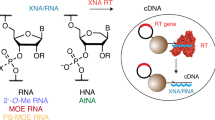
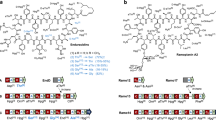
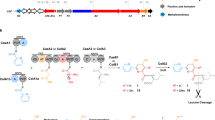
Abstract
Concatenation of engineered biocatalysts into multistep pathways markedly increases their utility, but the development of generalizable assembly methods remains a major challenge. Herein we evaluate 'bioretrosynthesis', which is an application of the retrograde evolution hypothesis, for biosynthetic pathway construction. To test bioretrosynthesis, we engineered a pathway for synthesis of the antiretroviral nucleoside analog didanosine (2′,3′-dideoxyinosine). Applying both directed evolution– and structure-based approaches, we began pathway construction with a retro-extension from an engineered purine nucleoside phosphorylase and evolved 1,5-phosphopentomutase to accept the substrate 2,3-dideoxyribose 5-phosphate with a 700-fold change in substrate selectivity and threefold increased turnover in cell lysate. A subsequent retrograde pathway extension, via ribokinase engineering, resulted in a didanosine pathway with a 9,500-fold change in nucleoside production selectivity and 50-fold increase in didanosine production. Unexpectedly, the result of this bioretrosynthetic step was not a retro-extension from phosphopentomutase but rather the discovery of a fortuitous pathway-shortening bypass via the engineered ribokinase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Savile, C.K. et al. Biocatalytic asymmetric synthesis of chiral amines from ketones applied to sitagliptin manufacture. Science 329, 305–309 (2010).
Liang, J. et al. Development of a biocatalytic process as an alternative to the (–)-DIP-Cl–mediated asymmetric reduction of a key intermediate of montelukast. Org. Process Res. Dev. 14, 193–198 (2010).
Gao, X. et al. Directed evolution and structural characterization of a simvastatin synthase. Chem. Biol. 16, 1064–1074 (2009).
Bornscheuer, U.T. et al. Engineering the third wave of biocatalysis. Nature 485, 185–194 (2012).
Paddon, C.J. et al. High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 496, 528–532 (2013).
Ajikumar, P.K. et al. Isoprenoid pathway optimization for Taxol precursor overproduction in Escherichia coli. Science 330, 70–74 (2010).
Ito, T. et al. Deciphering pactamycin biosynthesis and engineered production of new pactamycin analogues. ChemBioChem 10, 2253–2265 (2009).
Zhang, M.-Q. et al. Optimizing natural products by biosynthetic engineering: discovery of nonquinone Hsp90 inhibitors. J. Med. Chem. 51, 5494–5497 (2008).
Niu, W., Molefe, M.N. & Frost, J.W. Microbial synthesis of the energetic material precursor 1,2,4-butanetriol. J. Am. Chem. Soc. 125, 12998–12999 (2003).
Yim, H. et al. Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol. Nat. Chem. Biol. 7, 445–452 (2011).
Ma, S.K. et al. A green-by-design biocatalytic process for atorvastatin intermediate. Green Chem. 12, 81 (2010).
Pinheiro, E. et al. Examining the production costs of antiretroviral drugs. AIDS 20, 1745–1752 (2006).
Medema, M.H., van Raaphorst, R., Takano, E. & Breitling, R. Computational tools for the synthetic design of biochemical pathways. Nat. Rev. Microbiol. 10, 191–202 (2012).
Eriksen, D.T., Lian, J. & Zhao, H. Protein design for pathway engineering. J. Struct. Biol. 185, 234 (2014).
Horowitz, N.H. On the evolution of biochemical syntheses. Proc. Natl. Acad. Sci. USA 31, 153–157 (1945).
Bachmann, B.O. Biosynthesis: is it time to go retro? Nat. Chem. Biol. 6, 390–393 (2010).
Corey, E.J. The logic of chemical synthesis—multistep synthesis of complex natural carbogenic molecules. Angew. Chem. Int. Edn Engl. 30, 455–465 (1991).
Turner, N.J. & O'Reilly, E. Biocatalytic retrosynthesis. Nat. Chem. Biol. 9, 285–288 (2013).
Nannemann, D.P., Kaufmann, K.W., Meiler, J. & Bachmann, B.O. Design and directed evolution of a dideoxy purine nucleoside phosphorylase. Protein Eng. Des. Sel. 23, 607–616 (2010).
Hamamoto, T., Noguchi, T. & Midorikawa, Y. Phosphopentomutase of Bacillus stearothermophilus TH6–2: the enzyme and its gene ppm. Biosci. Biotechnol. Biochem. 62, 1103–1108 (1998).
Panosian, T.D. et al. Bacillus cereus phosphopentomutase is an alkaline phosphatase family member that exhibits an altered entry point into the catalytic cycle. J. Biol. Chem. 286, 8043–8054 (2011).
Patrick, W.M., Firth, A.E. & Blackburn, J.M. User-friendly algorithms for estimating completeness and diversity in randomized protein-encoding libraries. Protein Eng. 16, 451–457 (2003).
Scism, R.A. & Bachmann, B.O. Five-component cascade synthesis of nucleotide analogues in an engineered self-immobilized enzyme aggregate. ChemBioChem 11, 67–70 (2010).
Li, J. et al. Crystal structure of Sa239 reveals the structural basis for the activation of ribokinase by monovalent cations. J. Struct. Biol. 177, 578–582 (2012).
Nocek, B. et al. Structural studies of ROK fructokinase YdhR from Bacillus subtilis: insights into substrate binding and fructose specificity. J. Mol. Biol. 406, 325–342 (2011).
Charrier, V. et al. Cloning and sequencing of two Enterococcal glpK genes and regulation of the encoded glycerol kinases by phosphoenolpyruvate dependent, phosphotransferase system–catalyzed phosphorylation of a single histidyl residue. J. Biol. Chem. 272, 14166–14174 (1997).
Campobasso, N., Mathews, I.I., Begley, T.P. & Ealick, S.E. Crystal structure of 4-methyl-5-β-hydroxyethylthiazole kinase from Bacillus subtilis at 1.5 Å resolution. Biochemistry 39, 7868–7877 (2000).
Andersson, C.E. & Mowbray, S.L. Activation of ribokinase by monovalent cations. J. Mol. Biol. 315, 409–419 (2002).
Sigrell, J.A., Cameron, A.D., Jones, T.A. & Mowbray, S.L. Structure of Escherichia coli ribokinase in complex with ribose and dinucleotide determined to 1.8 Å resolution: insights into a new family of kinase structures. Structure 6, 183–193 (1998).
Datta, R. et al. Homology-model–guided site-specific mutagenesis reveals the mechanisms of substrate binding and product-regulation of adenosine kinase from Leishmania donovani. Biochem. J. 394, 35–42 (2006).
Zhang, Y., Dougherty, M., Downs, D.M. & Ealick, S.E. Crystal structure of an aminoimidazole riboside kinase from Salmonella enterica: implications for the evolution of the ribokinase superfamily. Structure 12, 1809–1821 (2004).
Currie, M.A. et al. ADP-dependent 6-phosphofructokinase from Pyrococcus horikoshii OT3—structure determination and biochemical characterization of PH1645. J. Biol. Chem. 284, 22664–22671 (2009).
Trinh, C.H., Asipu, A., Bonthron, D.T. & Phillips, S.E. Structures of alternatively spliced isoforms of human ketohexokinase. Acta Crystallogr. D Biol. Crystallogr. 65, 201–211 (2009).
Cabrera, R. et al. The crystal complex of phosphofructokinase-2 of Escherichia coli with fructose-6-phosphate—kinetic and structural analysis of the allosteric ATP inhibition. J. Biol. Chem. 286, 5774–5783 (2011).
Miallau, L., Hunter, W.N., McSweeney, S.M. & Leonard, G.A. Structures of Staphylococcus aureus d-tagatose-6-phosphate kinase implicate domain motions in specificity and mechanism. J. Biol. Chem. 282, 19948–19957 (2007).
Sutherland, J.D., Wilson, E.J. & Wright, M.C. Directed evolution of novel biosynthetic pathways: growth of an Escherichia coli proline auxotroph on Δ1-pyrroline-2-carboxylic acid. Bioorg. Med. Chem. Lett. 3, 1185–1188 (1993).
Bhabha, G. et al. Divergent evolution of protein conformational dynamics in dihydrofolate reductase. Nat. Struct. Mol. Biol. 20, 1243–1249 (2013).
Bhabha, G. et al. A dynamic knockout reveals that conformational fluctuations influence the chemical step of enzyme catalysis. Science 332, 234–238 (2011).
Pfleger, B.F., Pitera, D.J., Smolke, D.C. & Keasling, J.D. Combinatorial engineering of intergenic regions in operons tunes expression of multiple genes. Nat. Biotechnol. 24, 1027–1032 (2006).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Iverson, T.M., Panosian, T.D., Birmingham, W.R., Nannemann, D.P. & Bachmann, B.O. Molecular differences between a mutase and a phosphatase: investigations of the activation step in Bacillus cereus phosphopentomutase. Biochemistry 51, 1964–1975 (2012).
Panosian, T.D., Nannemann, D.P., Bachmann, B.O. & Iverson, T.M. Crystallization and preliminary X-ray analysis of a phosphopentomutase from Bacillus cereus. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 66, 811–814 (2010).
deGroot, H. & Noll, T. Enzymic determination of inorganic phosphates, organic phosphates and phosphate-liberating enzymes by use of nucleoside phosphorylase-xanthine oxidase (dehydrogenase)-coupled reactions. Biochem. J. 230, 255–260 (1985).
Ball, E.G. Xanthine oxidase: purification and properties. J. Biol. Chem. 128, 51–67 (1939).
Bezy, V., Morin, P., Couerbe, P., Leleu, G. & Agrofoglio, L. Simultaneous analysis of several antiretroviral nucleosides in rat-plasma by high-performance liquid chromatography with UV using acetic acid/hydroxylamine buffer—test of this new volatile medium-pH for HPLC-ESI-MS/MS. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 821, 132–143 (2005).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Brunger, A.T. Version 1.2 of the crystallography and NMR system. Nat. Protoc. 2, 2728–2733 (2007).
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Cryst. 30, 1022–1025 (1997).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Winn, M.D., Isupov, M.N. & Murshudov, G.N. Use of TLS parameters to model anisotropic displacements in macromolecular refinement. Acta Crystallogr. D Biol. Crystallogr. 57, 122–133 (2001).
Holm, L. & Park, J. DaliLite workbench for protein structure comparison. Bioinformatics 16, 566–567 (2000).
Acknowledgements
We thank V. Phelan, K. McCulloch and C. Goodwin for assistance with data acquisition. We also thank J. Zang (Chinese Academy of Sciences), A. Joachimiak (Argonne National Laboratory and University of Chicago), J. Deutscher (Centre national de la recherche scientifique) and S. Ealick (Cornell University) for expression plasmids used in this study. This work was supported by the US National Institutes of Health (NIH) grant T32 GM065086 and the D. Stanley and Ann T. Tarbell Endowment fund (W.R.B.), NIH training grant 5T32GM008320, a US National Science Foundation individual graduate fellowship DGE:0909667 (C.A.S.), NIH training grants T32NS07491 and 5T32GM008320 (T.D.P.), NIH training grant T90 DA022873 (D.P.N.), NIH grants GM079419 (to T.M.I.) and GM077189 (B.O.B.) and the Vanderbilt Institute of Chemical Biology. The use of the Advanced Photon Source, an Office of Science User Facility operated for the US Department of Energy (DOE) Office of Science by Argonne National Laboratory, was supported by the US DOE under contract no. DE-AC02-06CH11357. The use of the LS-CAT Sector 21 was supported by the Michigan Economic Development Corporation and the Michigan Technology Tri-Corridor (grant 085P1000817). The use of the Vanderbilt robotic crystallization facility was supported by NIH grant S10 RR026915.
Author information
Authors and Affiliations
Contributions
B.O.B. supervised bioretrosynthetic studies, and T.M.I. supervised the X-ray crystallographic work. W.R.B. and D.P.N. designed assays. W.R.B. performed assays, screened mutagenesis libraries, performed kinetic characterization and tested enzymes in the biosynthetic pathway studies. C.A.S. determined the crystal structures of V158L and 4H11 PPM variants. T.D.P. determined crystal structures of wild-type PPM and the S154A and S154G variants. D.P.N. established initial synthesis routes of dideoxyribose and dideoxyribose 5-phosphate. W.R.B., C.A.S., T.M.I. and B.O.B. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Results, Supplementary Figures 1–20 and Supplementary Tables 1–7. (PDF 4014 kb)
Rights and permissions
About this article
Cite this article
Birmingham, W., Starbird, C., Panosian, T. et al. Bioretrosynthetic construction of a didanosine biosynthetic pathway. Nat Chem Biol 10, 392–399 (2014). https://doi.org/10.1038/nchembio.1494
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1494
This article is cited by
-
Multistep enzyme cascades as a route towards green and sustainable pharmaceutical syntheses
Nature Chemistry (2022)
-
Alkaloid diversification in the genus Palicourea (Rubiaceae: Palicoureeae) viewed from a (retro-)biogenetic perspective
Phytochemistry Reviews (2022)
-
Metabolic engineering of Escherichia coli for de novo production of 3-phenylpropanol via retrobiosynthesis approach
Microbial Cell Factories (2021)
-
Biocatalysis
Nature Reviews Methods Primers (2021)
-
Technical Advances to Accelerate Modular Type I Polyketide Synthase Engineering towards a Retro-biosynthetic Platform
Biotechnology and Bioprocess Engineering (2019)