Abstract
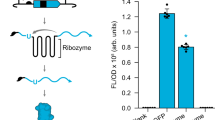
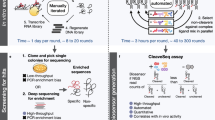
Biosensors that transduce target chemical and biochemical inputs into genetic outputs are essential for bioengineering and synthetic biology. Current biosensor design strategies are often limited by a low signal-to-noise ratio, the extensive optimization required for each new input, and poor performance in mammalian cells. Here we report the development of a proximity-dependent split RNA polymerase (RNAP) as a general platform for biosensor engineering. After discovering that interactions between fused proteins modulate the assembly of a split T7 RNAP, we optimized the split RNAP components for protein–protein interaction detection by phage-assisted continuous evolution (PACE). We then applied the resulting activity-responsive RNAP (AR) system to create biosensors that can be activated by light and small molecules, demonstrating the 'plug-and-play' nature of the platform. Finally, we validated that ARs can interrogate multidimensional protein–protein interactions and trigger RNA nanostructure production, protein synthesis, and gene knockdown in mammalian systems, illustrating the versatility of ARs in synthetic biology applications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ruder, W.C., Lu, T. & Collins, J.J. Synthetic biology moving into the clinic. Science 333, 1248–1252 (2011).
Lienert, F., Lohmueller, J.J., Garg, A. & Silver, P.A. Synthetic biology in mammalian cells: next generation research tools and therapeutics. Nat. Rev. Mol. Cell Biol. 15, 95–107 (2014).
Packer, M.S. & Liu, D.R. Methods for the directed evolution of proteins. Nat. Rev. Genet. 16, 379–394 (2015).
Zhang, J., Jensen, M.K. & Keasling, J.D. Development of biosensors and their application in metabolic engineering. Curr. Opin. Chem. Biol. 28, 1–8 (2015).
Benenson, Y. RNA-based computation in live cells. Curr. Opin. Biotechnol. 20, 471–478 (2009).
Green, A.A., Silver, P.A., Collins, J.J. & Yin, P. Toehold switches: de-novo-designed regulators of gene expression. Cell 159, 925–939 (2014).
Brophy, J.A. & Voigt, C.A. Principles of genetic circuit design. Nat. Methods 11, 508–520 (2014).
Copeland, M.F., Politz, M.C., Johnson, C.B., Markley, A.L. & Pfleger, B.F. A transcription activator-like effector (TALE) induction system mediated by proteolysis. Nat. Chem. Biol. 12, 254–260 (2016).
Putz, U., Skehel, P. & Kuhl, D. A tri-hybrid system for the analysis and detection of RNA--protein interactions. Nucleic Acids Res. 24, 4838–4840 (1996).
Fields, S. & Song, O. A novel genetic system to detect protein-protein interactions. Nature 340, 245–246 (1989).
SenGupta, D.J. et al. A three-hybrid system to detect RNA-protein interactions in vivo. Proc. Natl. Acad. Sci. USA 93, 8496–8501 (1996).
Baker, K. et al. Chemical complementation: a reaction-independent genetic assay for enzyme catalysis. Proc. Natl. Acad. Sci. USA 99, 16537–16542 (2002).
Konermann, S. et al. Optical control of mammalian endogenous transcription and epigenetic states. Nature 500, 472–476 (2013).
Polstein, L.R. & Gersbach, C.A. A light-inducible CRISPR-Cas9 system for control of endogenous gene activation. Nat. Chem. Biol. 11, 198–200 (2015).
Nuñez, J.K., Harrington, L.B. & Doudna, J.A. Chemical and biophysical modulation of Cas9 for tunable genome engineering. ACS Chem. Biol. 11, 681–688 (2016).
Martin, F. Fifteen years of the yeast three-hybrid system: RNA-protein interactions under investigation. Methods 58, 367–375 (2012).
Church, G.M., Elowitz, M.B., Smolke, C.D., Voigt, C.A. & Weiss, R. Realizing the potential of synthetic biology. Nat. Rev. Mol. Cell Biol. 15, 289–294 (2014).
Yen, L. et al. Exogenous control of mammalian gene expression through modulation of RNA self-cleavage. Nature 431, 471–476 (2004).
Winkler, W.C. & Breaker, R.R. Regulation of bacterial gene expression by riboswitches. Annu. Rev. Microbiol. 59, 487–517 (2005).
Culler, S.J., Hoff, K.G. & Smolke, C.D. Reprogramming cellular behavior with RNA controllers responsive to endogenous proteins. Science 330, 1251–1255 (2010).
Feng, J. et al. A general strategy to construct small molecule biosensors in eukaryotes. eLife 4, e10606 (2015).
Lee, J.H. et al. Highly multiplexed subcellular RNA sequencing in situ. Science 343, 1360–1363 (2014).
Zamft, B.M. et al. Measuring cation dependent DNA polymerase fidelity landscapes by deep sequencing. PLoS One 7, e43876 (2012).
Pu, J., Chronis, I., Ahn, D. & Dickinson, B.C. A panel of protease-responsive RNA polymerases respond to biochemical signals by production of defined RNA outputs in live cells. J. Am. Chem. Soc. 137, 15996–15999 (2015).
Segall-Shapiro, T.H., Meyer, A.J., Ellington, A.D., Sontag, E.D. & Voigt, C.A.A. A 'resource allocator' for transcription based on a highly fragmented T7 RNA polymerase. Mol. Syst. Biol. 10, 742 (2014).
Shis, D.L. & Bennett, M.R. Library of synthetic transcriptional AND gates built with split T7 RNA polymerase mutants. Proc. Natl. Acad. Sci. USA 110, 5028–5033 (2013).
Shekhawat, S.S. & Ghosh, I. Split-protein systems: beyond binary protein-protein interactions. Curr. Opin. Chem. Biol. 15, 789–797 (2011).
Kerppola, T.K. Visualization of molecular interactions using bimolecular fluorescence complementation analysis: characteristics of protein fragment complementation. Chem. Soc. Rev. 38, 2876–2886 (2009).
Hu, C.D. & Kerppola, T.K. Simultaneous visualization of multiple protein interactions in living cells using multicolor fluorescence complementation analysis. Nat. Biotechnol. 21, 539–545 (2003).
Dickinson, B.C., Leconte, A.M., Allen, B., Esvelt, K.M. & Liu, D.R. Experimental interrogation of the path dependence and stochasticity of protein evolution using phage-assisted continuous evolution. Proc. Natl. Acad. Sci. USA 110, 9007–9012 (2013).
Ellefson, J.W. et al. Directed evolution of genetic parts and circuits by compartmentalized partnered replication. Nat. Biotechnol. 32, 97–101 (2014).
Ghosh, I., Hamilton, A.D. & Regan, L. Antiparallel leucine zipper-directed protein reassembly: application to the green fluorescent protein. J. Am. Chem. Soc. 122, 5658–5659 (2000).
Magliery, T.J. et al. Detecting protein-protein interactions with a green fluorescent protein fragment reassembly trap: scope and mechanism. J. Am. Chem. Soc. 127, 146–157 (2005).
Steitz, T.A. The structural changes of T7 RNA polymerase from transcription initiation to elongation. Curr. Opin. Struct. Biol. 19, 683–690 (2009).
Esvelt, K.M., Carlson, J.C. & Liu, D.R. A system for the continuous directed evolution of biomolecules. Nature 472, 499–503 (2011).
Badran, A.H. et al. Continuous evolution of Bacillus thuringiensis toxins overcomes insect resistance. Nature 533, 58–63 (2016).
Carlson, J.C., Badran, A.H., Guggiana-Nilo, D.A. & Liu, D.R. Negative selection and stringency modulation in phage-assisted continuous evolution. Nat. Chem. Biol. 10, 216–222 (2014).
Dickinson, B.C., Packer, M.S., Badran, A.H. & Liu, D.R. A system for the continuous directed evolution of proteases rapidly reveals drug-resistance mutations. Nat. Commun. 5, 5352 (2014).
Hubbard, B.P. et al. Continuous directed evolution of DNA-binding proteins to improve TALEN specificity. Nat. Methods 12, 939–942 (2015).
Leconte, A.M. et al. A population-based experimental model for protein evolution: effects of mutation rate and selection stringency on evolutionary outcomes. Biochemistry 52, 1490–1499 (2013).
Cheetham, G.M. & Steitz, T.A. Structure of a transcribing T7 RNA polymerase initiation complex. Science 286, 2305–2309 (1999).
Tahirov, T.H. et al. Structure of a T7 RNA polymerase elongation complex at 2.9 A resolution. Nature 420, 43–50 (2002).
Guntas, G. et al. Engineering an improved light-induced dimer (iLID) for controlling the localization and activity of signaling proteins. Proc. Natl. Acad. Sci. USA 112, 112–117 (2015).
Rivera, V.M. et al. A humanized system for pharmacologic control of gene expression. Nat. Med. 2, 1028–1032 (1996).
Gossen, M. & Bujard, H. Tight control of gene expression in mammalian cells by tetracycline-responsive promoters. Proc. Natl. Acad. Sci. USA 89, 5547–5551 (1992).
Paulmurugan, R. et al. A novel estrogen receptor intramolecular folding-based titratable transgene expression system. Mol. Ther. 17, 1703–1711 (2009).
Blakeley, B.D., Chapman, A.M. & McNaughton, B.R. Split-superpositive GFP reassembly is a fast, efficient, and robust method for detecting protein-protein interactions in vivo. Mol. Biosyst. 8, 2036–2040 (2012).
Kerppola, T.K. Bimolecular fluorescence complementation (BiFC) analysis as a probe of protein interactions in living cells. Annu. Rev. Biophys. 37, 465–487 (2008).
Filonov, G.S., Kam, C.W., Song, W. & Jaffrey, S.R. In-gel imaging of RNA processing using broccoli reveals optimal aptamer expression strategies. Chem. Biol. 22, 649–660 (2015).
Ringquist, S. et al. Translation initiation in Escherichia coli: sequences within the ribosome-binding site. Mol. Microbiol. 6, 1219–1229 (1992).
Gibson, D.G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Acknowledgements
We thank I. Chronis (University of Chicago), Y. Koh (University of Chicago), and D. Ahn (University of Chicago) for technical assistance, and D. Liu (Harvard University), B. McNaughton (Colorado State University), A. Deiters (University of Pittsburgh), B. Glick (University of Chicago), M. Glotzer (University of Chicago), Y. Krishnan (University of Chicago), J. Thornton (University of Chicago), and Y. Weizmann (University of Chicago) for supplying equipment and materials. This work was supported by the University of Chicago, the Cancer Research Foundation, the National Institute of General Medical Sciences of the National Institutes of Health (R35 GM119840) to B.C.D., the University of Chicago Medicine Comprehensive Cancer Center (P30 CA14599), and the National Center for Advancing Translational Sciences of the National Institutes of Health (UL1 TR000430). J.Z.-B. was supported by a Chemical Biology Training Grant from the US National Institutes of Health (T32GM008720).
Author information
Authors and Affiliations
Contributions
J.Z.-B., J.P., and B.C.D. cloned and validated all materials and performed transcription assays. J.P. and B.C.D. performed PACE experiments. J.Z.-B. and J.P. performed cell culture experiments. J.Z.-B., J.P., and B.C.D. designed experimental strategies and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors have filed a provisional patent application on the AR system.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–3 and Supplementary Figures 1–6. (PDF 4992 kb)
Rights and permissions
About this article
Cite this article
Pu, J., Zinkus-Boltz, J. & Dickinson, B. Evolution of a split RNA polymerase as a versatile biosensor platform. Nat Chem Biol 13, 432–438 (2017). https://doi.org/10.1038/nchembio.2299
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2299
This article is cited by
-
An orthogonalized PYR1-based CID module with reprogrammable ligand-binding specificity
Nature Chemical Biology (2024)
-
Synthetic microbiology in sustainability applications
Nature Reviews Microbiology (2024)
-
Evolution of an adenine base editor into a small, efficient cytosine base editor with low off-target activity
Nature Biotechnology (2023)
-
Target-dependent RNA polymerase as universal platform for gene expression control in response to intracellular molecules
Nature Communications (2023)
-
Synthetic virology approaches to improve the safety and efficacy of oncolytic virus therapies
Nature Communications (2023)