Abstract
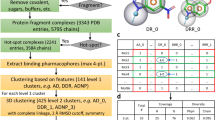
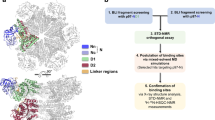
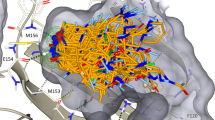
Chemical probes that form a covalent bond with a protein target often show enhanced selectivity, potency and utility for biological studies. Despite these advantages, protein-reactive compounds are usually avoided in high-throughput screening campaigns. Here we describe a general method (DOCKovalent) for screening large virtual libraries of electrophilic small molecules. We apply this method prospectively to discover reversible covalent fragments that target distinct protein nucleophiles, including the catalytic serine of AmpC β-lactamase and noncatalytic cysteines in RSK2, MSK1 and JAK3 kinases. We identify submicromolar to low-nanomolar hits with high ligand efficiency, cellular activity and selectivity, including what are to our knowledge the first reported reversible covalent inhibitors of JAK3. Crystal structures of inhibitor complexes with AmpC and RSK2 confirm the docking predictions and guide further optimization. As covalent virtual screening may have broad utility for the rapid discovery of chemical probes, we have made the method freely available through an automated web server (http://covalent.docking.org/).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
22 December 2014
In the version of this article initially published, the scaffold structure shown for compounds 33 and 34 was incorrect in Figure 4a, with a nitrogen atom misplaced within the five-membered ring moiety. The error has been corrected in the HTML and PDF versions of the article.
References
Weerapana, E., Simon, G.M. & Cravatt, B.F. Disparate proteome reactivity profiles of carbon electrophiles. Nat. Chem. Biol. 4, 405–407 (2008).
Blum, G., von Degenfeld, G., Merchant, M.J., Blau, H.M. & Bogyo, M. Noninvasive optical imaging of cysteine protease activity using fluorescently quenched activity-based probes. Nat. Chem. Biol. 3, 668–677 (2007).
Cohen, M.S., Zhang, C., Shokat, K.M. & Taunton, J. Structural bioinformatics-based design of selective, irreversible kinase inhibitors. Science 308, 1318–1321 (2005).
Chang, J.W., Nomura, D.K. & Cravatt, B.F. A potent and selective inhibitor of KIAA1363/AADACL1 that impairs prostate cancer pathogenesis. Chem. Biol. 18, 476–484 (2011).
Robertson, J.G. Mechanistic basis of enzyme-targeted drugs. Biochemistry 44, 5561–5571 (2005).
Drahl, C., Cravatt, B.F. & Sorensen, E.J. Protein-reactive natural products. Angew. Chem. Int. Edn Engl. 44, 5788–5809 (2005).
Kathman, S.G., Xu, Z. & Statsyuk, A.V. A fragment-based method to discover irreversible covalent inhibitors of cysteine proteases. J. Med. Chem. 57, 4969–4974 (2014).
Sirois, S., Hatzakis, G., Wei, D., Du, Q. & Chou, K.C. Assessment of chemical libraries for their druggability. Comput. Biol. Chem. 29, 55–67 (2005).
Baell, J.B. & Holloway, G.A. New substructure filters for removal of pan assay interference compounds (PAINS) from screening libraries and for their exclusion in bioassays. J. Med. Chem. 53, 2719–2740 (2010).
Potashman, M.H. & Duggan, M.E. Covalent modifiers: an orthogonal approach to drug design. J. Med. Chem. 52, 1231–1246 (2009).
Weerapana, E. et al. Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature 468, 790–795 (2010).
Totrov, M. & Abagyan, R. Flexible ligand docking to multiple receptor conformations: a practical alternative. Curr. Opin. Struct. Biol. 18, 178–184 (2008).
Shoichet, B.K. Virtual screening of chemical libraries. Nature 432, 862–865 (2004).
Kitchen, D.B., Decornez, H., Furr, J.R. & Bajorath, J. Docking and scoring in virtual screening for drug discovery: methods and applications. Nat. Rev. Drug Discov. 3, 935–949 (2004).
Glick, M. & Jacoby, E. The role of computational methods in the identification of bioactive compounds. Curr. Opin. Chem. Biol. 15, 540–546 (2011).
Ouyang, X. et al. CovalentDock: automated covalent docking with parameterized covalent linkage energy estimation and molecular geometry constraints. J. Comput. Chem. 34, 326–336 (2013).
Toledo Warshaviak, D., Golan, G., Borrelli, K.W., Zhu, K. & Kalid, O. Structure-based virtual screening approach for discovery of covalently bound ligands. J. Chem. Inf. Model. 54, 1941–1950 (2014).
Zhu, K. et al. Docking covalent inhibitors: a parameter free approach to pose prediction and scoring. J. Chem. Inf. Model. 54, 1932–1940 (2014).
Schröder, J. et al. Docking-based virtual screening of covalently binding ligands: an orthogonal lead discovery approach. J. Med. Chem. 56, 1478–1490 (2013).
De Cesco, S. et al. Virtual screening and computational optimization for the discovery of covalent prolyl oligopeptidase inhibitors with activity in human cells. J. Med. Chem. 55, 6306–6315 (2012).
Mysinger, M.M. & Shoichet, B.K. Rapid context-dependent ligand desolvation in molecular docking. J. Chem. Inf. Model. 50, 1561–1573 (2010).
Jacoby, G.A. AmpC β-lactamases. Clin. Microbiol. Rev. 10.1128/CMR.00036-08 (2009).
Livermore, D.M. & Mushtaq, S. Activity of biapenem (RPX2003) combined with the boronate β-lactamase inhibitor RPX7009 against carbapenem-resistant Enterobacteriaceae. J. Antimicrob. Chemother. 68, 1825–1831 (2013).
Eidam, O. et al. Fragment-guided design of subnanomolar β-lactamase inhibitors active in vivo. Proc. Natl. Acad. Sci. USA 109, 17448–17453 (2012).
Wikler, M.A. Performance Standards for Antimicrobial Susceptibility Testing: Twentieth Informational Supplement (Clinical and Laboratory Standards Institute, 2010).
Serafimova, I.M. et al. Reversible targeting of noncatalytic cysteines with chemically tuned electrophiles. Nat. Chem. Biol. 8, 471–476 (2012).
Miller, R.M., Paavilainen, V.O., Krishnan, S., Serafimova, I.M. & Taunton, J. Electrophilic fragment-based design of reversible covalent kinase inhibitors. J. Am. Chem. Soc. 135, 5298–5301 (2013).
Doehn, U. et al. RSK is a principal effector of the RAS-ERK pathway for eliciting a coordinate promotile/invasive gene program and phenotype in epithelial cells. Mol. Cell 35, 511–522 (2009).
Kang, S. et al. p90 ribosomal S6 kinase 2 promotes invasion and metastasis of human head and neck squamous cell carcinoma cells. J. Clin. Invest. 120, 1165–1177 (2010).
Park, J. et al. RAS-MAPK-MSK1 pathway modulates ataxin 1 protein levels and toxicity in SCA1. Nature 498, 325–331 (2013).
Le, N.T. et al. A crucial role for p90RSK-mediated reduction of ERK5 transcriptional activity in endothelial dysfunction and atherosclerosis. Circulation 127, 486–499 (2013).
Irwin, J.J., Sterling, T., Mysinger, M.M., Bolstad, E.S. & Coleman, R.G. ZINC: a free tool to discover chemistry for biology. J. Chem. Inf. Model. 52, 1757–1768 (2012).
Yamaoka, K. et al. The Janus kinases (Jaks). Genome Biol. 5, 253 (2004).
Kremer, J.M. et al. A phase IIb dose-ranging study of the oral JAK inhibitor tofacitinib (CP-690,550) versus placebo in combination with background methotrexate in patients with active rheumatoid arthritis and an inadequate response to methotrexate alone. Arthritis Rheum. 64, 970–981 (2012).
Jiang, J.K. et al. Examining the chirality, conformation and selective kinase inhibition of 3-((3R,4R)-4-methyl-3-(methyl(7H-pyrrolo[2,3-d]pyrimidin-4-yl)amino)piperidin-1-y l)-3-oxopropanenitrile (CP-690,550). J. Med. Chem. 51, 8012–8018 (2008).
Fleischmann, R. et al. Placebo-controlled trial of tofacitinib monotherapy in rheumatoid arthritis. N. Engl. J. Med. 367, 495–507 (2012).
Clark, J.D., Flanagan, M.E. & Telliez, J.B. Discovery and development of Janus kinase (JAK) inhibitors for inflammatory diseases. J. Med. Chem. 57, 5023–5038 (2014).
Honigberg, L.A. et al. The Bruton tyrosine kinase inhibitor PCI-32765 blocks B-cell activation and is efficacious in models of autoimmune disease and B-cell malignancy. Proc. Natl. Acad. Sci. USA 107, 13075–13080 (2010).
Smith, A.J., Zhang, X., Leach, A.G. & Houk, K.N. Beyond picomolar affinities: quantitative aspects of noncovalent and covalent binding of drugs to proteins. J. Med. Chem. 52, 225–233 (2009).
Hermann, J.C. et al. Structure-based activity prediction for an enzyme of unknown function. Nature 448, 775–779 (2007).
Schwöbel, J.A. et al. Prediction of michael-type acceptor reactivity toward glutathione. Chem. Res. Toxicol. 23, 1576–1585 (2010).
Fischer, M., Coleman, R.G., Fraser, J.S. & Shoichet, B.K. Incorporation of protein flexibility and conformational energy penalties in docking screens to improve ligand discovery. Nat. Chem. 6, 575–583 (2014).
Gasteiger, J., Rudolph, C. & Sadowski, J. Automatic generation of 3D-atomic coordinates for organic molecules. Tetrahedron Computer Methodology 3, 537–547 (1990).
Li, J. et al. Extension of the platform of applicability of the SM5. 42R universal solvation model. Theor. Chem. Acc. 103, 9–63 (1999).
Hawkins, P.C., Skillman, A.G., Warren, G.L., Ellingson, B.A. & Stahl, M.T. Conformer generation with OMEGA: algorithm and validation using high quality structures from the Protein Databank and Cambridge Structural Database. J. Chem. Inf. Model. 50, 572–584 (2010).
Mysinger, M.M., Carchia, M., Irwin, J.J. & Shoichet, B.K. Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better benchmarking. J. Med. Chem. 55, 6582–6594 (2012).
Gilson, M.K., Sharp, K.A. & Honig, B.H. Calculating the electrostatic potential of molecules in solution: method and error assessment. J. Comput. Chem. 9, 327–335 (1988).
Gaulton, A. et al. ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res. 40, D1100–D1107 (2012).
ADAMS, J. et al. Boronic acids and esters as inhibitors of fatty acid amide hydrolase. WO Patent 2,008,063,300 (2008).
Rogers, D. & Hahn, M. Extended-connectivity fingerprints. J. Chem. Inf. Model. 50, 742–754 (2010).
Brozell, S.R. et al. Evaluation of DOCK 6 as a pose generation and database enrichment tool. J. Comput. Aided Mol. Des. 26, 749–773 (2012).
Del Mar, E.G., Largman, C., Brodrick, J.W., Fassett, M. & Geokas, M.C. Substrate specificity of human pancreatic elastase 2. Biochemistry 19, 468–472 (1980).
Pouvreau, L. et al. Effect of pea and bovine trypsin inhibitors on wild-type and modified trypsins. FEBS Lett. 423, 167–172 (1998).
Rodríguez-Martínez, J.A., Rivera-Rivera, I., Sola, R.J. & Griebenow, K. Enzymatic activity and thermal stability of PEG-alpha-chymotrypsin conjugates. Biotechnol. Lett. 31, 883–887 (2009).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Moriarty, N.W., Grosse-Kunstleve, R.W. & Adams, P.D. electronic Ligand Builder and Optimization Workbench (eLBOW): a tool for ligand coordinate and restraint generation. Acta Crystallogr. D Biol. Crystallogr. 65, 1074–1080 (2009).
Painter, J. & Merritt, E.A. TLSMD web server for the generation of multi-group TLS models. J. Appl. Crystallogr. 39, 109–111 (2006).
Knight, Z.A., Feldman, M.E., Balla, A., Balla, T. & Shokat, K.M. A membrane capture assay for lipid kinase activity. Nat. Protoc. 2, 2459–2466 (2007).
Acknowledgements
Computational methods supported by US National Institutes of Health (NIH) grant GM59957, and the web portal was supported by NIH GM71896. This work was also supported by the Ministère de la Recherche et de la Technologie, the Institut national de la santé et de la recherche médicale (UMR Inserm U1071), the Institut National de la Recherche Agronomique (USC-2018) and the Centre Hospitalier Régional Universitaire de Clermont-Ferrand, France (to R.B.). We thank M. Fischer and D. Shaya for help with X-ray data collection, A. O'Donoghue (UCSF) for protease substrates, P. Coffino and S. Menant (UCSF) for the proteasome and the PA26 complex sample, X. Ouyang (Nanyang Technological University) for the r.m.s. calculation software used for the β-lactam benchmark, and S. Barelier for reading of this manuscript. N.L. was supported by an EMBO long-term fellowship (ALTF 1121-2011) and the University of California–San Francisco Program for Breakthrough Biomedical Research, which is funded in part by the Sandler Foundation. S.K. was supported by a fellowship from the California Tobacco-Related Disease Research Program (no. 19FT-0091). P.C. was supported by Howard Hughes Medical Institute Predoctoral Fellowship.
Author information
Authors and Affiliations
Contributions
B.K.S. and J.T. directed the project. N.L. designed the algorithms, wrote the covalent docking code and executed the docking simulations. N.L. performed β-lactamase assays and crystallography with help from O.E. J.J.I. designed and implemented the DOCKovalent web tool. P.C. performed the proteasome experiments. L.G. and R.B. performed bacterial cell culture experiments. R.M.M. executed synthetic chemistry, kinase assays, crystallography and cell-based assays for RSK2 and MSK1. S.K. and K.U. performed synthetic chemistry for JAK3, S.K. performed JAK1 and JAK3 kinase assays. N.L., R.M.M., B.K.S. and J.T. wrote the paper. All authors contributed to the manuscript in its final form.
Corresponding authors
Ethics declarations
Competing interests
J.T., R.M.M. and S.K. have filed patent applications on cyanoacrylamide kinase inhibitors (licensed to Principia Biopharma, of which J.T. is a cofounder).
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–11, Supplementary Figures 1–17 and Supplementary Notes 1 and 2. (PDF 7163 kb)
Rights and permissions
About this article
Cite this article
London, N., Miller, R., Krishnan, S. et al. Covalent docking of large libraries for the discovery of chemical probes. Nat Chem Biol 10, 1066–1072 (2014). https://doi.org/10.1038/nchembio.1666
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1666
This article is cited by
-
Docking covalent targets for drug discovery: stimulating the computer-aided drug design community of possible pitfalls and erroneous practices
Molecular Diversity (2023)
-
An update on the discovery and development of reversible covalent inhibitors
Medicinal Chemistry Research (2023)
-
Assessing the performance of docking, FEP, and MM/GBSA methods on a series of KLK6 inhibitors
Journal of Computer-Aided Molecular Design (2023)
-
Advances in covalent drug discovery
Nature Reviews Drug Discovery (2022)
-
Covalent docking in CDOCKER
Journal of Computer-Aided Molecular Design (2022)