Abstract
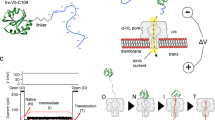
Membrane proteins are designed to fold and function in a lipid membrane, yet folding experiments within a native membrane environment are challenging to design. Here we show that single-molecule forced unfolding experiments can be adapted to study helical membrane protein folding under native-like bicelle conditions. Applying force using magnetic tweezers, we find that a transmembrane helix protein, Escherichia coli rhomboid protease GlpG, unfolds in a highly cooperative manner, largely unraveling as one physical unit in response to mechanical tension above 25 pN. Considerable hysteresis is observed, with refolding occurring only at forces below 5 pN. Characterizing the energy landscape reveals only modest thermodynamic stability (ΔG = 6.5 kBT) but a large unfolding barrier (21.3 kBT) that can maintain the protein in a folded state for long periods of time (t1/2 ∼3.5 h). The observed energy landscape may have evolved to limit the existence of troublesome partially unfolded states and impart rigidity to the structure.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Engelman, D.M. et al. Membrane protein folding: beyond the two stage model. FEBS Lett. 555, 122–125 (2003).
Bowie, J.U. Solving the membrane protein folding problem. Nature 438, 581–589 (2005).
White, S.H. & von Heijne, G. How translocons select transmembrane helices. Annu. Rev. Biophys. 37, 23–42 (2008).
Hong, H., Blois, T.M., Cao, Z. & Bowie, J.U. Method to measure strong protein-protein interactions in lipid bilayers using a steric trap. Proc. Natl. Acad. Sci. USA 107, 19802–19807 (2010).
Chang, Y.C. & Bowie, J.U. Measuring membrane protein stability under native conditions. Proc. Natl. Acad. Sci. USA 111, 219–224 (2014).
Oesterhelt, F. et al. Unfolding pathways of individual bacteriorhodopsins. Science 288, 143–146 (2000).
Kedrov, A., Janovjak, H., Sapra, K.T. & Muller, D.J. Deciphering molecular interactions of native membrane proteins by single-molecule force spectroscopy. Annu. Rev. Biophys. Biomol. Struct. 36, 233–260 (2007).
Engel, A. & Gaub, H.E. Structure and mechanics of membrane proteins. Annu. Rev. Biochem. 77, 127–148 (2008).
Zocher, M. et al. Single-molecule force spectroscopy from nanodiscs: an assay to quantify folding, stability, and interactions of native membrane proteins. ACS Nano 6, 961–971 (2012).
Cecconi, C., Shank, E.A., Bustamante, C. & Marqusee, S. Direct observation of the three-state folding of a single protein molecule. Science 309, 2057–2060 (2005).
Shank, E.A., Cecconi, C., Dill, J.W., Marqusee, S. & Bustamante, C. The folding cooperativity of a protein is controlled by its chain topology. Nature 465, 637–640 (2010).
Faham, S. & Bowie, J.U. Bicelle crystallization: a new method for crystallizing membrane proteins yields a monomeric bacteriorhodopsin structure. J. Mol. Biol. 316, 1–6 (2002).
Joh, N.H. et al. Modest stabilization by most hydrogen-bonded side-chain interactions in membrane proteins. Nature 453, 1266–1270 (2008).
Dürr, U.H., Gildenberg, M. & Ramamoorthy, A. The magic of bicelles lights up membrane protein structure. Chem. Rev. 112, 6054–6074 (2012).
Wang, Y.C., Zhang, Y.J. & Ha, Y. Crystal structure of a rhomboid family intramembrane protease. Nature 444, 179–183 (2006).
Wu, Z.R. et al. Structural analysis of a rhomboid family intramembrane protease reveals a gating mechanism for substrate entry. Nat. Struct. Mol. Biol. 13, 1084–1091 (2006).
Ben-Shem, A., Fass, D. & Bibi, E. Structural basis for intramembrane proteolysis by rhomboid serine proteases. Proc. Natl. Acad. Sci. USA 104, 462–466 (2007).
Vinothkumar, K.R. Structure of rhomboid protease in a lipid environment. J. Mol. Biol. 407, 232–247 (2011).
Lemberg, M.K. & Freeman, M. Cutting proteins within lipid bilayers: rhomboid structure and mechanism. Mol. Cell 28, 930–940 (2007).
Freeman, M. Rhomboid proteases and their biological functions. Annu. Rev. Genet. 42, 191–210 (2008).
Ha, Y., Akiyama, Y. & Xue, Y. Structure and mechanism of rhomboid protease. J. Biol. Chem. 288, 15430–15436 (2013).
Vinothkumar, K.R. & Freeman, M. Intramembrane proteolysis by rhomboids: catalytic mechanisms and regulatory principles. Curr. Opin. Struct. Biol. 23, 851–858 (2013).
Lemberg, M.K. Sampling the membrane: function of rhomboid-family proteins. Trends Cell Biol. 23, 210–217 (2013).
Baker, R.P. & Urban, S. Architectural and thermodynamic principles underlying intramembrane protease function. Nat. Chem. Biol. 8, 759–768 (2012).
Paslawski, W. et al. Cooperative folding of a polytopic α-helical membrane protein involves a compact N-terminal nucleus and nonnative loops. Proc. Natl. Acad. Sci. USA 112, 7978–7983 (2015).
Cecconi, C., Shank, E.A., Marqusee, S. & Bustamante, C. in DNA Nanotechnology: Methods and Protocols 1st edn. Vol. 749 (eds. Zuccheri, G. & Samorì, B.) 255–271 (Humana Press, 2011).
Min, D. et al. Mechanical unzipping and rezipping of a single SNARE complex reveals hysteresis as a force-generating mechanism. Nat. Commun. 4, 1705–1714 (2013).
Bae, W. et al. Programmed folding of DNA origami structures through single-molecule force control. Nat. Commun. 5, 5654–5661 (2014).
Gosse, C. & Croquette, V. Magnetic tweezers: micromanipulation and force measurement at the molecular level. Biophys. J. 82, 3314–3329 (2002).
Saleh, O.A., Allemand, J.F., Croquette, V. & Bensimon, D. Single-molecule manipulation measurements of DNA transport proteins. ChemPhysChem 6, 813–818 (2005).
Kim, K. & Saleh, O.A. A high-resolution magnetic tweezer for single-molecule measurements. Nucleic Acids Res. 37, 136–142 (2009).
Lipfert, J., Hao, X.M. & Dekker, N.H. Quantitative modeling and optimization of magnetic tweezers. Biophys. J. 96, 5040–5049 (2009).
De Vlaminck, I. & Dekker, C. Recent advances in magnetic tweezers. Annu. Rev. Biophys. 41, 453–472 (2012).
Greenleaf, W.J., Frieda, K.L., Foster, D.A.N., Woodside, M.T. & Block, S.M. Direct observation of hierarchical folding in single riboswitch aptamers. Science 319, 630–633 (2008).
Alegre-Cebollada, J., Kosuri, P., Rivas-Pardo, J.A. & Fernandez, J.M. Direct observation of disulfide isomerization in a single protein. Nat. Chem. 3, 882–887 (2011).
Stigler, J., Ziegler, F., Gieseke, A., Gebhardt, J.C.M. & Rief, M. The complex folding network of single calmodulin molecules. Science 334, 512–516 (2011).
Carrion-Vazquez, M. et al. Mechanical and chemical unfolding of a single protein: A comparison. Proc. Natl. Acad. Sci. USA 96, 3694–3699 (1999).
Liphardt, J., Onoa, B., Smith, S.B., Tinoco, I. Jr. & Bustamante, C. Reversible unfolding of single RNA molecules by mechanical force. Science 292, 733–737 (2001).
Beaugrand, M. et al. Lipid concentration and molar ratio boundaries for the use of isotropic bicelles. Langmuir 30, 6162–6170 (2014).
Curnow, P. & Booth, P.J. Combined kinetic and thermodynamic analysis of α-helical membrane protein unfolding. Proc. Natl. Acad. Sci. USA 104, 18970–18975 (2007).
Otzen, D.E. Mapping the folding pathway of the transmembrane protein DsbB by protein engineering. Protein Eng. Des. Sel. 24, 139–149 (2011).
Watters, A.L. et al. The highly cooperative folding of small naturally occurring proteins is likely the result of natural selection. Cell 128, 613–624 (2007).
Cymer, F., von Heijne, G. & White, S.H. Mechanisms of integral membrane protein insertion and folding. J. Mol. Biol. 427, 999–1022 (2015).
Kim, S.J. & Skach, W.R. Mechanisms of CFTR folding at the endoplasmic reticulum. Front. Pharmacol. 3, 201 (2012).
Curnow, P. et al. Stable folding core in the folding transition state of an α-helical integral membrane protein. Proc. Natl. Acad. Sci. USA 108, 14133–14138 (2011).
Jefferson, R.E., Blois, T.M. & Bowie, J.U. Membrane proteins can have high kinetic stability. J. Am. Chem. Soc. 135, 15183–15190 (2013).
Otzen, D.E. Folding of DsbB in mixed micelles: a kinetic analysis of the stability of a bacterial membrane protein. J. Mol. Biol. 330, 641–649 (2003).
Kim, B.L., Schafer, N.P. & Wolynes, P.G. Predictive energy landscapes for folding α-helical transmembrane proteins. Proc. Natl. Acad. Sci. USA 111, 11031–11036 (2014).
Urban, S. & Wolfe, M.S. Reconstitution of intramembrane proteolysis in vitro reveals that pure rhomboid is sufficient for catalysis and specificity. Proc. Natl. Acad. Sci. USA 102, 1883–1888 (2005).
Faham, S., Ujwal, R., Abramson, J. & Bowie, J.U. in Membrane Protein Crystallization 1st edn. Vol. 63 (eds. DeLucas, L.J.) 109–125 (Academic Press, 2009).
Ujwal, R. & Bowie, J.U. Crystallizing membrane proteins using lipidic bicelles. Methods 55, 337–341 (2011).
Whitelegge, J.P. et al. Toward the bilayer proteome, electrospray ionization-mass spectrometry of large, intact transmembrane proteins. Proc. Natl. Acad. Sci. USA 96, 10695–10698 (1999).
Kessler, D.A. & Rabin, Y. Distribution functions for filaments under tension. J. Chem. Phys. 121, 1155–1164 (2004).
Schwaiger, I., Sattler, C., Hostetter, D.R. & Rief, M. The myosin coiled-coil is a truly elastic protein structure. Nat. Mater. 1, 232–235 (2002).
Schuler, B., Lipman, E.A. & Eaton, W.A. Probing the free-energy surface for protein folding with single-molecule fluorescence spectroscopy. Nature 419, 743–747 (2002).
Yang, W.Y. & Gruebele, M. Folding at the speed limit. Nature 423, 193–197 (2003).
Rhoades, E., Cohen, M., Schuler, B. & Haran, G. Two-state folding observed in individual protein molecules. J. Am. Chem. Soc. 126, 14686–14687 (2004).
Kubelka, J., Hofrichter, J. & Eaton, W.A. The protein folding 'speed limit'. Curr. Opin. Struct. Biol. 14, 76–88 (2004).
Chung, H.S., Louis, J.M. & Eaton, W.A. Experimental determination of upper bound for transition path times in protein folding from single-molecule photon-by-photon trajectories. Proc. Natl. Acad. Sci. USA 106, 11837–11844 (2009).
Gebhardt, J.C.M., Bornschlogla, T. & Rief, M. Full distance-resolved folding energy landscape of one single protein molecule. Proc. Natl. Acad. Sci. USA 107, 2013–2018 (2010).
Acknowledgements
This work was supported by the National Creative Research Initiative Program (Center for Single-Molecule Systems Biology to T.-Y.Y.) funded by the National Research Foundation of Korea and Marine Biotechnology Program (20150220 to T.-Y.Y.) funded by the Ministry of Oceans and Fisheries of Korea, and supported by US National Institutes of Health grant 2R01GM063919 to J.U.B.
Author information
Authors and Affiliations
Contributions
D.M., R.E.J., J.U.B. and T.-Y.Y. designed the experiments. R.E.J. expressed and purified proteins. D.M. prepared the DNA-protein hybrid sample and performed the magnetic tweezers experiments. All of the authors analyzed the data and contributed to writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–10 and Supplementary Tables 1–3. (PDF 2696 kb)
Rights and permissions
About this article
Cite this article
Min, D., Jefferson, R., Bowie, J. et al. Mapping the energy landscape for second-stage folding of a single membrane protein. Nat Chem Biol 11, 981–987 (2015). https://doi.org/10.1038/nchembio.1939
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1939