Abstract
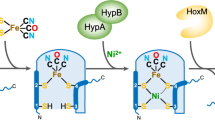
Hydrogenases are highly active enzymes for hydrogen production and oxidation. [NiFeSe] hydrogenases, in which selenocysteine is a ligand to the active site Ni, have high catalytic activity and a bias for H2 production. In contrast to [NiFe] hydrogenases, they display reduced H2 inhibition and are rapidly reactivated after contact with oxygen. Here we report an expression system for production of recombinant [NiFeSe] hydrogenase from Desulfovibrio vulgaris Hildenborough and study of a selenocysteine-to-cysteine variant (Sec489Cys) in which, for the first time, a [NiFeSe] hydrogenase was converted to a [NiFe] type. This modification led to severely reduced Ni incorporation, revealing the direct involvement of this residue in the maturation process. The Ni-depleted protein could be partly reconstituted to generate an enzyme showing much lower activity and inactive states characteristic of [NiFe] hydrogenases. The Ni-Sec489Cys variant shows that selenium has a crucial role in protection against oxidative damage and the high catalytic activities of the [NiFeSe] hydrogenases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Vincent, K.A., Parkin, A. & Armstrong, F.A. Investigating and exploiting the electrocatalytic properties of hydrogenases. Chem. Rev. 107, 4366–4413 (2007).
King, P.W. Designing interfaces of hydrogenase-nanomaterial hybrids for efficient solar conversion. Biochim. Biophys. Acta 1827, 949–957 (2013).
Lubitz, W., Ogata, H., Rüdiger, O. & Reijerse, E. Hydrogenases. Chem. Rev. 114, 4081–4148 (2014).
Simmons, T.R., Berggren, G., Bacchi, M., Fontecave, M. & Artero, V. Mimicking hydrogenases: From biomimetics to artificial enzymes. Coord. Chem. Rev. 270–271, 127–150 (2014).
Friedrich, B., Fritsch, J. & Lenz, O. Oxygen-tolerant hydrogenases in hydrogen-based technologies. Curr. Opin. Biotechnol. 22, 358–364 (2011).
Wakerley, D.W. & Reisner, E. Oxygen-tolerant proton reduction catalysis: much O2 about nothing? Energy Environ. Sci. 8, 2283–2295 (2015).
Brown, K.A., Wilker, M.B., Boehm, M., Dukovic, G. & King, P.W. Characterization of photochemical processes for H2 production by CdS nanorod-[FeFe] hydrogenase complexes. J. Am. Chem. Soc. 134, 5627–5636 (2012).
Hambourger, M. et al. [FeFe]-hydrogenase-catalyzed H2 production in a photoelectrochemical biofuel cell. J. Am. Chem. Soc. 130, 2015–2022 (2008).
De Lacey, A.L., Fernandez, V.M., Rousset, M. & Cammack, R. Activation and inactivation of hydrogenase function and the catalytic cycle: spectroelectrochemical studies. Chem. Rev. 107, 4304–4330 (2007).
Fritsch, J., Lenz, O. & Friedrich, B. Structure, function and biosynthesis of O-tolerant hydrogenases. Nat. Rev. Microbiol. 11, 106–114 (2013).
Baltazar, C.S.A. et al. Nickel–iron–selenium hydrogenases—an overview. Int. J. Inorg. Chem. 2011, 948–962 (2011).
Wombwell, C., Caputo, C.A. & Reisner, E. [NiFeSe]-hydrogenase chemistry. Acc. Chem. Res. 48, 2858–2865 (2015).
Valente, F.M.A. et al. Hydrogenases in Desulfovibrio vulgaris Hildenborough: structural and physiologic characterisation of the membrane-bound [NiFeSe] hydrogenase. J. Biol. Inorg. Chem. 10, 667–682 (2005).
Rüdiger, O. et al. Enzymatic anodes for hydrogen fuel cells based on covalent attachment of Ni-Fe hydrogenases and direct electron transfer to SAM-modified gold electrodes. Electroanalysis 22, 776–783 (2010).
Riethausen, J., Rüdiger, O., Gärtner, W., Lubitz, W. & Shafaat, H.S. Spectroscopic and electrochemical characterization of the [NiFeSe] hydrogenase from Desulfovibrio vulgaris Miyazaki F: reversible redox behavior and interactions between electron transfer centers. ChemBioChem 14, 1714–1719 (2013).
Parkin, A., Goldet, G., Cavazza, C., Fontecilla-Camps, J.C. & Armstrong, F.A. The difference a Se makes? Oxygen-tolerant hydrogen production by the [NiFeSe]-hydrogenase from Desulfomicrobium baculatum. J. Am. Chem. Soc. 130, 13410–13416 (2008).
Reisner, E., Fontecilla-Camps, J.C. & Armstrong, F.A. Catalytic electrochemistry of a [NiFeSe]-hydrogenase on TiO2 and demonstration of its suitability for visible-light driven H2 production. Chem. Commun. (Camb.) 5, 550–552 (2009).
Teixeira, M. et al. Nickel-[iron-sulfur]-selenium-containing hydrogenases from Desulfovibrio baculatus (DSM 1743). Redox centers and catalytic properties. Eur. J. Biochem. 167, 47–58 (1987).
De Lacey, A.L., Gutiérrez-Sánchez, C., Fernández, V.M., Pacheco, I. & Pereira, I.A.C. FTIR spectroelectrochemical characterization of the Ni-Fe-Se hydrogenase from Desulfovibrio vulgaris Hildenborough. J. Biol. Inorg. Chem. 13, 1315–1320 (2008).
Marques, M.C., Coelho, R., De Lacey, A.L., Pereira, I.A. & Matias, P.M. The three-dimensional structure of [NiFeSe] hydrogenase from Desulfovibrio vulgaris Hildenborough: a hydrogenase without a bridging ligand in the active site in its oxidised, “as-isolated” state. J. Mol. Biol. 396, 893–907 (2010).
Ceccaldi, P., Marques, M.C., Fourmond, V., Pereira, I.C. & Léger, C. Oxidative inactivation of NiFeSe hydrogenase. Chem. Commun. (Camb.) 51, 14223–14226 (2015).
Reisner, E., Powell, D.J., Cavazza, C., Fontecilla-Camps, J.C. & Armstrong, F.A. Visible light-driven H(2) production by hydrogenases attached to dye-sensitized TiO(2) nanoparticles. J. Am. Chem. Soc. 131, 18457–18466 (2009).
Caputo, C.A. et al. Photocatalytic hydrogen production using polymeric carbon nitride with a hydrogenase and a bioinspired synthetic Ni catalyst. Angew. Chem. Int. Ed. Engl. 53, 11538–11542 (2014).
Sakai, T., Mersch, D. & Reisner, E. Photocatalytic hydrogen evolution with a hydrogenase in a mediator-free system under high levels of oxygen. Angew. Chem. Int. Ed. Engl. 52, 12313–12316 (2013).
Caputo, C.A., Wang, L.D., Beranek, R. & Reisner, E. Carbon nitride-TiO2 hybrid modified with hydrogenase for visible light driven hydrogen production. Chem. Sci. 6, 5690–5694 (2015).
Mersch, D. et al. Wiring of photosystem II to hydrogenase for photoelectrochemical water splitting. J. Am. Chem. Soc. 137, 8541–8549 (2015).
Gutiérrez-Sanz, Ó. et al. H2-fueled ATP synthesis on an electrode: mimicking cellular respiration. Angew. Chem. Int. Ed. Engl. 55, 6216–6220 (2016).
Berghöfer, Y., Agha-Amiri, K. & Klein, A. Selenium is involved in the negative regulation of the expression of selenium-free [NiFe] hydrogenases in Methanococcus voltae. Mol. Gen. Genet. 242, 369–373 (1994).
Valente, F.M. et al. Selenium is involved in regulation of periplasmic hydrogenase gene expression in Desulfovibrio vulgaris Hildenborough. J. Bacteriol. 188, 3228–3235 (2006).
Hatfield, D.L., Tsuji, P.A., Carlson, B.A. & Gladyshev, V.N. Selenium and selenocysteine: roles in cancer, health, and development. Trends Biochem. Sci. 39, 112–120 (2014).
Reich, H.J. & Hondal, R.J. Why nature chose selenium. ACS Chem. Biol. 11, 821–841 (2016).
Hondal, R.J., Marino, S.M. & Gladyshev, V.N. Selenocysteine in thiol/disulfide-like exchange reactions. Antioxid. Redox Signal. 18, 1675–1689 (2013).
Hondal, R.J. & Ruggles, E.L. Differing views of the role of selenium in thioredoxin reductase. Amino Acids 41, 73–89 (2011).
Snider, G.W., Ruggles, E., Khan, N. & Hondal, R.J. Selenocysteine confers resistance to inactivation by oxidation in thioredoxin reductase: comparison of selenium and sulfur enzymes. Biochemistry 52, 5472–5481 (2013).
Reddie, K.G. & Carroll, K.S. Expanding the functional diversity of proteins through cysteine oxidation. Curr. Opin. Chem. Biol. 12, 746–754 (2008).
Valente, F.M.A. et al. The [NiFeSe] hydrogenase from Desulfovibrio vulgaris Hildenborough is a bacterial lipoprotein lacking a typical lipoprotein signal peptide. FEBS Lett. 581, 3341–3344 (2007).
Marques, M.C., Coelho, R., Pereira, I.A.C. & Matias, P.M. Redox state-dependent changes in the crystal structure of [NiFeSe] hydrogenase from Desulfovibrio vulgaris Hildenborough. Int. J. Hydrogen Energy 38, 8664–8682 (2013).
Garcin, E. et al. The crystal structure of a reduced [NiFeSe] hydrogenase provides an image of the activated catalytic center. Structure 7, 557–566 (1999).
Volbeda, A. et al. Structural foundations for the O2 resistance of Desulfomicrobium baculatum [NiFeSe]-hydrogenase. Chem. Commun. (Camb.) 49, 7061–7063 (2013).
Dementin, S. et al. A glutamate is the essential proton transfer gate during the catalytic cycle of the [NiFe] hydrogenase. J. Biol. Chem. 279, 10508–10513 (2004).
Baltazar, C.S.A., Teixeira, V.H. & Soares, C.M. Structural features of [NiFeSe] and [NiFe] hydrogenases determining their different properties: a computational approach. J. Biol. Inorg. Chem. 17, 543–555 (2012).
Lacasse, M.J. & Zamble, D.B. [NiFe]-hydrogenase maturation. Biochemistry 55, 1689–1701 (2016).
Watanabe, S. et al. Structural basis of a Ni acquisition cycle for [NiFe] hydrogenase by Ni-metallochaperone HypA and its enhancer. Proc. Natl. Acad. Sci. USA 112, 7701–7706 (2015).
Gutiérrez-Sanz, O. et al. Influence of the protein structure surrounding the active site on the catalytic activity of [NiFeSe] hydrogenases. J. Biol. Inorg. Chem. 18, 419–427 (2013).
Leinfelder, W., Zehelein, E., Mandrand-Berthelot, M.A. & Böck, A. Gene for a novel tRNA species that accepts L-serine and cotranslationally inserts selenocysteine. Nature 331, 723–725 (1988).
Zhang, Y., Romero, H., Salinas, G. & Gladyshev, V.N. Dynamic evolution of selenocysteine utilization in bacteria: a balance between selenoprotein loss and evolution of selenocysteine from redox active cysteine residues. Genome Biol. 7, R94 (2006).
Ogata, H., Nishikawa, K. & Lubitz, W. Hydrogens detected by subatomic resolution protein crystallography in a [NiFe] hydrogenase. Nature 520, 571–574 (2015).
Keller, K.L., Wall, J.D. & Chhabra, S. Methods for engineering sulfate reducing bacteria of the genus Desulfovibrio. Methods Enzymol. 497, 503–517 (2011).
Li, M.Z. & Elledge, S.J.S.L.I.C. SLIC: a method for sequence- and ligation-independent cloning. Methods Mol. Biol. 852, 51–59 (2012).
Dementin, S. et al. Changing the ligation of the distal [4Fe4S] cluster in NiFe hydrogenase impairs inter- and intramolecular electron transfers. J. Am. Chem. Soc. 128, 5209–5218 (2006).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
McCoy, A.J. Solving structures of protein complexes by molecular replacement with Phaser. Acta Crystallogr. D Biol. Crystallogr. 63, 32–41 (2007).
Potterton, E., Briggs, P., Turkenburg, M. & Dodson, E. A graphical user interface to the CCP4 program suite. Acta Crystallogr. D Biol. Crystallogr. 59, 1131–1137 (2003).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of COOT. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Acknowledgements
We thank F. Grein and M. Martins for advice and helpful discussions, R. Coelho and S. Silva for help with crystallization procedures and S. Zacarias for experimental assistance; ESRF, DLS and SOLEIL light sources for X-ray data collection; V. Olieric (Swiss Light Source) for the 0.95-Å data collection of the anaerobically purified and crystallized r[NiFeSe] hydrogenase. This work was supported by grants PTDC/BBB-BEP/0934/2012 and PTDC/BBB-BEP/2885/2014 (to I.A.C.P. and P.M.M.) from the Fundação para a Ciência e Tecnologia (FCT/MCTES), by research units GREEN-IT (UID/Multi/04551/2013) funded by FCT/MCTES, and MOSTMICRO (project LISBOA-01-0145-FEDER-007660) co-funded by FCT/MCTES and FEDER funds through COMPETE2020—Programa Operacional Competitividade e Internacionalização (POCI); and by Spanish MINECO/FEDER project CTQ2015-71290-R (to A.L.D.L.). M.C.M. was a recipient of fellowship SFRH/BD/60879/2009 and C.T. was a recipient of predoctoral contract BES-2013-064099 from MINECO. This work was also supported by the European Community's Seventh Framework Program (FP7/2007–2013) under grant agreement 283570 (BioStruct-X).
Author information
Authors and Affiliations
Contributions
I.A.C.P., P.M.M. and M.C.M. conceived the study. A.R.R., K.L.K., J.D.W. and M.C.M. carried out the molecular biology work. M.C.M. and I.A.C.P. produced and characterized hydrogenase variants. M.C.M. and P.M.M. were involved in crystallization and structure determination. O.G.-S., C.T. and A.L.D.L. carried out FTIR and MS experiments. I.A.C.P., P.M.M. and M.C.M. wrote the manuscript with input from other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–2 and Supplementary Figures 1–9 (PDF 1516 kb)
Rights and permissions
About this article
Cite this article
Marques, M., Tapia, C., Gutiérrez-Sanz, O. et al. The direct role of selenocysteine in [NiFeSe] hydrogenase maturation and catalysis. Nat Chem Biol 13, 544–550 (2017). https://doi.org/10.1038/nchembio.2335
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2335
This article is cited by
-
Selenium chemistry for spatio-selective peptide and protein functionalization
Nature Reviews Chemistry (2024)
-
Promising approaches for the assembly of the catalytically active, recombinant Desulfomicrobium baculatum hydrogenase with substitutions at the active site
Microbial Cell Factories (2023)
-
Stepwise assembly of the active site of [NiFe]-hydrogenase
Nature Chemical Biology (2023)
-
Photosystem II for photoelectrochemical hydrogen production
Biophysical Reviews (2023)
-
Fast CO2 hydration kinetics impair heterogeneous but improve enzymatic CO2 reduction catalysis
Nature Chemistry (2022)