Abstract
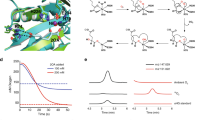
Substrate channeling has emerged as a common mechanism for enzymatic intermediate transfer. A conspicuous gap in knowledge concerns the use of covalent lysine imines in the transfer of carbonyl-group-containing intermediates, despite their wideuse in enzymatic catalysis. Here we show how imine chemistry operates in the transfer of covalent intermediates in pyridoxal 5′-phosphate biosynthesis by the Arabidopsis thaliana enzyme Pdx1. An initial ribose 5-phosphate lysine imine is converted to the chromophoric I320 intermediate, simultaneously bound to two lysine residues and partially vacating the active site, which creates space for glyceraldehyde 3-phosphate to bind. Crystal structures show how substrate binding, catalysis and shuttling are coupled to conformational changes around strand β6 of the Pdx1 (βα)8-barrel. The dual-specificity active site and imine relay mechanism for migration of carbonyl intermediates provide elegant solutions to the challenge of coordinating a complex sequence of reactions that follow a path of over 20 Å between substrate- and product-binding sites.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
23 January 2017
In the version of this article initially published online, reference to another recently published structure of the I320 complex was inadvertently omitted. The paper reporting the structure, which is now included as ref. 43 of the article, is Robinson, G.C., Kaufmann, M., Roux, C. & Fitzpatrick, T.B. Structural definition of the lysine swing in Arabidopsis thaliana PDX1: intermediate channeling facilitating vitamin B6 biosynthesis. Proc. Natl. Acad. Sci. USA 113, E5821–E5829 (2016). The error has been corrected in the print, PDF and HTML versions of this article.
References
Ehrenshaft, M., Bilski, P., Li, M.Y., Chignell, C.F. & Daub, M.E. A highly conserved sequence is a novel gene involved in de novo vitamin B6 biosynthesis. Proc. Natl. Acad. Sci. USA 96, 9374–9378 (1999).
Mooney, S. & Hellmann, H. Vitamin B6: killing two birds with one stone? Phytochemistry 71, 495–501 (2010).
Percudani, R. & Peracchi, A. The B6 database: a tool for the description and classification of vitamin B6-dependent enzymatic activities and of the corresponding protein families. BMC Bioinformatics 10, 273 (2009).
Mittenhuber, G. Phylogenetic analyses and comparative genomics of vitamin B6 (pyridoxine) and pyridoxal phosphate biosynthesis pathways. J. Mol. Microbiol. Biotechnol. 3, 1–20 (2001).
Das, S. et al. Biology's new Rosetta stone. Nature 385, 29–30 (1997).
Fitzpatrick, T.B. et al. Two independent routes of de novo vitamin B6 biosynthesis: not that different after all. Biochem. J. 407, 1–13 (2007).
Raschle, T., Amrhein, N. & Fitzpatrick, T.B. On the two components of pyridoxal 5′-phosphate synthase from Bacillus subtilis. J. Biol. Chem. 280, 32291–32300 (2005).
Burns, K.E., Xiang, Y., Kinsland, C.L., McLafferty, F.W. & Begley, T.P. Reconstitution and biochemical characterization of a new pyridoxal-5′-phosphate biosynthetic pathway. J. Am. Chem. Soc. 127, 3682–3683 (2005).
Hanes, J.W. et al. Mechanistic studies on pyridoxal phosphate synthase: the reaction pathway leading to a chromophoric intermediate. J. Am. Chem. Soc. 130, 3043–3052 (2008).
Raschle, T. et al. Reaction mechanism of pyridoxal 5′-phosphate synthase. Detection of an enzyme-bound chromophoric intermediate. J. Biol. Chem. 282, 6098–6105 (2007).
Strohmeier, M. et al. Structure of a bacterial pyridoxal 5′-phosphate synthase complex. Proc. Natl. Acad. Sci. USA 103, 19284–19289 (2006).
Zhu, J., Burgner, J.W., Harms, E., Belitsky, B.R. & Smith, J.L. A new arrangement of (β/α)8 barrels in the synthase subunit of PLP synthase. J. Biol. Chem. 280, 27914–27923 (2005).
Hanes, J.W., Keresztes, I. & Begley, T.P. 13C NMR snapshots of the complex reaction coordinate of pyridoxal phosphate synthase. Nat. Chem. Biol. 4, 425–430 (2008).
Tambasco-Studart, M. et al. Vitamin B6 biosynthesis in higher plants. Proc. Natl. Acad. Sci. USA 102, 13687–13692 (2005).
Guédez, G. et al. Assembly of the eukaryotic PLP-synthase complex from Plasmodium and activation of the Pdx1 enzyme. Structure 20, 172–184 (2012).
Smith, A.M., Brown, W.C., Harms, E. & Smith, J.L. Crystal structures capture three states in the catalytic cycle of a pyridoxal phosphate (PLP) synthase. J. Biol. Chem. 290, 5226–5239 (2015).
Zein, F. et al. Structural insights into the mechanism of the PLP synthase holoenzyme from Thermotoga maritime. Biochemistry 45, 14609–14620 (2006).
Wallner, S., Neuwirth, M., Flicker, K., Tews, I. & Macheroux, P. Dissection of contributions from invariant amino acids to complex formation and catalysis in the heteromeric pyridoxal 5-phosphate synthase complex from Bacillus subtilis. Biochemistry 48, 1928–1935 (2009).
Gengenbacher, M. et al. Vitamin B6 biosynthesis by the malaria parasite Plasmodium falciparum: biochemical and structural insights. J. Biol. Chem. 281, 3633–3641 (2006).
Raushel, F.M., Thoden, J.B. & Holden, H.M. Enzymes with molecular tunnels. Acc. Chem. Res. 36, 539–548 (2003).
Hanes, J.W., Keresztes, I. & Begley, T.P. Trapping of a chromophoric intermediate in the Pdx1-catalyzed biosynthesis of pyridoxal 5′-phosphate. Angew. Chem. Int. Ed. Engl. 47, 2102–2105 (2008).
Zhang, X. et al. Structural insights into the catalytic mechanism of the yeast pyridoxal 5-phosphate synthase Snz1. Biochem. J. 432, 445–450 (2010).
von Stetten, D. et al. In crystallo optical spectroscopy (icOS) as a complementary tool on the macromolecular crystallography beamlines of the ESRF. Acta Crystallogr. D Biol. Crystallogr. 71, 15–26 (2015).
Ho, M.-C., Ménétret, J.-F., Tsuruta, H. & Allen, K.N. The origin of the electrostatic perturbation in acetoacetate decarboxylase. Nature 459, 393–397 (2009).
Borshchevskiy, V. et al. Low-dose X-ray radiation induces structural alterations in proteins. Acta Crystallogr. D Biol. Crystallogr. 70, 2675–2685 (2014).
Dubnovitsky, A.P., Ravelli, R.B., Popov, A.N. & Papageorgiou, A.C. Strain relief at the active site of phosphoserine aminotransferase induced by radiation damage. Protein Sci. 14, 1498–1507 (2005).
Weik, M. et al. Specific chemical and structural damage to proteins produced by synchrotron radiation. Proc. Natl. Acad. Sci. USA 97, 623–628 (2000).
Pearson, A.R. & Owen, R.L. Combining X-ray crystallography and single-crystal spectroscopy to probe enzyme mechanisms. Biochem. Soc. Trans. 37, 378–381 (2009).
Berglund, G.I. et al. The catalytic pathway of horseradish peroxidase at high resolution. Nature 417, 463–468 (2002).
Zeldin, O.B., Gerstel, M. & Garman, E.F. Optimizing the spatial distribution of dose in X-ray macromolecular crystallography. J. Synchrotron Radiat. 20, 49–57 (2013).
Foadi, J. et al. Clustering procedures for the optimal selection of data sets from multiple crystals in macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 69, 1617–1632 (2013).
Derrer, B. et al. Defining the structural requirements for ribose 5-phosphate-binding and intersubunit cross-talk of the malarial pyridoxal 5-phosphate synthase. FEBS Lett. 584, 4169–4174 (2010).
Raschle, T. et al. Intersubunit cross-talk in pyridoxal 5′-phosphate synthase, coordinated by the C terminus of the synthase subunit. J. Biol. Chem. 284, 7706–7718 (2009).
Moccand, C., Kaufmann, M. & Fitzpatrick, T.B. It takes two to tango: defining an essential second active site in pyridoxal 5′-phosphate synthase. PLoS One 6, e16042 (2011).
Nagano, N., Orengo, C.A. & Thornton, J.M. One fold with many functions: the evolutionary relationships between TIM barrel families based on their sequences, structures and functions. J. Mol. Biol. 321, 741–765 (2002).
Reeksting, S.B. et al. Exploring inhibition of Pdx1, a component of the PLP synthase complex of the human malaria parasite Plasmodium falciparum. Biochem. J. 449, 175–187 (2013).
Neuwirth, M. et al. X-ray crystal structure of Saccharomyces cerevisiae Pdx1 provides insights into the oligomeric nature of PLP synthases. FEBS Lett. 583, 2179–2186 (2009).
Jenni, S. et al. Structure of fungal fatty acid synthase and implications for iterative substrate shuttling. Science 316, 254–261 (2007).
Wu, N., Tsuji, S.Y., Cane, D.E. & Khosla, C. Assessing the balance between protein–protein interactions and enzyme-substrate interactions in the channeling of intermediates between polyketide synthase modules. J. Am. Chem. Soc. 123, 6465–6474 (2001).
Tanovic, A., Samel, S.A., Essen, L.O. & Marahiel, M.A. Crystal structure of the termination module of a nonribosomal peptide synthetase. Science 321, 659–663 (2008).
Pares, S., Cohen-Addad, C., Sieker, L., Neuburger, M. & Douce, R. X-ray structure determination at 2.6-A resolution of a lipoate-containing protein: the H-protein of the glycine decarboxylase complex from pea leaves. Proc. Natl. Acad. Sci. USA 91, 4850–4853 (1994).
Reed, L.J. Multienzyme complexes. Acc. Chem. Res. 7, 40–46 (1974).
Robinson, G.C., Kaufmann, M., Roux, C. & Fitzpatrick, T.B. Structural definition of the lysine swing in Arabidopsis thaliana PDX1: intermediate channeling facilitating vitamin B6 biosynthesis Proc. Natl. Acad. Sci. USA 113, E5821–E5829 (2016).
McCarthy, A.A. et al. A decade of user operation on the macromolecular crystallography MAD beamline ID14-4 at the ESRF J. Synchrotron Radiat. 16, 803–812 (2009).
McGeehan, J. et al. Colouring cryo-cooled crystals: online microspectrophotometry. J. Synchrotron Radiat. 16, 163–172 (2009).
Wakatsuki, S. et al. ID14 'Quadriga', a beamline for protein crystallography at the ESRF. J. Synchrotron Radiat. 5, 215–221 (1998).
de Sanctis, D. et al. ID29: a high-intensity highly automated ESRF beamline for macromolecular crystallography experiments exploiting anomalous scattering. J. Synchrotron Radiat. 19, 455–461 (2012).
Nurizzo, D. et al. The ID23-1 structural biology beamline at the ESRF. J. Synchrotron Radiat. 13, 227–238 (2006).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P.R. & Murshudov, G.N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
Gildea, R.J. et al. New methods for indexing multi-lattice diffraction data. Acta Crystallogr. D Biol. Crystallogr. 70, 2652–2666 (2014).
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 1022–1025 (1997).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Adams, P.D. et al. The Phenix software for automated determination of macromolecular structures. Methods 55, 94–106 (2011).
Lebedev, A.A. et al. JLigand: a graphical tool for the CCP4 template-restraint library. Acta Crystallogr. D Biol. Crystallogr. 68, 431–440 (2012).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Zeldin, O.B., Gerstel, M. & Garman, E.F. RADDOSE-3D: time- and space-resolved modelling of dose in macromolecular crystallography. J. Appl. Crystallogr. 46, 1225–1230 (2013).
Acknowledgements
We thank W.J. Anderson, H. Clarke, C. Phippen (Southampton), A. Wessling and N. Kwak (Heidelberg), for their experimental contributions; T. Fitzpatrick (Geneva, Switzerland) for the generous gift of the expression plasmid for Arabidopsis Pdx1.3; S. Findlow and C. Holes at the Macromolecular Crystallisation Facility, Centre for Biological Sciences, and P. Horton and S. Coles at the Southampton Diffraction Centre, Chemistry, both University of Southampton; J. Kopp and C. Siegmann from the crystallization platform of the Cluster of Excellence CellNetworks, Heidelberg; staff at the Diamond Light Source and the European Synchrotron Radiation Facility for access and excellent user support without which this project would not have been possible; P. Carpentier, M. Weik, G. Gotthard and D. von Stetten at ESRF for support during data collection and online spectroscopy and D. Flot and G. Leonard for flexible access to the ESRF beamlines; A. Douangamath, P. Aller, R. Owen and M. Walsh for support at Diamond beamlines; B. Kappes (Erlangen) and P. Macheroux (Graz) for critical discussions; and L. Kinsland for assistance in preparation of the manuscript. This work was supported in parts by grants by the European Commission (VITBIOMAL-012158) and by the Deutsche Forschungsgemeinschaft (DFG) (TE368) to I.T., by the NIH (DK44083) and by the Robert A. Welch Foundation (A-0034) to T.P.B., by ESRF Mx1461, Mx1732, RADDAM and Diamond Light Source Mx8891 to I.T. M.J.R. was supported by a joint studentship between Diamond Light Source and the University of Southampton.
Author information
Authors and Affiliations
Contributions
M.J.R., V.W., G.G., S.W., M.S. and J.W.H. performed protein expression, purification and enzymatic essays. M.J.R., V.W., G.G., S.W., M.S. and I.T. performed crystallization and X-ray diffraction experiments. M.J.R., V.W., A.R. and I.T. performed online spectroscopy experiments. M.J.R., V.W., Y.Z., I.S., S.E.E., T.P.B. and I.T. performed crystallographic analysis and data deposition. M.J.R., A.R., G.E., T.P.B. and I.T. performed spectroscopic data analysis. M.J.R., G.E., I.S., S.E.E., T.P.B. and I.T. wrote the paper. M.J.R. and V.W. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Table 1 and Supplementary Figures 1–6. (PDF 970 kb)
Rights and permissions
About this article
Cite this article
Rodrigues, M., Windeisen, V., Zhang, Y. et al. Lysine relay mechanism coordinates intermediate transfer in vitamin B6 biosynthesis. Nat Chem Biol 13, 290–294 (2017). https://doi.org/10.1038/nchembio.2273
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2273
This article is cited by
-
Protein structural biology using cell-free platform from wheat germ
Advanced Structural and Chemical Imaging (2018)