Abstract
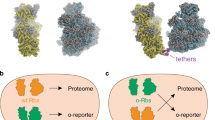
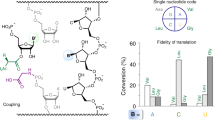
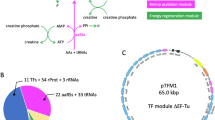
In ribosomal polypeptide synthesis the library of amino acid building blocks is limited by the manner in which codons are used. Of the proteinogenic amino acids, 18 are coded for by multiple codons and therefore many of the 61 sense codons can be considered redundant. Here we report a method to reduce the redundancy of codons by artificially dividing codon boxes to create vacant codons that can then be reassigned to non-proteinogenic amino acids and thereby expand the library of genetically encoded amino acids. To achieve this, we reconstituted a cell-free translation system with 32 in vitro transcripts of transfer RNASNN (tRNASNN) (S = G or C), assigning the initiator and 20 elongator amino acids. Reassignment of three redundant codons was achieved by replacing redundant tRNASNNs with tRNASNNs pre-charged with non-proteinogenic amino acids. As a demonstration, we expressed a 32-mer linear peptide that consists of 20 proteinogenic and three non-proteinogenic amino acids, and a 14-mer macrocyclic peptide that contains more than four non-proteinogenic amino acids.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Passioura, T., Katoh, T., Goto, Y. & Suga, H. Selection-based discovery of druglike macrocyclic peptides. Annu. Rev. Biochem. 83, 727–752 (2014).
Craik, D. J., Fairlie, D. P., Liras, S. & Price, D. The future of peptide-based drugs. Chem. Biol. Drug Des. 81, 136–147 (2013).
Heinis, C. & Winter, G. Encoded libraries of chemically modified peptides. Curr. Opin. Chem. Biol. 26, 89–98 (2015).
Arnison, P. G. et al. Ribosomally synthesized and post-translationally modified peptide natural products: overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 30, 108–160 (2013).
Horswill, A. R. & Benkovic, S. J. Cyclic peptides, a chemical genetics tool for biologists. Cell Cycle 4, 552–555 (2005).
Yamagishi, Y. et al. Natural product-like macrocyclic N-methyl-peptide inhibitors against a ubiquitin ligase uncovered from a ribosome-expressed de novo library. Chem. Biol. 18, 1562–1570 (2011).
Hayashi, Y., Morimoto, J. & Suga, H. In vitro selection of anti-Akt2 thioether-macrocyclic peptides leading to isoform-selective inhibitors. ACS Chem. Biol. 7, 607–613 (2012).
Morimoto, J., Hayashi, Y. & Suga, H. Discovery of macrocyclic peptides armed with a mechanism-based warhead: isoform-selective inhibition of human deacetylase SIRT2. Angew. Chem. Int. Ed. 51, 3423–3427 (2012).
Tanaka, Y. et al. Structural basis for the drug extrusion mechanism by a MATE multidrug transporter. Nature 496, 247–251 (2013).
Nemoto, N., Miyamoto-Sato, E., Husimi, Y. & Yanagawa, H. In vitro virus: bonding of mRNA bearing puromycin at the 3′-terminal end to the C-terminal end of its encoded protein on the ribosome in vitro. FEBS Lett. 414, 405–408 (1997).
Roberts, R. W. & Szostak, J. W. RNA-peptide fusions for the in vitro selection of peptides and proteins. Proc. Natl Acad. Sci. USA 94, 12297–12302 (1997).
Murakami, H., Ohta, A., Ashigai, H. & Suga, H. A highly flexible tRNA acylation method for non-natural polypeptide synthesis. Nature Methods 3, 357–359 (2006).
Goto, Y. et al. Reprogramming the translation initiation for the synthesis of physiologically stable cyclic peptides. ACS Chem. Biol. 3, 120–129 (2008).
Goto, Y., Murakami, H. & Suga, H. Initiating translation with D-amino acids. RNA 14, 1390–1398 (2008).
Kawakami, T., Murakami, H. & Suga, H. Messenger RNA-programmed incorporation of multiple N-methyl-amino acids into linear and cyclic peptides. Chem. Biol. 15, 32–42 (2008).
Xiao, H., Murakami, H., Suga, H. & Ferre-D'Amare, A. R. Structural basis of specific tRNA aminoacylation by a small in vitro selected ribozyme. Nature 454, 358–361 (2008).
Ohta, A., Murakami, H., Higashimura, E. & Suga, H. Synthesis of polyester by means of genetic code reprogramming. Chem. Biol. 14, 1315–1322 (2007).
Goto, Y., Katoh, T. & Suga, H. Flexizymes for genetic code reprogramming. Nature Protocols 6, 779–790 (2011).
Shimizu, Y. et al. Cell-free translation reconstituted with purified components. Nature Biotechnol. 19, 751–755 (2001).
Forster, A. C. et al. Programming peptidomimetic syntheses by translating genetic codes designed de novo. Proc. Natl Acad. Sci. USA 100, 6353–6357 (2003).
Josephson, K., Hartman, M. C. & Szostak, J. W. Ribosomal synthesis of unnatural peptides. J. Am. Chem. Soc. 127, 11727–11735 (2005).
Noren, C. J., Anthony-Cahill, S. J., Griffith, M. C. & Schultz, P. G. A general method for site-specific incorporation of unnatural amino acids into proteins. Science 244, 182–188 (1989).
Wang, L., Brock, A., Herberich, B. & Schultz, P. G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Mukai, T. et al. Codon reassignment in the Escherichia coli genetic code. Nucleic Acids Res. 38, 8188–8195 (2010).
Mukai, T. et al. Genetic-code evolution for protein synthesis with non-natural amino acids. Biochem. Biophys. Res. Commun. 411, 757–761 (2011).
Johnson, D. B. F. et al. RF1 knockout allows ribosomal incorporation of unnatural amino acids at multiple sites. Nature Chem. Biol. 7, 779–786 (2011).
Lajoie, M. J. et al. Genomically recoded organisms expand biological functions. Science 342, 357–360 (2013).
Hohsaka, T., Ashizuka, Y., Murakami, H. & Sisido, M. Incorporation of nonnatural amino acids into streptavidin through in vitro frame-shift suppression. J. Am. Chem. Soc. 118, 9778–9779 (1996).
Magliery, T. J., Anderson, J. C. & Schultz, P. G. Expanding the genetic code: selection of efficient suppressors of four-base codons and identification of shifty’ four-base codons with a library approach in Escherichia coli. J. Mol. Biol. 307, 755–769 (2001).
Ohtsuki, T., Manabe, T. & Sisido, M. Multiple incorporation of non-natural amino acids into a single protein using tRNAs with non-standard structures. FEBS Lett. 579, 6769–6774 (2005).
Neumann, H., Wang, K., Davis, L., Garcia-Alai, M. & Chin, J. W. Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome. Nature 464, 441–444 (2010).
Lajoie, M. J. et al. Probing the limits of genetic recoding in essential genes. Science 342, 361–363 (2013).
Komine, Y., Adachi, T., Inokuchi, H. & Ozeki, H. Genomic organization and physical mapping of the transfer RNA genes in Escherichia coli K12. J. Mol. Biol. 212, 579–598 (1990).
Murphy, F. V. IV & Ramakrishnan, V. Structure of a purine–purine wobble base pair in the decoding center of the ribosome. Nature Struct. Mol. Biol. 11, 1251–1252 (2004).
Gustilo, E. M., Vendeix, F. A. & Agris, P. F. tRNA's modifications bring order to gene expression. Curr. Opin. Microbiol. 11, 134–140 (2008).
Crick, F. H. Codon–anticodon pairing: the wobble hypothesis. J. Mol. Biol. 19, 548–555 (1966).
Vasil'eva, I. A. & Moor, N. A. Interaction of aminoacyl-tRNA synthetases with tRNA: general principles and distinguishing characteristics of the high-molecular-weight substrate recognition. Biochemistry. Biokhim. 72, 247–263 (2007).
Giege, R., Sissler, M. & Florentz, C. Universal rules and idiosyncratic features in tRNA identity. Nucleic Acids Res. 26, 5017–5035 (1998).
Tamura, K., Himeno, H., Asahara, H., Hasegawa, T. & Shimizu, M. In vitro study of E. coli tRNAArg and tRNALys identity elements. Nucleic Acids Res. 20, 2335–2339 (1992).
Nureki, O. et al. Molecular recognition of the identity-determinant set of isoleucine transfer RNA from Escherichia coli. J. Mol. Biol. 236, 710–724 (1994).
Sylvers, L. A., Rogers, K. C., Shimizu, M., Ohtsuka, E. & Soll, D. A 2-thiouridine derivative in tRNAGlu is a positive determinant for aminoacylation by Escherichia coli glutamyl-tRNA synthetase. Biochemistry 32, 3836–3841 (1993).
Cui, Z., Stein, V., Tnimov, Z., Mureev, S. & Alexandrov, K. Semisynthetic tRNA complement mediates in vitro protein synthesis. J. Am. Chem. Soc. 137, 4404–4413 (2015).
Park, H. S. et al. Expanding the genetic code of Escherichia coli with phosphoserine. Science 333, 1151–1154 (2011).
Schlippe, Y. V., Hartman, M. C., Josephson, K. & Szostak, J. W. In vitro selection of highly modified cyclic peptides that act as tight binding inhibitors. J. Am. Chem. Soc. 134, 10469–10477 (2012).
Acknowledgements
We thank M. E. Harris and M. Ohuchi for a gift of plasmids that coded E. coli M1 RNA and C5 protein and their products. We thank S. Jongkees and J. Rogers for proofreading the manuscript. This research was supported by the Japan Science and Technology Agency (JST) Core Research for Evolutional Science and Technology (CREST) of Molecular Technologies to H.S., Japan Society for the Promotion of Science (JSPS) Grant-in-Aid for Young Scientists (B) to Y.G. (22750145) and Grants-in-Aid for JSPS Fellows to Y.I. (26-9576).
Author information
Authors and Affiliations
Contributions
Y.I., A.H., H.M., T.K., Y.G. and H.S. designed the experiments and analysed the data. Y.I. and A.H. performed the experiments. Y.I., T.K., Y.G. and H.S. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 2239 kb)
Rights and permissions
About this article
Cite this article
Iwane, Y., Hitomi, A., Murakami, H. et al. Expanding the amino acid repertoire of ribosomal polypeptide synthesis via the artificial division of codon boxes. Nature Chem 8, 317–325 (2016). https://doi.org/10.1038/nchem.2446
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2446
This article is cited by
-
Cell-free Biosynthesis of Peptidomimetics
Biotechnology and Bioprocess Engineering (2023)
-
Evolution of KIPPIS as a versatile platform for evaluating intracellularly functional peptide aptamers
Scientific Reports (2021)
-
Reconstituted cell-free protein synthesis using in vitro transcribed tRNAs
Communications Biology (2020)
-
Deciphering protein post-translational modifications using chemical biology tools
Nature Reviews Chemistry (2020)
-
Ribosome-mediated polymerization of long chain carbon and cyclic amino acids into peptides in vitro
Nature Communications (2020)