Abstract
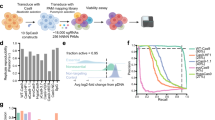
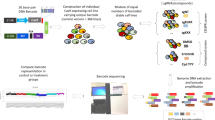
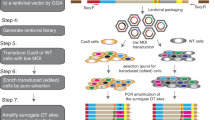
Identification and validation of drug-resistant mutations can provide important insights into the mechanism of action of a compound. Here we demonstrate the feasibility of such an approach in mammalian cells using next-generation sequencing of drug-resistant clones and CRISPR-Cas9–mediated gene editing on two drug-target pairs, 6-thioguanine–HPRT1 and triptolide-ERCC3. We showed that disrupting functional HPRT1 allele or introducing ERCC3 point mutations by gene editing can confer drug resistance in cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Heitman, J., Movva, N.R. & Hall, M.N. Science 253, 905–909 (1991).
Wacker, S.A., Houghtaling, B.R., Elemento, O. & Kapoor, T.M. Nat. Chem. Biol. 8, 235–237 (2012).
Jinek, M. et al. Science 337, 816–821 (2012).
Mali, P. et al. Science 339, 823–826 (2013).
Ran, F.A. et al. Nat. Protoc. 8, 2281–2308 (2013).
Fu, Y. et al. Nat. Biotechnol. 31, 822–826 (2013).
Andersson, B.S. et al. Leukemia 9, 2100–2108 (1995).
Glaab, W.E. et al. Carcinogenesis 19, 1931–1937 (1998).
Chugh, R. et al. Sci. Transl. Med. 4, 156ra139 (2012).
Titov, D.V. et al. Nat. Chem. Biol. 7, 182–188 (2011).
Corson, T.W., Cavga, H., Aberle, N. & Crews, C.M. ChemBioChem. 12, 1767–1773 (2011).
Leuenroth, S.J. et al. Proc. Natl. Acad. Sci. USA 104, 4389–4394 (2007).
Lu, Y. et al. Chem. Biol. 21, 246–256 (2014).
Coin, F., Oksenych, V. & Egly, J.M. Mol. Cell 26, 245–256 (2007).
Jawhari, A. et al. J. Biol. Chem. 277, 31761–31767 (2002).
Soundararajan, R., Sayat, R., Robertson, G.S. & Marignani, P.A. Cancer Biol. Ther. 8, 2054–2062 (2009).
Wang, H. et al. Cell 153, 910–918 (2013).
Auer, T.O., Duroure, K., De Cian, A., Concordet, J.P. & Del Bene, F. Genome Res. 24, 142–153 (2014).
Chen, F. et al. Nat. Methods 8, 753–755 (2011).
You, F.M. et al. BMC Bioinformatics 9, 253 (2008).
McKenna, A. et al. Genome Res. 20, 1297–1303 (2010).
Van der Auwera, G.A. et al. Curr. Prot. Bioinformatics 43, 11.10 (2013).
DePristo, M.A. et al. Nat. Genet. 43, 491–498 (2011).
Li, H. & Durbin, R. Bioinformatics 25, 1754–1760 (2009).
Wang, K., Li, M. & Hakonarson, H. Nucleic Acids Res. 38, e164 (2010).
Sherry, S.T. et al. Nucleic Acids Res. 29, 308–311 (2001).
Yang, L. et al. Nucleic Acids Res. 41, 9049–9061 (2013).
Bedell, V.M. et al. Nature 491, 114–118 (2012).
Maresca, M., Lin, V.G., Guo, N. & Yang, Y. Genome Res. 23, 539–546 (2013).
Acknowledgements
The authors would like to thank N. Guo for technical help on CRISPR-Cas9 cloning. Y.S. and M.C. would like to thank the Novartis Postdoc Office for support during their training.
Author information
Authors and Affiliations
Contributions
Y.F., Y.Y., J.A.T. and Y.S. conceived the study, Y.S., M.C., H.W. and E.M. conducted experiments, Y.S. performed bioinformatic analysis, and Y.S. and Y.F. wrote the manuscript with additional help from all authors.
Corresponding author
Ethics declarations
Competing interests
At the time the research was performed, the authors were employees of Novartis, and the research was fully funded by Novartis.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–14 and Supplementary Figures 1–11. (PDF 1443 kb)
Supplementary Data Set 1
Results of next generation exome sequencing of Triptolide resistant KBM7 clones (XLSX 36 kb)
Rights and permissions
About this article
Cite this article
Smurnyy, Y., Cai, M., Wu, H. et al. DNA sequencing and CRISPR-Cas9 gene editing for target validation in mammalian cells. Nat Chem Biol 10, 623–625 (2014). https://doi.org/10.1038/nchembio.1550
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1550
This article is cited by
-
Targeting pan-essential pathways in cancer with cytotoxic chemotherapy: challenges and opportunities
Cancer Chemotherapy and Pharmacology (2023)
-
An update on CRISPR-Cas12 as a versatile tool in genome editing
Molecular Biology Reports (2023)
-
Receptor clustering and pathogenic complement activation in myasthenia gravis depend on synergy between antibodies with multiple subunit specificities
Acta Neuropathologica (2022)
-
Modular deep learning enables automated identification of monoclonal cell lines
Nature Machine Intelligence (2021)
-
Target identification of small molecules using large-scale CRISPR-Cas mutagenesis scanning of essential genes
Nature Communications (2018)