adrenergic receptor. J. Med. Chem. 51, 2907–2914 (2008).
Huber, T., Botelho, A.V., Beyer, K. & Brown, M.F. Membrane model for the G-protein-coupled receptor rhodopsin: hydrophobic interface and dynamical structure. Biophys. J. 86, 2078–2100 (2004).
Khelashvili, G. et al. Why GPCRs behave differently in cubic and lamellar lipidic mesophases. J. Am. Chem. Soc. 134, 15858–15868 (2012).
Mathiasen, S. et al. Nanoscale high-content analysis using compositional heterogeneities of single proteoliposomes. Nat. Methods 11, 931–934 (2014).
Gonen, T., Sliz, P., Kistler, J., Cheng, Y. & Walz, T. Aquaporin-0 membrane junctions reveal the structure of a closed water pore. Nature 429, 193–197 (2004).
Warne, T. et al. Structure of a β1-adrenergic G-protein-coupled receptor. Nature 454, 486–491 (2008).
Nygaard, R. et al. The dynamic process of β2-adrenergic receptor activation. Cell 152, 532–542 (2013).
Salaita, K. et al. Restriction of receptor movement alters cellular response: physical force sensing by EphA2. Science 327, 1380–1385 (2010).
Sharpe, H.J., Stevens, T.J. & Munro, S. A comprehensive comparison of transmembrane domains reveals organelle-specific properties. Cell 142, 158–169 (2010).
Pedersen, S.L. et al. Improving membrane binding as a design strategy for amphipathic peptide hormones: 2-helix variants of PYY3-36. J. Pept. Sci. 18, 579–587 (2012).
Reihani, S.N.S., Mir, S.A., Richardson, A.C. & Oddershede, L.B. Significant improvement of optical traps by tuning standard water immersion objectives. J. Opt. 13, 105301 (2011).
Richardson, A.C., Reihani, N., & Oddershede, L.B. Combining confocal microscopy with precise force-scope optical tweezers. Proc. SPIE 6326, 632628 (2006).
Pontes, B. et al. Cell cytoskeleton and tether extraction. Biophys. J. 101, 43–52 (2011).
Berk, D.A. & Hochmuth, R.M. Lateral mobility of integral proteins in red blood cell tethers. Biophys. J. 61, 9–18 (1992).
Prévost, C. et al. IRSp53 senses negative membrane curvature and phase separates along membrane tubules. Nat. Commun. 6, 8529 (2015).
Kralj-Iglic, V., Heinrich, V., Svetina, S. & Zeks, B. Free energy of closed membrane with anisotropic inclusions. Eur. Phys. J. B 10, 5–8 (1999).
Callan-Jones, A., Durand, M. & Fournier, J.B. Hydrodynamics of bilayer membranes with diffusing transmembrane proteins. Soft Matter 12, 1791–1800 (2016).
Dill, K.A. & Bromberg, S. Molecular Driving Forces: Statistical Thermodynamics in Chemistry and Biology (Garland Science, New York, 2003).
Gatz, D.F. & Smith, L. The standard error of a weighted mean concentration. 1. Bootstrapping vs other methods. Atmos. Environ. 29, 1185–1193 (1995).
Acknowledgements
This work was supported by the Lundbeck Foundation (Center of Excellence Biomembranes in Nanomedicine), the Danish Council for Strategic Research (1311-00002B), and the Innovation Fund Denmark (5184-00048B) to D.S.'s research group, by the Danish National Research Foundation (DNRF116) to L.B.O.'s research group and by the Villum Kann Rasmussen Foundation (VKR022593 to P.M.B.).
Author information
Authors and Affiliations
Contributions
D.S. designed and supervised the project. K.R.R. conducted all experiments and data analysis, A.M. assisted with experiments and data analysis, V.T. helped with cell culturing, and N.S.H. helped design experiments and discuss results. A.C.-J. developed the theoretical sorting model and performed the numerical fits. L.B.O. provided the optical tweezers setup and supervised the optical trapping experiments. N.L. and K.R.R. conducted the pulling tether experiments. N.L. provided the expertise in using the laser tweezers setup, and P.M.B. discussed results and data treatment. K.L.M. and V.F.W. provided the plasmid and know-how for expressing the SNAP-tagged Y2R, β1AR and β2AR. K.J.J. and S.L.P. synthesized the PYY3-36 peptide. All authors discussed the results and commented on the manuscript, which was written by K.R.R. and D.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Table 1 and Supplementary Figures 1–5 (PDF 4662 kb)
Rights and permissions
About this article
Cite this article
Rosholm, K., Leijnse, N., Mantsiou, A. et al. Membrane curvature regulates ligand-specific membrane sorting of GPCRs in living cells. Nat Chem Biol 13, 724–729 (2017). https://doi.org/10.1038/nchembio.2372
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2372
This article is cited by
-
Molecular mechanism of GPCR spatial organization at the plasma membrane
Nature Chemical Biology (2024)
-
Membrane mediated mechanical stimuli produces distinct active-like states in the AT1 receptor
Nature Communications (2023)
-
Anionic phospholipids control mechanisms of GPCR-G protein recognition
Nature Communications (2023)
-
Membrane curvature governs the distribution of Piezo1 in live cells
Nature Communications (2022)
-
Lipidomic and biophysical homeostasis of mammalian membranes counteracts dietary lipid perturbations to maintain cellular fitness
Nature Communications (2020)