Abstract
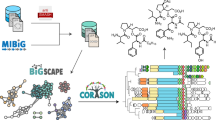
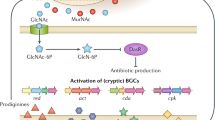
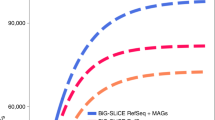
Actinobacteria encode a wealth of natural product biosynthetic gene clusters, whose systematic study is complicated by numerous repetitive motifs. By combining several metrics, we developed a method for the global classification of these gene clusters into families (GCFs) and analyzed the biosynthetic capacity of Actinobacteria in 830 genome sequences, including 344 obtained for this project. The GCF network, comprising 11,422 gene clusters grouped into 4,122 GCFs, was validated in hundreds of strains by correlating confident mass spectrometric detection of known small molecules with the presence or absence of their established biosynthetic gene clusters. The method also linked previously unassigned GCFs to known natural products, an approach that will enable de novo, bioassay-free discovery of new natural products using large data sets. Extrapolation from the 830-genome data set reveals that Actinobacteria encode hundreds of thousands of future drug leads, and the strong correlation between phylogeny and GCFs frames a roadmap to efficiently access them.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Bérdy, J. Bioactive microbial metabolites. J. Antibiot. (Tokyo) 58, 1–26 (2005).
Bérdy, J. Thoughts and facts about antibiotics: where we are now and where we are heading. J. Antibiot. (Tokyo) 65, 385–395 (2012).
Bentley, S.D. et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417, 141–147 (2002).
Lautru, S., Deeth, R.J., Bailey, L.M. & Challis, G.L. Discovery of a new peptide natural product by Streptomyces coelicolor genome mining. Nat. Chem. Biol. 1, 265–269 (2005).
Kersten, R.D. et al. A mass spectrometry–guided genome mining approach for natural product peptidogenomics. Nat. Chem. Biol. 7, 794–802 (2011).
Ziemert, N. et al. The natural product domain seeker NaPDoS: a phylogeny based bioinformatic tool to classify secondary metabolite gene diversity. PLoS ONE 7, e34064 (2012).
Medema, M.H. et al. antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. 39, W339–W346 (2011).
Conway, K.R. & Boddy, C.N. ClusterMine360: a database of microbial PKS/NRPS biosynthesis. Nucleic Acids Res. 41, D402–D407 (2013).
Diminic, J. et al. Databases of the thiotemplate modular systems (CSDB) and their in silico recombinants (r-CSDB). J. Ind. Microbiol. Biotechnol. 40, 653–659 (2013).
Yadav, G., Gokhale, R.S. & Mohanty, D. SEARCHPKS: a program for detection and analysis of polyketide synthase domains. Nucleic Acids Res. 31, 3654–3658 (2003).
Tae, H., Kong, E.-B. & Park, K. ASMPKS: an analysis system for modular polyketide synthases. BMC Bioinformatics 8, 327 (2007).
Ichikawa, N. et al. DoBISCUIT: a database of secondary metabolite biosynthetic gene clusters. Nucleic Acids Res. 41, D408–D414 (2013).
Caboche, S. et al. NORINE: a database of nonribosomal peptides. Nucleic Acids Res. 36, D326–D331 (2008).
Kim, J. & Yi, G.-S. PKMiner: a database for exploring type II polyketide synthases. BMC Microbiol. 12, 169 (2012).
Fischbach, M.A. & Walsh, C. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: logic, machinery, and mechanisms. Chem. Rev. 106, 3468–3496 (2006).
Raghupathy, N. & Durand, D. Gene cluster statistics with gene families. Mol. Biol. Evol. 26, 957–968 (2009).
Wang, X. et al. Identification and characterization of the actinomycin G gene cluster of Streptomyces iakyrus. Mol. Biosyst. 9, 1286–1289 (2013).
Colwell, R.K. et al. Models and estimators linking individual-based and sample-based rarefaction, extrapolation and comparison of assemblages. J. Plant Ecol. 5, 3–21 (2012).
Doroghazi, J.R. & Metcalf, W.W. Comparative genomics of actinomycetes with a focus on natural product biosynthetic genes. BMC Genomics 14, 611 (2013).
Jensen, P.R., Williams, P.G., Oh, D.C., Zeigler, L. & Fenical, W. Species-specific secondary metabolite production in marine actinomycetes of the genus Salinispora. Appl. Environ. Microbiol. 73, 1146–1152 (2007).
Dunbar, K.L., Melby, J.O. & Mitchell, D.A. YcaO domains use ATP to activate amide backbones during peptide cyclodehydrations. Nat. Chem. Biol. 8, 569–575 (2012).
Charlop-Powers, Z., Owen, J.G., Reddy, B.V.B., Ternei, M.A. & Brady, S.F. Chemical-biogeographic survey of secondary metabolism in soil. Proc. Natl. Acad. Sci. USA 111, 3757–3762 (2014).
Bunge, J., Willis, A. & Walsh, F. Estimating the number of species in microbial diversity studies. Annual Review of Statistics and Its Application 1, 427–445 (2014).
Nguyen, D.D. et al. MS/MS networking guided analysis of molecule and gene cluster families. Proc. Natl. Acad. Sci. USA 110, E2611–E2620 (2013).
Cote, R. in ATCC Bacteria and Bacteriophages 19th edn (eds. Pienta, P., Tang, J. & Cote, R.) 484 (American Type Culture Collection, 1996).
Zerbino, D.R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Chaisson, M.J., Brinza, D. & Pevzner, P.A. De novo fragment assembly with short mate-paired reads: does the read length matter? Genome Res. 19, 336–346 (2009).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005); erratum Nature 4, 120 (2006).
Boetzer, M., Henkel, C.V., Jansen, H.J., Butler, D. & Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27, 578–579 (2011).
Peng, Y., Leung, H.C., Yiu, S.-M. & Chin, F.Y. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28, 1420–1428 (2012).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Li, L., Stoeckert, C.J. & Roos, D.S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Ester, M., Kriegel, H.-P., Sander, J. & Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. in Proceedings of the Second International Conference on Knowledge Discovery and Data Mining (KDD-96) (eds. Simoudis, E., Han, J. & Fayyad, U.) 226–231 (AAAI Press, 1996).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Price, M.N., Dehal, P.S. & Arkin, A.P. FastTree 2–approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Zhang, Q., Yu, Y., Vélasquez, J.E. & van der Donk, W.A. Evolution of lanthipeptide synthetases. Proc. Natl. Acad. Sci. USA 109, 18361–18366 (2012).
Yutin, N., Puigbò, P., Koonin, E.V. & Wolf, Y.I. Phylogenomics of prokaryotic ribosomal proteins. PLoS ONE 7, e36972 (2012).
Larkin, M.A. et al. Clustal W and clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Huerta-Cepas, J., Dopazo, J. & Gabaldón, T. ETE: a python Environment for Tree Exploration. BMC Bioinformatics 11, 24 (2010).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
El-Nakeeb, M.A. & Lechevalier, H.A. Selective isolation of aerobic actinomycetes. Appl. Microbiol. 11, 75–77 (1963).
Smith, S.E. et al. Comparative genomic and phylogenetic approaches to characterize the role of genetic recombination in mycobacterial evolution. PLoS ONE 7, e50070 (2012).
Acknowledgements
J.R.D. was funded through an Institute for Genomic Biology fellowship. This work was supported in part by US National Institutes of Health grants GM PO1 GM077596 and GM 067725 (N.L.K.) and an Institute for Genomic Biology Proof of Concept grant. D.P.L. and the Agricultural Research Service (ARS) Culture Collection Current Research Information System project is funded through ARS National Program 301.
Author information
Authors and Affiliations
Contributions
J.R.D. designed and performed bioinformatic analyses. J.R.D., R.R.H. and K.A.T. produced microbial extracts. J.C.A. designed LC/MS experiments, and J.C.A. and A.W.G. collected and analyzed LC/MS data. K.-S.J. received and processed germplasm and contributed to genomic library preparation. D.P.L. selected and provided germplasm from the ARS Culture Collection. W.W.M. and N.L.K. designed and directed the work. J.R.D., J.C.A., W.W.M. and N.L.K. wrote the manuscript. Draft genomes sequenced as part of this project are available through NCBI BioProject PRJNA238534.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–3, Supplementary Figures 1–6 and Supplementary Note. (PDF 2814 kb)
Supplementary Data Set 1
A spreadsheet that lists the locus tags and corresponding organism for each genome in this study (XLSX 33 kb)
Supplementary Data Set 2
All gene cluster families that have at least one characterized natural product are listed in this Excel spreadsheet. (XLSX 50 kb)
Supplementary Data Set 3
A spreadsheet containing raw MS data for the known compounds identified in this study. (XLSX 33 kb)
Rights and permissions
About this article
Cite this article
Doroghazi, J., Albright, J., Goering, A. et al. A roadmap for natural product discovery based on large-scale genomics and metabolomics. Nat Chem Biol 10, 963–968 (2014). https://doi.org/10.1038/nchembio.1659
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1659