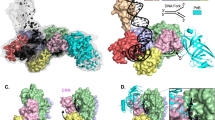
Abstract
The bidirectional replication of a circular chromosome by many bacteria necessitates proper termination to avoid the head-on collision of the opposing replisomes. In Escherichia coli, replisome progression beyond the termination site is prevented by Tus proteins bound to asymmetric Ter sites. Structural evidence indicates that strand separation on the blocking (nonpermissive) side of Tus–Ter triggers roadblock formation, but biochemical evidence also suggests roles for protein-protein interactions. Here DNA unzipping experiments demonstrate that nonpermissively oriented Tus–Ter forms a tight lock in the absence of replicative proteins, whereas permissively oriented Tus–Ter allows nearly unhindered strand separation. Quantifying the lock strength reveals the existence of several intermediate lock states that are impacted by mutations in the lock domain but not by mutations in the DNA-binding domain. Lock formation is highly specific and exceeds reported in vivo efficiencies. We postulate that protein-protein interactions may actually hinder, rather than promote, proper lock formation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Hill, T.M., Henson, J.M. & Kuempel, P.L. The terminus region of the E. coli chromosome contains two separate loci that exhibit polar inhibition of replication. Proc. Natl. Acad. Sci. USA 84, 1754–1758 (1987).
Hidaka, M., Kobayashi, T., Takenaka, S., Takeya, H. & Horiuchi, T. Purification of a DNA replication terminus (ter) site–binding protein in Escherichia coli and identification of the structural gene. J. Biol. Chem. 264, 21031–21037 (1989).
Hill, T.M. Arrest of bacterial DNA replication. Annu. Rev. Microbiol. 46, 603–633 (1992).
Kamada, K., Horiuchi, T., Ohsumi, K., Shimamoto, N. & Morikawa, K. Structure of a replication-terminator protein complexed with DNA. Nature 383, 598–603 (1996).
Khatri, G.S., MacAllister, T., Sista, P.R. & Bastia, D. The replication terminator protein of E. coli is a DNA sequence-specific contra-helicase. Cell 59, 667–674 (1989).
Hill, T.M. & Marians, K.J. Escherichia coli Tus protein acts to arrest the progression of DNA replication forks in vitro. Proc. Natl. Acad. Sci. USA 87, 2481–2485 (1990).
Hill, T.M. in Escherichia coli and Salmonella: Cellular and Molecular Biology 2nd edn. (eds. Neidhardt, F.C. et al.) 1602–1614 (American Society for Microbiology, Washington, DC, USA, 1996).
Coskun-Ari, F.F., Skokotas, A., Moe, G.R. & Hill, T.M. Biophysical characteristics of Tus, the replication arrest protein of Escherichia coli. J. Biol. Chem. 269, 4027–4034 (1994).
Neylon, C., Kralicek, A.V., Hill, T.M. & Dixon, N.E. Replication termination in Escherichia coli: structure and anti-helicase activity of the Tus-Ter complex. Microbiol. Mol. Biol. Rev. 69, 501–526 (2005).
Lee, J.Y., Finkelstein, I.J., Arciszewska, L.K., Sherratt, D.J. & Greene, E.C. Single-molecule imaging of FtsK translocation reveals mechanistic features of protein-protein collisions on DNA. Mol. Cell 54, 832–843 (2014).
Kaul, S. et al. The replication terminator protein of the gram-positive bacterium Bacillus subtilis functions as a polar contrahelicase in gram-negative Escherichia coli. Proc. Natl. Acad. Sci. USA 91, 11143–11147 (1994).
Andersen, P.A., Griffiths, A.A., Duggin, I.G. & Wake, R.G. Functional specificity of the replication fork-arrest complexes of Bacillus subtilis and Escherichia coli: significant specificity of Tus-Ter functioning in E. coli. Mol. Microbiol. 36, 1327–1335 (2000).
Sahoo, T., Mohanty, B.K., Lobert, M., Manna, A.C. & Bastia, D. The contrahelicase activities of the replication terminator proteins of Escherichia coli and Bacillus subtilis are helicase-specific and impede both helicase translocation and authentic DNA unwinding. J. Biol. Chem. 270, 29138–29144 (1995).
Mulugu, S. et al. Mechanism of termination of DNA replication of Escherichia coli involves helicase-contrahelicase interaction. Proc. Natl. Acad. Sci. USA 98, 9569–9574 (2001).
Mohanty, B.K., Sahoo, T. & Bastia, D. The relationship between sequence-specific termination of DNA replication and transcription. EMBO J. 15, 2530–2539 (1996).
Mohanty, B.K., Sahoo, T. & Bastia, D. Mechanistic studies on the impact of transcription on sequence-specific termination of DNA replication and vice versa. J. Biol. Chem. 273, 3051–3059 (1998).
Lee, E.H., Kornberg, A., Hidaka, M., Kobayashi, T. & Horiuchi, T. Escherichia coli replication termination protein impedes the action of helicases. Proc. Natl. Acad. Sci. USA 86, 9104–9108 (1989).
Lee, E.H. & Kornberg, A. Features of replication fork blockage by the Escherichia coli terminus-binding protein. J. Biol. Chem. 267, 8778–8784 (1992).
Bedrosian, C.L. & Bastia, D. Escherichia coli replication terminator protein impedes simian virus 40 (SV40) DNA replication fork movement and SV40 large tumor antigen helicase activity in vitro at a prokaryotic terminus sequence. Proc. Natl. Acad. Sci. USA 88, 2618–2622 (1991).
Hidaka, M. et al. Termination complex in Escherichia coli inhibits SV40 DNA replication in vitro by impeding the action of T antigen helicase. J. Biol. Chem. 267, 5361–5365 (1992).
Mulcair, M.D. et al. A molecular mousetrap determines polarity of termination of DNA replication in E. coli. Cell 125, 1309–1319 (2006).
Neylon, C. et al. Interaction of the Escherichia coli replication terminator protein (Tus) with DNA: a model derived from DNA-binding studies of mutant proteins by surface plasmon resonance. Biochemistry 39, 11989–11999 (2000).
Bastia, D. et al. Replication termination mechanism as revealed by Tus-mediated polar arrest of a sliding helicase. Proc. Natl. Acad. Sci. USA 105, 12831–12836 (2008).
Liphardt, J., Onoa, B., Smith, S.B., Tinoco, I. Jr. & Bustamante, C. Reversible unfolding of single RNA molecules by mechanical force. Science 292, 733–737 (2001).
Woodside, M.T. et al. Nanomechanical measurements of the sequence-dependent folding landscapes of nucleic acid hairpins. Proc. Natl. Acad. Sci. USA 103, 6190–6195 (2006).
Lionnet, T., Spiering, M.M., Benkovic, S.J., Bensimon, D. & Croquette, V. Real-time observation of bacteriophage T4 gp41 helicase reveals an unwinding mechanism. Proc. Natl. Acad. Sci. USA 104, 19790–19795 (2007).
Schwarz, G. Estimating dimension of a model. Ann. Statist. 6, 461–464 (1978).
Cowan, G. Statistical Data Analysis (Oxford University Press, Oxford, UK, 1998).
Press, W.H., Flannery, B.P., Teukolsky, S.A. & Vetterling, W.T. Numerical Recipes in C: The Art of Scientific Computing 2nd edn. (Cambridge University Press, Cambridge, UK, 1992).
Dulin, D. et al. Elongation-competent pauses govern the fidelity of a viral RNA-dependent RNA polymerase. Cell Rep. 10, 983–992 (2015).
Janissen, R. et al. Invincible DNA tethers: covalent DNA anchoring for enhanced temporal and force stability in magnetic tweezers experiments. Nucleic Acids Res. 42, e137 (2014).
Coskun-Ari, F.F. & Hill, T.M. Sequence-specific interactions in the Tus-Ter complex and the effect of base pair substitutions on arrest of DNA replication in Escherichia coli. J. Biol. Chem. 272, 26448–26456 (1997).
Dantus, M., Bowman, R.M. & Zewail, A.H. Femtosecond laser observations of molecular vibration and rotation. Nature 343, 737–739 (1990).
Arasaki, Y., Takatsuka, K., Wang, K. & McKoy, V. Energy- and angle-resolved pump-probe femtosecond photoelectron spectroscopy: molecular rotation. J. Chem. Phys. 114, 7941–7950 (2001).
Middleton, C.T. et al. DNA excited-state dynamics: from single bases to the double helix. Annu. Rev. Phys. Chem. 60, 217–239 (2009).
Cnossen, J.P., Dulin, D. & Dekker, N.H. An optimized software framework for real-time, high-throughput tracking of spherical beads. Rev. Sci. Instrum. 85, 103712 (2014).
te Velthuis, A.J., Kerssemakers, J.W., Lipfert, J. & Dekker, N.H. Quantitative guidelines for force calibration through spectral analysis of magnetic tweezers data. Biophys. J. 99, 1292–1302 (2010).
Yu, Z. et al. A force calibration standard for magnetic tweezers. Rev. Sci. Instrum. 85, 123114 (2014).
Acknowledgements
We thank J. Cnossen for his countless efforts in adjusting the MT software to our needs, J. Kerssemakers for discussions and S. Hamdan for his contribution to the discussion by generously sharing his own experimental observations. This study was supported by a grant from the Australian Research Council (DP0877658) to N.E.D. and by Vici and TOP grants from the Netherlands Organisation for Scientific Research (NWO) and ERC Starting Grant DynGenome from the European Research Council to N.H.D.
Author information
Authors and Affiliations
Contributions
N.E.D. and N.H.D. designed the research. B.A.B. and N.H.D. designed the experiments. D.D. designed and assembled the magnetic tweezers apparatus. B.A.B. performed the experiments. B.A.B., B.C., T.v.L., R.J. and N.H.D. designed, and B.C. and T.v.L. made, the DNA hairpin constructs. Z.-Q.X. and S.J. purified the Tus proteins. B.A.B. and M.D. analyzed the data. M.D. developed the application of MLE to force spectroscopy data. B.A.B., D.D., M.D. and N.H.D. interpreted the data. S.J. and N.E.D. contributed to discussions concerning the model and in vivo observations. B.A.B., N.E.D. and N.H.D. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–4 and Supplementary Tables 1 and 2. (PDF 1507 kb)
Rights and permissions
About this article
Cite this article
Berghuis, B., Dulin, D., Xu, ZQ. et al. Strand separation establishes a sustained lock at the Tus–Ter replication fork barrier. Nat Chem Biol 11, 579–585 (2015). https://doi.org/10.1038/nchembio.1857
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1857
This article is cited by
-
Single molecule high-throughput footprinting of small and large DNA ligands
Nature Communications (2017)