Abstract
High-content screening is transforming drug discovery by enabling simultaneous measurement of multiple features of cellular phenotype that are relevant to therapeutic and toxic activities of compounds. High-content screening studies typically generate immense datasets of image-based phenotypic information, and how best to mine relevant phenotypic data is an unsolved challenge. Here, we introduce factor analysis as a data-driven tool for defining cell phenotypes and profiling compound activities. This method allows a large data reduction while retaining relevant information, and the data-derived factors used to quantify phenotype have discernable biological meaning. We used factor analysis of cells stained with fluorescent markers of cell cycle state to profile a compound library and cluster the hits into seven phenotypic categories. We then compared phenotypic profiles, chemical similarity and predicted protein binding activities of active compounds. By integrating these different descriptors of measured and potential biological activity, we can effectively draw mechanism-of-action inferences.
Similar content being viewed by others
Main
Drug discovery requires integration of chemical and biological knowledge about many compounds in an efficient manner1. Profiling compounds by chemical structure has become increasingly sophisticated, but profiling by biological activity has lagged owing to the difficulty of collecting and integrating different types of biological information, and also because of the large expense of data-rich methods such as mRNA expression profiling. High-content screening (HCS) combines automated microscopy with image analysis to enable phenotypic profiling of compounds based on activities on cells visualized by fluorescence cytology2,3,4. This rapidly developing technology is increasingly used to facilitate both target and lead characterization5,6. The instrumentation and image quantification aspects of HCS, while under constant improvement, are already well advanced7,8,9. Methods for downstream data processing and mining of biological data are by comparison significantly less refined. Most users score for predefined phenotypes of interest, such as nuclear translocation of a transcription factor, largely ignoring the wealth of phenotypic information present in most HCS datasets. Thus, the huge potential of HCS to inform on biological effects relevant to therapeutics and toxicity is largely untapped.
Two problems have limited the use of HCS to report broadly on phenotypic effects of compounds: the large size of the datasets, and the fact that the biological meaning of most of the measurements is unclear. A typical HCS experiment might generate terabytes of image data from which gigabytes of numbers are extracted describing the amount and location of biomolecules on a cell-to-cell basis. Most of these numbers have no obvious biological meaning; for example, though the amount of DNA per nucleus has obvious significance, the importance of other nuclear measures, such as DNA texture or nuclear ellipticity, is much less clear. This leads biologists to ignore the non-obvious measurements, even though they may report usefully on compound activities. Here, we introduce factor analysis to mine HCS datasets. This method was developed more than a century ago10, remains standard in other fields for analyzing large, multidimensional datasets11,12,13,14,15, and was implemented here using standard, commercially available statistics software. It allows a large data reduction and quantifies phenotype using data-derived factors that are biologically interpretable in many cases.
The basic supposition underlying factor analysis is that groups of variables within a multivariate dataset that are highly correlated with each other, but poorly correlated with other variables in the dataset, are likely to be measuring a common underlying trait, or “factor”16. In HCS, this translates to the reasonable supposition that groups of image-based cell features that show highly correlated changes between individual cells following different compound treatments are likely reporting on a common phenotypic property. If this supposition is true, we should often be able to interpret the biological meaning of the factors, even though they were generated directly from the data without biological assumptions. Here, we use cytological markers of cell cycle, HCS and factor analysis to profile the biological effects of a compound library. We find that six factors are sufficient to describe the biological responses, that several of them have interpretable biological meaning and that the responses group the active compounds into seven major categories by phenotypic effects. We then explore how phenotypic profiles of active compounds compare with chemical structure and predicted target profiles. The resulting structure-activity relationships are more information rich than would be possible with a single data type, and they allow us to infer mechanisms of action for some compounds.
Results
Factor analysis of high-content image data
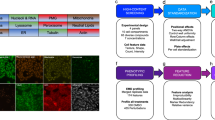
We designed an HCS assay to identify compounds that affect cell proliferation, and to profile their cell cycle phenotype, using fluorescent probes for DNA (Ch1), mitosis (Ch2) and DNA replication (Ch3) (Fig. 1). Probes for Ch1 (Hoechst 33342 dye) and Ch2 (anti-phosphoH3) were standard. To label sites of DNA replication in Ch3 we pulsed cells briefly with 5-ethynl-2-deoxyuridine (EdU) before fixation. Classic bromodeoxyuridine (BrdU) staining is not ideal for HCS because many steps are required to visualize the probe, including a DNA denaturation step that perturbs nuclear morphology. EdU is incorporated into DNA during replication like BrdU, but visualization requires only a single reaction using “click chemistry” to conjugate a rhodamine-azide dye to the ethynyl group (Supplementary Methods online). Images were acquired automatically using 10× objective and widefield imaging. For primary image analysis, the DNA stain was segmented to find nuclei. A nuclear mask was then used to generate 36 cytological features (all nuclear) from the three fluorescent channels (Supplementary Table 1 online). At least 500 cells were scored per treatment in two replicate experiments. We used the common factor model to map these 36 cytological features into a reduced dimensional space defined by a set of six orthogonal factors that reflect the major underlying phenotypic attributes measured in the assay (Fig. 2a,b, Supplementary Methods and Supplementary Data 1 online). The set of features that load substantially on a given factor was used to infer the underlying phenotypic attributes associated with that factor. A representative polar plot of loadings versus cytological features for factor 1 was shown (Fig. 2c). Complete factor structure and underlying phenotypic traits are outlined in Figure 2d, and representative images are shown in Supplementary Figure 1 online. Note the order of numbering the factors is based on the extent to which a given factor accounts for the common variance in the whole dataset.
HeLa cells were grown in 384-well optical plates for 24 h before compound treatment. Compounds were delivered in an automated manner for a final concentration of 10 μM and incubated for approximately 20 h. Cells were then pulsed for 40 min with 500 nM EdU to label sites of nascent DNA replication (yellow), followed by fixation in formaldehyde. Rhodamine-azide was conjugated to EdU by click chemistry. Cells were immunolabeled with rabbit anti-phosphohistone H3 Ser10 (pH3) and a Cy5-conjugated goat anti-rabbit secondary antibody (red). DNA was labeled with Hoechst dye (blue). Automated fluorescence microscopy was carried out using a Cellomics Arrayscan, and images were collected with a 10× PlanFluor objective. Representative images are shown with 20 μm size bars. Individual cell segmentation based on the DNA stain (cytological mask) and quantification was performed using the Cellomics Morphology Explorer algorithm, and 36 cytological features (Supplementary Table 1) were determined for each cell on DNA (Ch1), pH3 (Ch2) and EdU (Ch3) channels. Features were collected for at least 500 cells per well (treatment).
(a) High-content data are contained in an n × m matrix, X consisting of a set of n image-based cytological features measured on m cells. The common factor model maps the n cytological features to a reduced k-dimensional space described by a set of factors F that reflect the major underlying phenotypic attributes measured in the assay. The loading matrix L defines the relationship between the measurements in X to the underlying common factors. The diagonal matrix ε is a matrix of specific variances. (b) The dimensionality of the factor space is determined by an eigenanalysis of the correlation matrix of the data matrix X. The dimension k is determined by Kaiser criterion to be equal to the number of factors with variance greater than unity. Using this criterion we determined that there are six significant factors. (c) The loadings L reflect the correlations between cytological features and the common underlying factors. We used polar plots to visualize these loading patterns and interpret the biological meaning of the underlying factor. Shown here is the loading pattern for factor 1 as an example. (d) The complete factor structure is shown in this schematic. Each of the six factors are drawn with lines connected to the cytological features with which they are most significantly correlated. Our interpretation of the phenotypic attributes characterized by each factor is shown on the right (see also Supplementary Methods and Supplementary Data 1).
Factor 1, which accounts for most of the common variance, loads highly on 12 features, all of which describe the size of the nucleus. Examples of these features include Area-Ch1, TotalIntensity-Ch1, Length-Ch1 and Width-Ch1. Based on this loading pattern we conclude that this is a nuclear size factor. Thus, the most information-rich phenotypic characteristic given our labeling and imaging strategy is the size of the nucleus and the quantity of DNA. Factor 2 loads primarily with four features that describe the extent of EdU probe incorporation. Hence, factor 2 is a DNA replication, or S-phase factor. Factor 3 loads primarily with features that describe DNA concentration (and thus condensation; for example, AvgIntensity-Ch1) and phosphoH3 intensity (for example, AvgIntensity-Ch2) and is thus a mitosis and chromosome condensation factor. Factor 4 is loaded substantially by four features that refer to the shape contour of the nuclear perimeter and is thus a nuclear morphology factor. Factor 5 loads with four features that describe Ch2 texture; that is, the morphology of EdU incorporation. It is statistically distinct from factor 2 and must report on some particular aspect of DNA replication, such as early versus late S phase. Factor 6 reports mainly on nuclear shape. Taken together, we reduced a dataset of 36 measured cytological features from ∼106 cells (∼7 GB) to six common underlying factors scored for ∼104 wells (∼3 MB). Moreover, these common underlying factors reflect a set of orthogonal phenotypic attributes that account for almost all of the covariance relationships shown in the image-based cytological features measured on each cell in our assay.
Factor-based phenotypic compound profiling
We used our high-content image assay to screen and profile a library of 6,547 compounds derived from a diversity library (21%), a natural products library (58%) and a library of known bioactive compounds (21%); all compounds were assayed in duplicate at a single dose of 10 μM for 20 h (Fig. 3a). Dose-response studies using a panel of known cytotoxic compounds with diverse mechanisms of action indicated the appropriateness of these dose and time conditions for phenotypic profiling (Supplementary Fig. 2 online). Based on the six-factor model, we used regression to estimate scores for each factor (that is, nuclear size, replication, mitosis, nuclear morphology, EdU texture and nuclear ellipticity) on a cell-by-cell basis for each treatment. We summarized each compound treatment effect as the mean score on each of the six factors (that is, a well average).
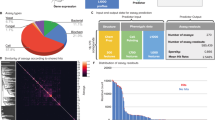
(a) We screened 6,547 compounds from three libraries. Our screen was performed in two replicate experiments. We established a factor-based phenotypic response metric that reflects the distance in factor space from a treatment to the untreated control population. This metric projects the multidimensional phenotype onto a single response dimension, enables a standard comparison between compounds with various bioactivities, and facilitates hit identification independent of the specific phenotype. Hits were defined as compounds in the top 5% based on phenotypic response in both replicate experiments. These filter criteria resulted in 211 bioactive compounds, or ∼3% of the total. (b) Pie charts indicating the fraction of each library in our screening set and hits set. We observe an enrichment of known bioactive compounds in our hit collection. (c) A scatter plot comparing the factor-based phenotypic response from both replicate experiments. Compounds in the top 2–5% are colored blue, the top 1% are green and non-hits are gray. (d) We performed hierarchical clustering of mean factor scores for each of the 211 hits. Clustering is based on Ward's linkage criteria and the half Euclidean distance metric. The position of compounds within the top 1% and 2–5% based on phenotypic response is shown. Cluster analysis of a reduced dataset consisting of all nonproprietary compounds (n = 173) retains the overall hierarchical structure (Supplementary Fig. 3).
We expected our compound library to contain multiple bioactive compounds with various distinct targets and mechanisms of action, and consequently we expected it to generate unique phenotypic readouts on the six orthogonal factors. To score the strength of phenotypic perturbation independent of precise phenotype, we computed the Euclidean distance between each compound and the average control (untreated) phenotype for a composite vector consisting of all factor scores for that compound. This Euclidean distance metric projects the multidimensional phenotype onto a single phenotypic response dimension and allows us to call “hits” independent of their exact phenotype.
We defined hits as compounds whose phenotypic response (that is, distance) was in the top 5% in both replicate experiments; this resulted in 211 compound hits, which is ∼3% of the total screening set. Our hit set was enriched in compounds derived from the library of bioactive compounds (Fig. 3b). This enrichment was most pronounced when we examined the strongest bioactive compounds in the top 1% distance group. In this set, 48% of the compounds were derived from the bioactive library, compared with 21% in the entire screening set. This indicates our strategy is effective at identifying compounds with substantial biological activity. We observed a generally good correspondence between the two replicate experiments (Fig. 3c).
We next profiled the biological activity of the hit compounds using unsupervised hierarchical clustering of the factor scores. This revealed seven primary clusters that we termed “phenotypes” (Fig. 3d, Supplementary Data 2 and Supplementary Fig. 3 online). We can begin to interpret these phenotypes by looking at how the factors change, and also where compounds with known biological activity are positioned (discussed below). For example, phenotypes 1 and 2, which show high chromosome condensation, correspond in large part to mitotic arrest; phenotype 4, which shows generally high chromosome condensation but also decreased nuclear size, corresponds in large part to apoptosis; phenotype 5, which shows increased DNA replication, and phenotypes 6 and 7, which show increased nuclear area, decreased DNA replication and decreased chromosome condensation, probably correspond to cell cycle exit in G1, which is generally understood to increase nuclear cross-sectional area. The strongest hits in our screen (top 1%) mostly affect mitotic progression and cell survival, whereas weaker hits (top 2–5%) seem to block cell cycle progression via a G1 arrest. This difference in phenotypic strength presumably reflects the more substantial cytological changes associated with mitosis and death, rather than differences in compound potency.
Comparison to metrics of chemical similarity
Compounds with similar structure have similar function17, and quantitative structure-activity relationships (SARs) are at the heart of drug discovery. As a step toward phenotype-based SARs, we investigated whether our phenotypic clustering groups together structurally similar compounds. For each compound we defined a circular molecular fingerprint using ECFP_4 descriptors that define molecular structure using radial atom neighborhoods (see Methods). We computed a similarity matrix based on Tanimoto similarities that describes the relationship between each of the 211 compounds in our hit set. Analogously, we generated a cosine distance-based phenotypic similarity matrix using our factor-based phenotypic profiles. These matrices, displayed as heat maps, are shown side by side (Fig. 4a). The compounds are ordered by phenotypic similarity using unsupervised clustering, so the seven primary phenotypes appear as blue boxes on the diagonal in the biological space panel.
(a) The relationship between phenotype (biological activity space) and compound structure (chemical space) was examined using similarity matrices. Phenotypic similarity between compounds is determined by the cosine distance between phenotypic vectors. Compound similarity is determined by the Tanimoto similarities in ECFP_4 fingerprints. Similarities are organized in 211 × 211 matrices ordered based on the clustering in Figure 3d. The corresponding dendrogram from Figure 3d is shown. A heat map is applied to phenotypic (black to blue) and compound structure (black to yellow) similarity matrices. Color bars are shown for each. The scale was selected so similarities at or below the 75th percentile are black, and those at the 99th percentile are maximally colored (blue or yellow). Black bars adjacent to the compound similarity matrix indicate positioning of the subclusters shown (Figs. 5 and 6). (b) The extent of structure activity concordance and discordance was assessed by comparing Tanimoto similarities and phenotypic distance between each pair of compounds. Analysis focused on comparisons where at least one compound was active. A 10% random sample of the entire similarity/distance dataset is plotted. Compound pairs with high structural similarity (Tanimoto score ≥ 0.3) and low phenotypic distance (Euclidean distance < 1) show structure-phenotype concordance. Compound pairs with high structural similarity and high phenotypic distance (Euclidean distance ≥ 1) show structure-phenotype discordance (“activity cliffs”). The red datum identifies the scoulerine-related compound pair shown in Supplementary Fig. 5.
In the chemical space side, we observed multiple blocks of structurally similar compounds that correspond to phenotypes 1, 2, 6 and 7. The blocks of chemical similarity were smaller than the phenotype blocks, because only a subset of compounds causing a given phenotype are similar, and in some cases, multiple blocks of chemical similarity were observed for a given phenotype, especially phenotype 6. These clusters evidently reflect regions in which biological effects are dominated by distinct structural compounds classes. The relationship between phenotype space and chemical space we observed (Fig. 4a) is perhaps expected, but to our knowledge it has not been visualized before in such quantitative detail.
To quantify the extent to which phenotypic clustering of the active compounds groups structurally related compounds, and to determine whether this structure-function concurrence is beyond what would be expected by random chance, we determined the Spearman correlation coefficient for rank-ordered phenotypic similarities and the corresponding compound similarities using the matrices from Figure 4a. We found an overall modest positive correlation (correlation = 0.0746), which presumably reflects strong correlation within small clusters, and lack of correlation elsewhere. We then generated 1,000 random compound similarity matrices by randomized sorting, computed the Spearman correlation coefficient with the phenotypic ordering, and used this to evaluate whether the observed correlation was statistically significant (Supplementary Fig. 4 online). This analysis indicates that the observed correlation is significant (P < 0.001) and approximately two-fold above that maximum chance observation. Thus, whereas this analysis comprises both structurally similar and structurally dissimilar compounds, the significance of the association between compound structure and function suggests that the molecular similarity principle17,18 holds for our phenotypic compound profiling.
In light of emerging evidence that the molecular similarity principle might not always hold true19, we sought to understand the extent to which small changes in structure are associated with large changes in function; for example, activity cliffs. To address this concept we compared Tanimoto similarities with phenotypic distance between each compound pair in our screening set. Because of the large number of comparisons we focused our analysis on only those comparisons in which at least one compound in a pair was active, and we examined phenotypic distance for those compound pairs that showed a Tanimoto structural similarity score ≥ 0.3; this threshold was selected based on the distribution of similarity scores in our dataset and is consistent with similarity scores observed between compounds showing activity against the same target20. Our analysis reveals that approximately 96% of the examined compounds with similar structure show substantially similar phenotypic readouts (Fig. 4b). Alternatively, of the structurally similar compounds active in our assays, 4% show significant phenotypic divergence (Fig. 4b). To understand this divergence further we examined a pair of scoulerine-related compounds more closely (Fig. 4b and Supplementary Fig. 5 online). These two compounds have high molecular similarity (top 0.1% based on similarity) and differ essentially by the presence of methoxy or hydroxyl groups (Supplementary Fig. 5). The compound pair has significantly different phenotypes (top 1% based on phenotypic distance), and this functional divergence is consistent with recent structure-activity studies on the two compounds21,22. Taken together we conclude that activity cliffs do emerge in our phenotypic screen. But, they represent the minority of cases. We are therefore more likely to observe phenotype concordance for structurally similar compounds.
Examples of phenotypic SARs
We chose several local SARs to examine in more detail (indicated by black bars adjacent to the compound structure similarity matrix in Fig. 4a). A subcluster that falls within phenotype 4 is shown in Figure 5a. This subcluster is characterized by decreased nuclear size, replication and EdU texture scores and increased nuclear morphology score. Unlike most of the compounds in cluster 4, this subcluster does not show a substantial increase in chromosome condensation. Thus, these compounds, though apparently cytotoxic, generate a phenotypic cytotoxicity signature that is distinct from that of classic apoptosis. This subcluster is enriched in antibiotic compounds that have known cytotoxic effects in mammalian cells. A small structural cluster contained three cyclic hexadepsipeptides, including aurantimycin (1) and diperamycin (2), which are derived from strains of Streptomyces23,24. These showed strong phenotypic similarity to a structurally divergent lysolipin derivative. The second region of local structural convergence within this phenotypic cluster contains several cyclic nonpeptide compounds. This includes kendomycin (3), an antibiotic with a C-glycosidic core and previously reported mammalian cytotoxicity and endothelin receptor antagonistic activity25, as well as other cyclic compounds that include two antibiotics of the concanamycin class that have cytotoxic activity and that are potent inhibitors of vacuolar ATPases26. We also note that this phenotypic subcluster contains a region with structurally distinct cytotoxic and antibiotic compounds, including heptelidic acid chlorohydrin (4)27. The phenotypic similarity of all these compounds presumably reflects a common target at the level of protein or pathway that may be vacuolar ATPases or other proteins that function in related areas of vesicular trafficking26.
Relationships between clusters containing similar phenotypic profiles (and corresponding structural similarities) and member compounds were examined. Three examples are shown with phenotypic profiles and subcluster dendrograms from Figure 3d. The heat map is maximally blue at or below a standardized factor score of −1.5 and maximally red at or above a standardized factor score of +1.5. The corresponding submatrices from the compound structure similarity matrix are also shown with the identical color map (Fig. 4). Yellow indicates high similarity, and black indicates low similarity. Structures are shown for several member compounds, and the position within the clusters is indicated by number. (a) Subcluster of compounds that result in a cell death phenotype characterized by low factor 1 (nuclear size) and increased factor 3 (chromosome condensation). The subcluster contains two cyclic depsipeptides with known cytotoxicity, 1 and 2, and cytotoxic antibiotics 3 and 4. (b) Subcluster of compounds resulting in G1 arrest characterized by large nuclear size (factor 1) and low DNA replication and mitosis (factors 2 and 3). This subcluster consists mainly of corticosteroids; for example, 5, 6 and 7. (c) A larger subcluster phenotypically dominated by low DNA replication, mitosis and EdU texture and average to high nuclear size. The top portion contains cardiac glycosides known to affect Na/K pumps, including 9, 10 and 11. The subcluster also contains protein translation inhibitors, 12 and 13. The lower portion contains steroid hormones, including 14 and 15.
A subcluster within phenotype 6 with high structural convergence is shown in Figure 5b. This subcluster is characterized by increased nuclear area and ellipticity, but decreased DNA replication, chromosome condensation, nuclear morphology and EdU texture. It contains 11 corticosteroid compounds with substantial structural similarity, including clobetasol-17-propionate (5), dexamethasone (6) and triamcinolone (7), and only one structurally dissimilar compound—entobex (8). Corticosteroids are known to cause a cell cycle arrest during G1 (ref. 28), which validates our interpretation of the parent cluster 6 as a G1 arrest phenotype. However, the local grouping of highly structurally similar compounds within this subcluster indicates a corticosteroid-specific G1 arrest phenotype. Such discrimination is surprising given our choice of cell types and fluorescent probes, and it indicates the power of relatively subtle morphology descriptors, such as nuclear shape metrics, to report on biological activity.
A larger phenotypic subcluster within phenotype 7, which has various effects on nuclear size, and a persistent decrease in DNA replication and EdU texture, was identified as consistent with a cell cycle arrest (Fig. 5c). Within this subcluster we found two groups of structurally similar compounds separated by a region of structurally distinct compounds. The two groups display a substantial degree of intergroup similarity, presumably because they share a steroid, or steroid-like, structure. The first group contains three cardiac glycosides: digitoxigenin (9), ouabain octahydrate (10) and digoxin (11). These are well-known inhibitors of Na/K pumps and have been shown to inhibit topoisomerase I in mammalian cells at nanomolar concentrations29. At high doses, cardiac glycosides cause a large drop in intracellular potassium levels, which leads to an inhibition of protein synthesis30,31. The protein translation inhibitor emetine (12)32 is also present within this cardiac glycoside subcluster, which suggests that the protein translation inhibition mechanism of action of these compounds may dominate their phenotypic effect in our assay. Supporting this interpretation, the non-structurally related translation inhibitor cycloheximide (13) shares this phenotype. The second group of related compounds contains a set of steroid hormones including progesterone (14) and danatrol (15). Progesterone signaling is known to result in growth arrest in G0/G1 (refs. 33,34).
Integration with ligand-target knowledge space
Our observation that multiple distinct structural classes of compounds can produce similar phenotypes, even at our highest phenotypic resolution, could be explained by compounds perturbing common targets via the same or different binding sites, or by compounds perturbing different components of common pathways. We investigated this possibility by implementing a structure-based target prediction method that has recently been reported35. Statistical models of substructural features were combined with an annotated chemogenomics database (WOMBAT) that associates ligand molecular structures with their cognate biological targets. We used these “known” ligand-target associations to train a naive Bayes model that we then used to predict the targets of our 211 active compounds. Using the top five most probable targets for each compound, we examined the extent to which phenotypic clustering of all the active compounds groups their cognate predicted targets. Notably, we found an increased positive correlation (correlation = 0.136, P < 0.001, Supplementary Fig. 6 online) between phenotypes and targets. This is twice the strength in correlation compared with the phenotype-to-structure comparison, which indicates that the observed divergence in SARs can in part be accounted for by structurally different compounds having common targets.
Although our results above point to the effectiveness of the target prediction method, predicting ligand-target association is an imperfect art. Thus, comparisons with the more robust phenotype and chemical similarity measures must be treated with caution. To provide a sense of its potential utility in pointing to a particular target, we illustrate results from a subcluster from mitotic arrest phenotype 1, which is primarily characterized by high chromosome condensation. Within this cluster we observed four distinct groups of structurally related compounds. The first, second, third and fourth groups are characterized by a colchicine derivative, a set of novel kinase inhibitors, a quinoline derivative and a pseudolarix acid B derivative. Our substructure-based method predicted multiple targets for each compound. We focused only on the top five targets, and for visualization purposes plotted only those targets that are predicted at least twice within the phenotypic subcluster (Fig. 6a). We find that a majority of all the compounds are predicted to target tubulin (7 out of 13), and as a consequence should affect mitotic spindle integrity. Additionally, the distinct group of novel kinase inhibitor compounds is predicted to hit both cyclin-dependent kinase 1 (CDK1) and CDK2. Colchicine is a well-known inhibitor of microtubule dynamics; it binds a distinct pocket within tubulin and causes depolymerization36. The colchicine derivative we found should have similar effects in cells. Several quinoline derivatives, including the one we found37, have been shown to also depolymerize microtubules via tubulin interactions38, and pseudolarix B has been recently shown to affect tubulin polymerization through a binding site distinct from the colchicine pocket39.
(a) A mitotic subcluster. Factor-based phenotypic profiles and subcluster dendrograms from Figure 3d are shown (−1.5 = blue, +1.5 = red). The corresponding compound structure similarity submatrix is also shown with the identical color map from Figure 4 (black, low similarity; yellow, high similarity). Structures are shown for several member compounds, and the position within the clusters is indicated by number. We predicted targets for each compound as described in the Methods. Blue boxes identify related predicted target with corresponding compound. Only genes encoding proteins that are targeted by two or more compounds within the cluster are shown. (b) The structures of three representative compounds are shown—16, 17 and 18. Also shown are images of cells treated with compound for 20 h and stained for DNA by Hoescht dye and the predicted target α-tubulin. Cell cycle profiles determined from HCS images using a decision tree–based classification scheme described elsewhere (C. Tao, Novartis Institutes for BioMedical Research, personal communication) are shown for each compound. Images and profiles of control cells with normal phenotype are shown. 20-μm size bars are shown in the control micrographs and correspond to all micrographs.
To gain mechanistic insight, we examined cytoskeletal morphology and cell cycle profiles for the set of putative tubulin-targeting compounds. We used immunofluorescence microscopy to detect α-tubulin in cells treated with each compound at the screening dose. As predicted, we observed depolymerization of microtubules and mitotic arrest in cells treated with each of the colchicine (16), quinoline (17) and pseudolarix acid B (18) derivatives (Fig. 6b). Thus, integration of compound structure with knowledge-based ligand-target predictions reveals that similar phenotypes produced by different compounds can in part be accounted for by targeting different components of common pathways, and by compounds hitting common targets via different binding sites. Moreover, our results indicate that phenotype and predicted targets constitute a useful SAR pair that can overcome the limitations of chemical similarities.
Discussion
The central goal of our study was to investigate SARs by integrating phenotypic information from HCS with chemical knowledge from profiles of chemical similarity and predicted targets. Such integration would be a powerful tool in drug discovery. This is not a novel concept, but it has been difficult to achieve at a practical level, in part because we lack conceptual frameworks for integrating high-dimensional biological and chemical data, and in part because high-dimensional datasets of biological activity (for example, microarray data) are typically too expensive to acquire across a large number of compounds. Our results represent considerable progress on the integrated structure-activity problem, using easy-to-adopt methods. The two chemical knowledge profiles we use, structural similarity (Figs. 4 and 5) and target predictions (Fig. 6), differ considerably in their rigor and degree of development, the former being a well-established science and the latter more of a ongoing challenge for computational chemists than a practical reality. Thus, our goals in comparing them to phenotypic profiles were rather different in the two cases. In the case of structural similarity, we knew that clusters of compounds that were related by phenotype and chemistry should exist in our library, and we used the comparison with phenotypes to find them and to examine them in detail to uncover new mechanistic information (Fig. 5). In the case of target prediction, we used the phenotype data to test how well the prediction algorithm was working, and also to point to one particular target (Fig. 6). Our analysis revealed that phenotypes correlate better with predicted compound targets than with the compound structures themselves (Supplementary Fig. 6). This result provided support for the effectiveness of the target prediction model and for the idea that different ligand-target interactions account, in part, for divergence in compound SARs.
Concordance between phenotypic and structural similarity profiles revealed the capability of HCS combined with factor analysis to make subtle phenotypic distinctions. For example, we readily discriminated the effects of corticosteroid-like and progesterone-like steroids, even though both cause cells to stop proliferating in G0/G1 (Fig. 5b,c). The subclustering of cytotoxic compounds (Fig. 5a) illustrates even finer phenotypic resolution. Obtaining this degree of distinction of therapeutically relevant mechanisms using a generic cancer cell line and cell cycle probes is notable, and it attests to the large amount of information that can be derived from microscope images when appropriate mining methods are implemented. Use of primary cells and more disease-relevant probes should further increase the resolution in areas relevant to drug discovery.
Lack of concordance between phenotypic and chemical similarity profiles is illustrated in the cytotoxicity cluster 4. One can envision cell death as a phenotypic end-point for multiple stress pathways that can be invoked by a variety of pharmacologic perturbations. In this regard we observe multiple distinct compound classes appearing within the cytotoxicity cluster, and consequently minimal correlation between structure and phenotype when examined with a low phenotypic resolution—that is, the cluster as whole. However, when examined at higher phenotypic resolution we can discriminate multiple small groups of structurally related compounds, within which we observed highly similar cytotoxicity signatures, for example the cyclic hexidepsipeptides versus the cyclic nonpeptide antibiotic compounds (Fig. 5a). This indicates that even at the end-point phenotype of cell death observed at a saturating dose we can still generate meaningful structure-function relationships.
Computational ligand-target prediction enabled us to demonstrate that by mapping compound structures to targets we improve our ability to discover meaningful SARs based on cytological phenotype (Supplementary Fig. 3). Furthermore, our data provide quantitative support for what is perhaps a logical explanation for divergence in structure versus phenotype concordance. To test the effectiveness of the target prediction method at higher phenotypic resolution, we looked closely at the predicted targets for four groups of phenotypically similar yet structurally distinct compounds. Our computational prediction pointed to tubulin as a common target for three of these groups, and our phenotypic data and follow-up experimental work supported this prediction (Fig. 6). Ligand-target prediction also revealed multiple highly probable targets that appear within each of four structural groups. Thus, parallel activity on these additional targets could account for subtle phenotypic differences between groups. Taken together, our results show that the combination of cytological phenotypes can improve confidence levels in target prediction both globally, as in our active compound set (Supplementary Fig. 2), and with respect to specific targets (Fig. 6). Thus quantitative cytological phenotypes, such as those derived here, may represent a new set of compound descriptor data that could be included directly into computational models to bolster compound-target prediction efficiency.
Despite progress on analysis of HCS data reported here and elsewhere40,41, the use of cytology to profile phenotypes in a broad and quantitative manner is still in its infancy. We believe this method has considerable potential. For example, new markers could be implemented that enable predictive toxicology of active lead compounds. Combined with chemical structure knowledge and ligand-target prediction, as shown here, such approaches could provide detailed mechanistic insight to help guide medicinal chemists early in the lead optimization process. Dealing with complexities of predictive toxicology will require breakthroughs in cytological image analysis, target prediction schemes and data mining. Our integration of image-based cytological phenotypes with chemical structure and computational ligand-target prediction represents a step forward in solving this and other difficult drug discovery problems.
Methods
Compound library.
We screened and profiled a library of 6,547 compounds derived from a diversity library (21%), a library of known bioactive compounds (21%) and a natural products library (58%). The bioactive set comprises those Novartis compounds that were recommended for promotion into development as drug candidates. This library has been compiled from multiple internal sources and includes entries irrespective of whether the compounds succeeded in preclinical or clinical development, or were introduced into the market. The natural products library consists of ∼3,800 compounds purified from plant extracts and other natural sources. In all cases, compounds were stored lyophilized and were determined by LC/MS to be at least 90% pure. Lyophilized compounds were resuspended in DMSO for a stock concentration of 10 mM. Immediately before, treatment samples of compound stock solution were diluted in DMEM to a 6× working concentration of 60 μM. We provide a table outlining all nonproprietary hit compound structures and available PubChem IDs (Supplementary Data 2).
Compound transfer.
HeLa cells (American Type Culture Collection) were plated in 384-well, black clear-bottomed plates (Greiner) at a density of 2,000 cells per well in 25 μl of growth medium (DMEM, 10% fetal bovine serum, penicillin and streptomycin; Invitrogen) for overnight incubation. Compounds were diluted in DMEM, and 5 μl of diluted compound was transferred to the 384-well culture plates at a final concentration of 10 μM per well using the BioMek FX (Beckman Coulter). Plates were transferred to 37 °C and incubated for 20 h.
Cell staining.
After 20 h of incubation with compound, cells were pulsed with 500 nM 5-ethynl-2′-deoxyuridine (Berry & Associates, Inc.) using a MultiDrop (Thermo Lab Systems) and incubated for 40 min at 37 °C. Cells were fixed in 3.7% paraformaldehyde for 30 min at 25 °C. Cells were washed once with phosphate-buffered saline (PBS; Invitrogen) with 0.5% Triton X-100 (PBST; Sigma Aldrich) using a Biotek Plate washer (Biotek Instruments) and then stained with rhodamine-azide (see Supplementary Methods). Plates were washed again with PBST and then incubated with primary antibodies. Rabbit anti-phosphohistone H3 Ser10 (Upstate) and mouse anti-α-tubulin (Sigma Aldrich) were added and plates were incubated at 25 °C for 3 h. Cells were washed once with PBST. Secondary antibodies donkey anti-mouse Alexa-488 (Invitrogen) and goat anti-rabbit Cy5 (Amersham) were added for 2 h at 25 °C. Cells were washed once with PBST and stained with Hoescht 33342 (Invitrogen) for 30 min at 25 °C. Wells were washed once with PBST, filled with PBS and sealed for imaging.
Imaging and image analysis.
Plates were imaged with a Cellomics Arrayscan. Images were collected using the XF93 filter set and 10× PlanFluor objective with camera binning set at 2 × 2. Individual cell segmentation was done using the Cellomics Morphology Explorer algorithm. Measurements for each cell were made on DNA intensity, nuclear area, deoxyuridine incorporation and phospho-H3 staining.
Analysis of image-based cytological phenotypes.
A detailed description of factor analysis and the phenotypic distance metric can be found in the Supplementary Methods.
Target prediction model.
Target prediction was performed using statistical models of substructural features, based on an annotated chemogenomics database that pairs ligand molecular structures and the biological targets they act on. The underlying assumption made is the “molecular similarity principle,” which assumes that similar molecules are likely to show similar properties17. We used the WOMBAT database42 in version 2006.1 as a knowledge base for training, which associates 154,236 ligands with 1,336 protein targets in 256,039 data entries. ECFP_4 fingerprints were calculated for washed and normalized structures, and multiple category naive Bayes models with Laplacian correction were trained on all data points, as implemented in PipelinePilot 5.1 (Scitegic). The five targets with the highest Bayes scores were considered for further analysis. For further details on the target prediction used see the original publication35 as well as a recent review that gives an overview of currently available methods and also highlights some recent applications43. The method used in this work is based on ECFP_4 descriptors, which are circular fingerprints encoding molecules as a set of radial patches that in their completeness again characterize the whole molecule. Circular fingerprints in general have been found to contain significant information regarding bioactivity44,45,46, but it was recently shown that three-dimensional descriptors show better generalization performance in case no bioactive structures similar to the one under consideration are known47. Though overall a high prediction performance of the correct target for >70% of the structures could be achieved in a validation study, the dependence of the method on the available knowledge base (training set) must be kept in mind. This is particularly true for new chemotypes.
Note: Supplementary information and chemical compound information is available on the Nature Chemical Biology website.
References
Haggarty, S.J. The principle of complementarity: chemical versus biological space. Curr. Opin. Chem. Biol. 9, 296–303 (2005).
Nichols, A. High content screening as a screening tool in drug discovery. Methods Mol. Biol. 356, 379–387 (2007).
Lang, P., Yeow, K., Nichols, A. & Scheer, A. Cellular imaging in drug discovery. Nat. Rev. Drug Discov. 5, 343–356 (2006).
Mitchison, T.J. Small-molecule screening and profiling by using automated microscopy. ChemBioChem 6, 33–39 (2005).
Blake, R.A. Target validation in drug discovery. Methods Mol. Biol. 356, 367–377 (2007).
Eggert, U.S. & Mitchison, T.J. Small molecule screening by imaging. Curr. Opin. Chem. Biol. 10, 232–237 (2006).
Carpenter, A. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Giuliano, K.A., Haskins, J.R. & Taylor, D.L. Advances in high content screening for drug discovery. Assay Drug Dev. Technol. 1, 565–577 (2003).
Lee, S. & Howell, B.J. High-content screening: emerging hardware and software technologies. Methods Enzymol. 414, 468–483 (2006).
Spearman, C. “General intelligence”, objectively determined and measured. Am. J. Psychol. 15, 201–293 (1904).
Carroll, J.B. & Schweiker, R.F. Factor analysis in educational research. Rev. Educ. Res. 21, 368–388 (1951).
Floyd, F.J. & Widaman, K.F. Factor analysis in the development and refinement of clinical assessment instruments. Psychol. Assess. 7, 286–299 (1995).
Malinowski, E.R. Factor Analysis in Chemistry 1–432 (John Wiley and Sons, Inc., New York, 2002).
Stewart, D.W. The application and misapplication of factor analysis in marketing research. J. Mark. Res. 18, 51–62 (1981).
Tinsley, H.E.A. & Tinsley, D.J. Uses of factor analysis in counseling psychology research. J. Couns. Psychol. 34, 414–424 (1987).
Johnson, R.A. & Wichern, D.W. Applied Multivariate Statistical Analysis 426–542 (Prentice Hall, Inc., Upper Saddle River, New Jersey, USA, 2002).
Bender, A. & Glen, R.C. Molecular similarity: a key technique in molecular informatics. Org. Biomol. Chem. 2, 3204–3218 (2004).
Johnson, M., Lajiness, M. & Maggiora, G. Molecular similarity: a basis for designing drug screening programs. Prog. Clin. Biol. Res. 291, 167–171 (1989).
Maggiora, G.M. On outliers and activity cliffs–why QSAR often disappoints. J. Chem. Inf. Model. 46, 1535 (2006).
Hert, J. et al. New methods for ligand-based virtual screening: use of data fusion and machine learning to enhance the effectiveness of similarity searching. J. Chem. Inf. Model. 46, 462–470 (2006).
Carrieri, A., Centeno, N.B., Rodrigo, J., Sanz, F. & Carotti, A. Theoretical evidence of a salt bridge disruption as the initiating process for the alpha1d-adrenergic receptor activation: a molecular dynamics and docking study. Proteins 43, 382–394 (2001).
Schaper, K. Free-Wilson-type analysis of non-additive substituent effects on THPB dopamine receptor affinity using artificial neural networks. Quant. Struct. Act. Relat. 18, 354–360 (1999).
Gräfe, U. et al. Aurantimycins, new depsipeptide antibiotics from Streptomyces aurantiacus IMET 43917. Production, isolation, structure elucidation, and biological activity. J. Antibiot. (Tokyo) 48, 119–125 (1995).
Matsumoto, N. et al. Diperamycin, a new antimicrobial antibiotic produced by Streptomyces griseoaurantiacus MK393–AF2. I. Taxonomy, fermentation, isolation, physico-chemical properties and biological activities. J. Antibiot. (Tokyo) 51, 1087–1092 (1998).
Yuan, Y., Men, H. & Lee, C. Total synthesis of kendomycin: a macro-C-glycosidation approach. J. Am. Chem. Soc. 126, 14720–14721 (2004).
Manabe, T., Yoshimori, T., Henomatsu, N. & Tashiro, Y. Inhibitors of vacuolar-type H(+)-ATPase suppresses proliferation of cultured cells. J. Cell. Physiol. 157, 445–452 (1993).
Kawashima, J. et al. Antitumor activity of heptelidic acid chlorohydrin. J. Antibiot. (Tokyo) 47, 1562–1563 (1994).
Samuelsson, M.K., Pazirandeh, A., Davani, B. & Okret, S. p57Kip2, a glucocorticoid-induced inhibitor of cell cycle progression in HeLa cells. Mol. Endocrinol. 12, 1811–1822 (1999).
Bielawski, K., Winnicka, K. & Bielawska, A. Inhibition of DNA topoisomerases I and II, and growth inhibition of breast cancer MCF-7 cells by ouabain, digoxin and proscillaridin A. Biol. Pharm. Bull. 29, 1493–1497 (2006).
Ramirez-Ortega, M. et al. Proliferation and apoptosis of HeLa cells induced by in vitro stimulation with digitalis. Eur. J. Pharmacol. 534, 71–76 (2006).
Pauw, P.G., Kaffer, C.R., Petersen, R.J., Semerad, S.A. & Williams, D.C. Inhibition of myogenesis by ouabain: effect on protein synthesis. In Vitro Cell. Dev. Biol. Anim. 36, 133–138 (2000).
Schweighoffer, T. et al. Cytometric analysis of DNA replication inhibited by emetine and cyclosporin A. Histochemistry 96, 93–97 (1991).
Horiuchi, S. et al. Expression of progesterone receptor B is associated with G0/G1 arrest of the cell cycle and growth inhibition in NIH3T3 cells. Exp. Cell Res. 305, 233–243 (2005).
Owen, G.I., Richer, J.K., Tung, L., Takimoto, G. & Horwitz, K. Progesterone regulates transcription of the p21(WAF1) cyclin-dependent kinase inhibitor gene through Sp1 and CBP/p300. J. Biol. Chem. 273, 10696–10701 (1998).
Nidhi, G., Glick, M., Davies, J.W. & Jenkins, J.L. Prediction of biological targets for compounds using multiple-category Bayesian models trained on chemogenomics databases. J. Chem. Inf. Model. 46, 1124–1133 (2006).
Checchi, P.M., Nettles, J.H., Zhou, J., Snyder, J.P. & Joshi, H.C. Microtubule-interacting drugs for cancer treatment. Trends Pharmacol. Sci. 24, 361–365 (2003).
Li, L. et al. Antitumor agents 155. Synthesis and biological evaluation of 3′,6,7-substituted 2-phenyl-4-quinolones as antimicrotubule agents. J. Med. Chem. 37, 3400–3407 (1994).
Shi, Q., Chen, K., Morris-Natschke, S.L. & Lee, K.H. Recent progress in the development of tubulin inhibitors as antimitotic antitumor agents. Curr. Pharm. Des. 4, 219–248 (1998).
Tong, Y.G. et al. Pseudolarix acid B, a new tubulin-binding agent, inhibits angiogenesis by interacting with a novel binding site on tubulin. Mol. Pharmacol. 69, 1226–1233 (2006).
Clemons, P.A. Complex phenotypic assays in high-throughput screening. Curr. Opin. Chem. Biol. 8, 334–338 (2004).
Loo, L.H., Wu, L.F. & Altschuler, S.J. Image-based multivariate profiling of drug responses from single cells. Nat. Methods 4, 445–453 (2007).
Olah, M. et al. in Chemoinformatics in Drug Discovery WOMBAT: World of Molecular Bioactivity (ed. Oprea, T.I.) 223–239 (Wiley-VCH, New York, 2004).
Jenkins, J.L., Bender, A. & Davies, J.W. In silico target fishing: predicting biological targets from chemical structure. Drug Discov. Today Technol. 3, 413–421 (2007).
Bender, A. & Glen, R.C. A discussion of measures of enrichment in virtual screening: comparing the information content of descriptors with increasing levels of sophistication. J. Chem. Inf. Model. 45, 1369–1375 (2005).
Bender, A., Mussa, H.Y., Reiling, S. & Glen, R.C. Similarity searching of chemical databases using atom environment descriptors (MOLPRINT 2D): evaluation of performance. J. Chem. Inf. Model. 44, 1708–1718 (2004).
Glen, R.C. et al. Circular fingerprints: flexible molecular descriptors with applications from physical chemistry to ADME. IDrugs 9, 199–204 (2006).
Nettles, J.H. et al. Bridging chemical and biological space: “target fishing” using 2D and 3D molecular descriptors. J. Med. Chem. 49, 6802–6810 (2006).
Acknowledgements
We thank L. Martell, M. Thoma, J. Nettles, B. Dwyer and M. Pflumm for insightful comments and discussions, G. Paris for assembly of the climax screening collection, A. Salic (Harvard Medical School) for the gift of rhodamine azide, C. Mickanin and S. Zhao for automation support, and Q. Yang for database support. D.W.Y. and A.B. are both Novartis Presidential Postdoctoral Fellows. Work in the T.J.M. lab is supported by US National Institutes of Health grant CA78048.
Author information
Authors and Affiliations
Contributions
D.W.Y., A.B., J.L.J., T.J.M. and Y.F. conceived the work. J.H., D.W.Y. and E.M. performed experiments. D.W.Y. developed and implemented factor analysis of HCS data. A.B. performed ligand-target and compound structure analysis. D.W.Y. and A.B. performed integrated statistical analysis of biological and chemical data. D.W.Y., T.J.M. and Y.F. analyzed phenotypes. C.Y.T. performed cell cycle analysis. J.A.T. and M.L. contributed to experimental design and interpretation. G.-W.C. assisted in data processing and analysis. D.W.Y. and A.B. wrote the paper with assistance from J.L.J., T.J.M. and Y.F.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Table 1 and Supplementary Methods (PDF 1158 kb)
Supplementary Data 1
Workbook containing results from the factor analysis. (XLS 86 kb)
Supplementary Data 2
Table of hit data. (XLS 445 kb)
Rights and permissions
About this article
Cite this article
Young, D., Bender, A., Hoyt, J. et al. Integrating high-content screening and ligand-target prediction to identify mechanism of action. Nat Chem Biol 4, 59–68 (2008). https://doi.org/10.1038/nchembio.2007.53
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2007.53
This article is cited by
-
High-content analysis identified synergistic drug interactions between INK128, an mTOR inhibitor, and HDAC inhibitors in a non-small cell lung cancer cell line
BMC Cancer (2024)
-
Yearning for machine learning: applications for the classification and characterisation of senescence
Cell and Tissue Research (2023)
-
Assignment of unimodal probability distribution models for quantitative morphological phenotyping
BMC Biology (2022)
-
A statistical framework for high-content phenotypic profiling using cellular feature distributions
Communications Biology (2022)
-
Orthogonally-tunable and ER-targeting fluorophores detect avian influenza virus early infection
Nature Communications (2022)