Abstract
Site-selective functionalization of complex molecules is one of the most significant challenges in chemistry. Typically, protecting groups or catalysts must be used to enable the selective modification of one site among many that are similarly reactive, and general strategies that selectively tune the local chemical environment around a target site are rare. Here, we show a four-amino-acid sequence (Phe-Cys-Pro-Phe), which we call the ‘π-clamp’, that tunes the reactivity of its cysteine thiol for site-selective conjugation with perfluoroaromatic reagents. We use the π-clamp to selectively modify one cysteine site in proteins containing multiple endogenous cysteine residues. These examples include antibodies and cysteine-based enzymes that would be difficult to modify selectively using standard cysteine-based methods. Antibodies modified using the π-clamp retained binding affinity to their targets, enabling the synthesis of site-specific antibody–drug conjugates for selective killing of HER2-positive breast cancer cells. The π-clamp is an unexpected approach to mediate site-selective chemistry and provides new avenues to modify biomolecules for research and therapeutics.
Similar content being viewed by others
Main
Site-selective chemistry1,2,3,4,5 is essential for creating homogeneously modified biologics6,7, studying protein structure and function8, generating materials with defined composition9, and on-demand modification of complex small molecules10,11. Existing approaches for site-selective chemistry use either reaction pairs that are orthogonal to other functional groups on the target of interest (Fig. 1a, strategy 1)12,13 or catalysts that mediate selective reactions at a particular site among many competing ones (Fig. 1a, strategy 2)14,15,16,17,18,19. These strategies have been widely used in protein modification and have led to the development of multiple bio-orthogonal handles20,21,22,23,24,25 and enzyme–tag pairs26,27,28,29,30,31.
a, Existing strategies for site-selective chemistry. Strategy 1: selectivity arises from orthogonal chemistry between site Z and reagent Y. Strategy 2: catalyst mediates the reaction between a particular site X (highlighted in red) and reagent Y. b, This work demonstrates a new strategy for site-selective chemistry by fine-tuning the local chemical environment around the target site. A particular site X (highlighted in red) is tuned to react with reagent Y in the presence of other competing X sites. c, A cysteine residue inside the π-clamp selectively reacts with perfluoroaromatic probes in the presence of other competing cysteine residues and thiol species.
Natural proteins precisely control selective reactions and interactions by building large three-dimensional structures from polypeptides usually much greater than 100 residues32. For example, enzymes have folded structures where particular amino acids are placed in a specialized active-site environment33. Inspired by this, we envisioned a new strategy for site-selective chemistry on proteins by fine-tuning the local environment around an amino-acid residue in a small peptide sequence (Fig. 1b). This is difficult, because peptides are highly dynamic and unstructured, thereby presenting a formidable challenge to building defined environments for selective chemical transformations.
Our design efforts leveraged cysteine, because nature has shown its robust catalytic role in enzymes34,35, and previous efforts indicate that the reactivity of a cysteine residue can vary in different protein environments36. Furthermore, cysteine is the first choice in bioconjugation to modify proteins, often via maleimide ligation or alkylation37,38. However, these traditional cysteine-based bioconjugations are significantly limited because they are not site-specific. When these methods are applied to protein targets with multiple cysteine residues, a mixture of heterogeneous products are generated, as exemplified by recent efforts to conjugate small-molecule drugs to antibodies through cysteine-based reactions39.
Small peptide tags that contain multiple cysteine residues have been used for bioconjugation. Tsien and co-workers have developed biarsenic reagents that selectively react with tetra-cysteine motifs in peptides and proteins40,41. More recently, organic arsenics have been used to modify two cysteine residues generated from reducing a disulfide bond42. These methods can present challenges with thiol selectivity43, and none report the site-specific modification of one cysteine residue in the presence of many, as enzymes or multiple chemical steps must be used to accomplish this feat44,45. An enzyme-free and one-step method for site-selective cysteine conjugation has yet to be developed.
We have previously described a perfluoroaryl-cysteine SNAr approach for peptide and protein modifications46,47,48,49. The reactions between perfluoroaryl groups and cysteine residues are fast in organic solvent, but extremely sluggish in water unless an enzyme is used47,48. This observation inspired us to develop small peptides to promote the SNAr reaction in an analogous fashion to enzymes.
Results
Design of the π-clamp
Here we describe the identification of the π-clamp sequence to mediate site-specific cysteine modification in water without an enzyme, which overcomes the selectivity challenge for cysteine bioconjugation (Fig. 1c). This offers a new mode for site-specific chemistry by fine-tuning the microenvironment of a four-residue stretch within a complex protein or peptide.
We serendipitously discovered the π-clamp via a library selection approach designed to identify peptide sequences that promote arylation reactions in water. To accomplish this, we prepared a peptide library (Zaa-Cys-Zaa-Zaa-Gly-Leu-Leu-Lys, where Zaa is any one of the 20 natural amino acids except cysteine) and reacted the library with a biotin-perfluoroaryl probe in solution (Supplementary Fig. 26). Following streptavidin pull-down and liquid chromatography–tandem mass spectrometry (LC-MS/MS) analysis to identify reaction products (Supplementary Section 3), we found that the sequence Phe-Cys-Pro-Trp reacted with the perfluoroaryl-cysteine moiety in water (Supplementary Fig. 1). This observation is in stark contrast to our earlier efforts47, which showed that cysteine residues and perfluoroaryl moieties do not react in water. The Phe-Cys-Pro-Trp sequence thus appears to modify the reactivity of the cysteine thiol. Further mutating the Phe and Trp to Gly eliminated the reaction. Based on these findings and a molecular model of Phe-Cys-Pro-Trp, we hypothesize that the Phe and Trp side chains activate the cysteine thiol and interact with the incoming perfluoroaryl group, while the Pro serves to position the Cys, Phe and Trp residues into a conformation that promotes the reaction. We refer to this distinctive amino-acid sequence Xaa-Cys-Pro-Xaa (Xaa = electron-rich aromatic amino acids including Phe, Trp or Tyr) as a π-clamp.
Studies of the π-clamp-mediated conjugation
To investigate the π-clamp-mediated conjugation, we mutated the aromatic residues. Each of nine peptides (Xaa-Cys-Pro-Xaa-Gly-Leu-Leu-Lys-Asn-Lys, where Xaa was Phe, Trp, or Tyr) was tested for reaction with a perfluoroaryl-probe (2) in 0.2 M phosphate buffer at pH 8.0 and 37 °C with 20 mM tris(2-carboxylethyl)phosphine (TCEP) added as the reducing agent. All nine peptides reacted with probe 2 (rate constants of 0.076–0.73 M−1 s−1, Supplementary Table 2). In contrast, the double glycine mutant (1A) formed no product (Table 1, entry 1). The Phe-Phe π-clamp peptide (1E) gave quantitative conversion in 30 min (rate constants = 0.73 M−1 s−1, Table 1, entry 5). Single mutations of each Phe to Gly (1B and 1C, Table 1, entries 2 and 3) or converting the L-Pro to D-Pro (1D, Table 1, entry 4) significantly decreased the rate of the arylation reaction. These studies indicate that each amino acid in the π-clamp is essential for product formation.
π-Clamp mediated conjugation is highly selective, as indicated by our thiol competition experiments. The π-clamp peptide 1E was found to undergo quantitative conversion with perfluoroaryl probe 2 in the presence of a double glycine mutant peptide (1A) that served as the competing thiol species. Only the π-clamp peptide reacted quantitatively to form conjugated product in 30 min (Fig. 2).
Site-specific conjugation at the π-clamp in the presence of another competing cysteine peptide. π-Clamp peptide 1E was fully converted to the arylated product 2E while a competing cysteine peptide 1A remained unmodified. Chromatograms shown are total ion currents (TIC) from LC-MS analysis of crude reaction mixtures at 0 min (black) and 30 min (red). The mass spectrum of product 2E is shown as the inset.
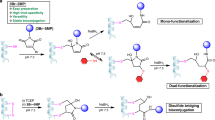
To further investigate the π-clamp-mediated cysteine conjugation, we carried out additional studies to understand whether location mattered, and also the substrate scope. We found that the π-clamp was efficiently modified irrespective of its position on the polypeptide chain (Fig. 3, Table 2 and Supplementary Figs 3–6). The π-clamp at the N terminus (1E), the C terminus (1N) and the middle (1O) of the polypeptide chain were readily modified with a diverse set of perfluoroaryl-linked probes including peptide, biotin, fluorescein, alkyne and polyethylene glycol (2–6).
We next investigated the regioselectivity on a 55 kDa protein substrate (Fig. 4a). Model protein 7 was designed to contain an N-terminal cysteine and a C-terminal π-clamp. A protease cleavage site was positioned upstream of the π-clamp, thereby allowing for unequivocal verification of the regioselectivity. On reacting protein 7 with probe 2 for 2 h, we observed >95% formation of the mono-labelled product 7A. The N-terminal free cysteine was subsequently labelled with fluorescein-5-maleimide, producing the dual-labelled product 7B. On protease cleavage, only two products were generated: a protein with maleimide-labelled N-terminal cysteine (7C) and a π-clamp arylated species, confirming the absolute regioselectivity endowed by the π-clamp.
a, Protecting-group-free one-pot dual labelling of a 55 kDa protein. The protein used was a fusion protein of the anthrax toxin lethal factor 1–263 (LFN) and diphtheria toxin domain A (DTA) with an engineered N-terminal cysteine, a C-terminal π-clamp and a protease cleavage site upstream of the π-clamp. After sequential modifications of the two cysteines with perfluoroaryl probe 2 and fluorescein-5-maleimide, dual-labelled product 7B was protease-digested to illustrate the selectivity of the π-clamp-mediated conjugation. Reaction conditions: (i) 50 µM 7, 1 mM 2, 0.2 M phosphate, 20 mM TCEP, 37 °C, 2 h; (ii) 50 µM 7A, 1 mM fluorescein-5-maleimide, 0.2 M phosphate pH 7.0, room temperature, 10 min; (iii) 25 µM protein 7B, 0.1 mg ml–1 TEV protease, 50 mM Tris, 0.1 mM EDTA, 1 mM DTT, pH 8.0, room temperature, 15 h. TEV, tobacco itch virus; EDTA, ethylenediaminetetraacetic acid; DTT, dithiothreitol; Tris, 2-amino-2-hydroxylmethyl-propane-1,3-diol. b, Top: quantitative and selective labelling of π-clamp SrtA (PDB entry: 1T2P). Bottom: control shows no labelling of SrtA. Reaction conditions: 38 µM 8 or 9, 1 mM 2, 0.2 M phosphate, 20 mM TCEP, 37 °C, 6 h.
Next, we site-specifically modified a cysteine-containing transpeptidase, Sortase A (SrtA)31 (Fig. 4b). An N-terminal π-clamp SrtA variant (8) reacted with probe 2 to produce >95% mono-labelled product 8A. The modified variant displayed full catalytic activity (Supplementary Fig. 10). No reaction took place with SrtA without the π-clamp (9). In sharp contrast, when the π-clamp Sortase (8) was reacted with bromoacetamide, a mixture of products was produced, with labelling of both cysteine residues (Supplementary Fig. 9).
Synthesis of site-specific antibody–drug conjugates
IgG molecules modified with small-molecule drugs (antibody–drug conjugates, ADCs) are currently used as therapeutic agents50. However, attaching small-molecule agents site-specifically to cysteines in IgGs is impossible so far, so commercial ADCs are heterogeneous mixtures of conjugates50. Approaches to engineering cysteine substitutions in antibodies produce mixed disulfides with cysteine or glutathione, so a fine-tuned reduction–oxidation protocol must be used to afford the free cysteine thiols for selective drug conjugation in the presence of disulfide bonds51,52.
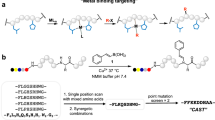
We anticipated that the π-clamp IgG could be used to overcome this specificity problem in ADC synthesis, which is notably challenging because IgGs harbour 32 native cysteine residues. The π-clamp-mediated modification on antibodies will be a single-step and site-specific antibody–drug conjugation technology that does not require significant antibody engineering or extra chemical steps51,52. To this end, we inserted the Phe-Cys-Pro-Phe sequence into the C termini of the heavy chains of trastuzumab53. Reacting the π-clamp trastuzumab (protein 10) with either a biotin–perfluoroaryl probe (11-biotin) or a drug–perfluoroaryl probe (11-MMAF) under reducing conditions, we observed facile formation of the heavy-chain mono-labelled products (10-biotin or 10-MMAF) by LC-MS analysis (Fig. 5a). Antibodies without the π-clamp showed no desired modification under the same conditions (Supplementary Fig. 27), highlighting the specificity of the conjugation. Moreover, this selective conjugation reaction works with other antibodies; reacting a π-clamp C225 antibody54,55 with 11-Biotin resulted in only the selective modifications on the π-clamp cysteine residues (Supplementary Fig. 28), suggesting that the π-clamp could be a general strategy for site-selective antibody modification.
a, Site-specific conjugation of biotin or monomethyl auristatin F (MMAF) to π-clamp trastuzumab (protein 10). LC-MS analysis showed site-specific labelling of the π-clamp cysteine residues on the trastuzumab heavy chain. The antibodies were treated with PNGase F to remove the N-linked glycans before LC-MS analysis. The deconvoluted mass spectra of the light chain (top) and the deglycosylated heavy chain (bottom) of π-clamp trastuzumab (10, left), the biotin labelled π-clamp trastuzumab (10-Biotin, centre) and the drug-conjugated π-clamp trastuzumab (10-MMAF, right) are shown. b, Biotin-conjugated π-clamp trastuzumab (10-Biotin) binds to HER2 in the Octet binding assay (KD = 0.2 ± 0.2 nM). The concentration of recombinant HER2 in each experiment is shown next to the curve (see Supplementary Section 9 for details). c, 10-Biotin retained binding to BT474 cells (HER2 positive) compared to the controls. Cells were treated with 10-Biotin or controls, washed with PBS with 0.1% BSA and then treated with streptavidin-AlexaFluor-647 before analysis by flow cytometer. 10-(PEG)4-Biotin and trastuzumab-(PEG)4-Biotin were prepared by reacting biotin-(PEG)4-NHS with protein 10 or trastuzumab, respectively (see Supplementary Section 3 and Fig. 36 for details). d, 10-MMAF killed BT474 cells (HER2 positive) but was not effective against CHO cells (HER2 negative). EC50 values for BT474 cells were 0.19 nM for 10-MMAF and 41 nM for auristatin F. The EC50 value of auristatin F for CHO cells is 1.3 µM. Cell viability was quantified using a CellTiter Glo assay and was normalized to cell only (see Methods for details). Experiments were performed in triplicate for each dose. Error bars indicate the standard deviation from the average of three experiments.
Under the developed reaction conditions (0.2 M phosphate, 20 mM TCEP, pH 8.0, at 37 °C), only the inter-chain disulfides and the π-clamp cysteine residues are reduced (Supplementary Fig. 38), and the modified antibodies retained binding affinity to their targets. Biotin-modified π-clamp trastuzumab (10-biotin) showed a similar binding affinity to HER2 (KD = 0.2 ± 0.2 nM) when compared to native trastuzumab non-selectively modified with a (PEG)4-biotin (trastuzumab-(PEG)4-biotin, KD = 0.3 ± 0.1 nM) (Fig. 5b and Supplementary Fig. 31). In addition, both proteins 10 and 10-biotin readily bound to BT474 cells (HER2-positive) (Fig. 5c; Supplementary Figs 32 and 33). As another antibody test case, biotin-modified C225 antibody (12-biotin) showed similar binding to A431 cells (EGFR-positive) compared to the native C225 antibody (Supplementary Figs 34 and 35). Collectively, insertion of the π-clamp into the heavy chains of antibodies and subsequent modification with probes did not significantly alter the binding properties.
Using the π-clamp-mediated cysteine conjugation, we synthesized a site-specific antibody–drug conjugate using π-clamp trastuzumab (protein 10) and a monomethyl auristatin F (MMAF) linked to a perfluoroaryl group (11-MMAF, see Supplementary Information for synthesis). LC-MS analysis of the conjugation reaction showed selective labelling of the heavy chain π-clamp cysteine residues (Fig. 5a). The prepared ADC selectively killed BT474 cells (HER2 positive) but was not effective for CHO cells (HER2 negative), indicating that the observed toxicity is receptor-dependent.
Mechanism of the π-clamp-mediated conjugation
To investigate the mechanism of the π-clamp-mediated reaction, we first used molecular dynamics (MD) to sample the conformational arrangements of the π-clamp peptide (1E) (Fig. 6a). Simulations indicated that 1E adopts four primary conformations when a cis-Pro is present: a ‘π-clamp’ (S1) with the phenyl rings of Phe-1 and Phe-4 interacting face-on with the Cys-2 thiol; a ‘half-clamp’ (S2) where only the Phe-4 side chain interacts with the Cys-2 thiol; S3 in which the Phe-1 and Phe-4 side chains are stacked together, leaving the Cys-2 thiol exposed; and an open configuration (S4) where all side chains are too far apart to interact. An MD simulation for π-clamp peptide 1E with a trans-Pro indicated two ‘open’ structures with the cysteine thiol not interacting with a Phe residue and one structure with a Phe-4 side chain interacting with a Cys-2 thiol (Supplementary Fig. 37).
a, Four primary structures S1–S4 were identified from MD simulation of π-clamp peptide 1E. The phenyl rings and cysteine thiol are shown as spheres, and the rest of the peptide is drawn as sticks. Hydrogen atoms are omitted for the sake of clarity. The heat map was plotted by summarizing all the obtained structures according to the distances between the sulfur atom and the centres of the phenyl rings. b, Conjugation to the π-clamp is energetically favoured over the double glycine mutant. Left: proposed nucleophilic aromatic substitution pathway for arylation at the π-clamp. Right: computed geometries and free energy surface of the nucleophilic aromatic substitution at the π-clamp (red). The free energy surface of the double glycine control is also shown (grey).
With these MD structures in hand, we used density functional theory (DFT) to investigate the nucleophilic aromatic substitution energy pathway for structures with a cis-Pro. We found that the half-clamp structure S2 stabilized the arylation product by ∼5 kcal mol–1 compared to the double glycine mutant, indicating the important role of Phe-4 in promoting the arylation reaction. This is consistent with our mutation studies showing that Phe-4 alone can partially mediate the arylation reaction (Table 1, entry 3). The product generated from the open structure (S4) has a similar free energy to that of the double glycine mutant, further substantiating the hypothesis that the two phenylalanine side chains are important for the arylation reaction with perfluoroaryl groups.
The most stable product was observed with the π-clamp structure S1, for which the free energy was ∼7 kcal mol–1 lower than that of the double glycine mutant. We further found that the activation energy for the formation of the transition state56 (III in Fig. 6b) was decreased by ∼3 kcal mol–1 when the π-clamp (S1) was present (see further discussion in Supplementary Sections 11–12), presumably because of the phenyl rings recognizing the perfluoroaryl group and activating the cysteine sulfur before conjugation. Collectively, these DFT calculations indicated that the π-clamp offers both a kinetic advantage (lower activation energy) and a thermodynamic advantage (lower free energy) over the double glycine mutant for the selective reaction with the perfluoroaryl reagents.
Discussion and conclusion
Here, we have described the discovery of a π-clamp to mediate site-selective cysteine conjugation. The π-clamp is composed of natural amino acids and shares some essential features of large enzymes, yet it mediates a purely abiotic cysteine perfluoroarylation reaction. The π-clamp tunes the reactivity of a cysteine thiol in its ‘active site’, recognizes the perfluoroaromatic reaction partner and decreases the activation energy for the reaction. In addition, the π-clamp has practical applications in protein labelling4. The reported reaction is site-specific, operational under physiologically relevant conditions, enzyme-free, and as efficient as the commonly used azide–alkyne click chemistry57,58 (π-clamp rate constant, 0.73 M−1 s−1).
Compared to existing bioconjugation techniques38, the advantages of the π-clamp include (1) its small size, which offers minimal structural perturbation to the target protein; (2) its genetic encodability for straightforward incorporation; (3) its ability to perform protecting-group-free dual cysteine modification; and (4) its reaction mode, which tunes the kinetic parameters to favour the cysteine perfluoroarylation reaction. This mode of reaction is distinct when compared to other advanced cysteine bioconjugations that use entropy to favour conjugation40,41,42.
The unexpected mode of site specificity provided by the π-clamp requires further mention. In all existing conjugation methods38, selectivity results from the judicious choice of certain functional groups so that each reaction pair undergoes conjugation in the presence of many other potentially reactive groups. For example, the unnatural handles used for click reactions are orthogonal to other functional groups on the target of interest12. In contrast, selectivity in π-clamp-mediated conjugation is achieved by fine-tuning the local chemical environment and reactivity, as proteins do. This provides a complementary strategy to non-natural amino-acid-mediated bioconjugation59. By fine-tuning the peptide microenvironment to allow for selective modification, the π-clamp significantly expands the chemistry available for selectively tailoring biomolecules.
Methods
Labelling of antibodies
Antibody conjugation reactions were performed on a 20 µl scale in polymerase chain reaction (PCR) tubes. A volume (8.33 µl) of π-clamp trastuzumab (10) stock solution (240 µM in PBS) was mixed with 2 µl reaction buffer (2 M phosphate, 200 mM TCEP, pH 8.0), 1 µl labelling biotin probe (11-Biotin, 20 mM in water) or drug probe (11-MMAF, 20 mM in DMSO) and 8.67 µl water. The reaction mixture was pipetted up and down for 20 times and was incubated in PCR machine at 37 °C. The final reaction conditions for biotin conjugation were 100 µM 10, 1 mM 11-Biotin, 0.2 M phosphate, 20 mM TCEP, 37 °C, 4 h, and the final reaction conditions for MMAF conjugation were 100 µM 10, 1 mM 11-MMAF, 0.2 M phosphate, 20 mM TCEP, 5% DMSO, 37 °C, 16 h. After reaction, the reaction mixture was diluted into 4 ml PBS and was buffer exchanged five times with PBS using 15 ml 10 K spin concentrator (EMD Millipore) to remove the excess labelling reagents. The final concentrated samples were used in LC-MS analysis, the Octet binding assay and the cell viability assay.
Cell viability assay
Cells were seeded in a 96-well white opaque plate at a density of 5 × 103/well (CHO) or 10 × 103/well (BT474). Cells were allowed to attach for 24 h at 37 °C and 5% CO2 in humidified atmosphere. Cells were then treated with serial dilutions of auristatin F, 10-MMAF or 10 for 96 h (BT474) or 72 h (CHO, treatment time was shortened to prevent overgrowth). The viability of cells was measured using CellTiter Glo reagents following the manufacturer's protocol and was normalized to the viability of cells without any treatment. The data were plotted using OriginLab software, and the half-maximal effective concentration (EC50) values were obtained by fitting the viability curves with a sigmoidal Boltzmann fit (see Supplementary Information).
Accession codes
References
Carrico, I. S. Chemoselective modification of proteins: hitting the target. Chem. Soc. Rev. 37, 1423–1431 (2008).
Hackenberger, C. P. R. & Schwarzer, D. Chemoselective ligation and modification strategies for peptides and proteins. Angew. Chem. Int. Ed. 47, 10030–10074 (2008).
Rabuka, D. Chemoenzymatic methods for site-specific protein modification. Curr. Opin. Chem. Biol. 14, 790–796 (2010).
Spicer, C. D. & Davis, B. G. Selective chemical protein modification. Nature Commun. 5, 4740 (2014).
Editorial. Take aim. Nature Chem. 4, 955 (2012).
Panowski, S., Bhakta, S., Raab, H., Polakis, P. & Junutula, J. R. Site-specific antibody drug conjugates for cancer therapy. MAbs 6, 34–45 (2014).
Kochendoerfer, G. G. Site-specific polymer modification of therapeutic proteins. Curr. Opin. Chem. Biol. 9, 555–560 (2005).
Uttamapinant, C. et al. A fluorophore ligase for site-specific protein labeling inside living cells. Proc. Natl Acad. Sci. USA 107, 10914–10919 (2010).
Krishna, O. D. & Kiick, K. L. Protein- and peptide-modified synthetic polymeric biomaterials. Peptide Sci. 94, 32–48 (2010).
Lichtor, P. A. & Miller, S. J. Combinatorial evolution of site- and enantioselective catalysts for polyene epoxidation. Nature Chem. 4, 990–995 (2012).
Wilcock, B. C. et al. Electronic tuning of site-selectivity. Nature Chem. 4, 996–1003 (2012).
Sletten, E. M. & Bertozzi, C. R. Bioorthogonal chemistry: fishing for selectivity in a sea of functionality. Angew. Chem. Int. Ed. 48, 6974–6998 (2009).
Afagh, N. A. & Yudin, A. K. Chemoselectivity and the curious reactivity preferences of functional groups. Angew. Chem. Int. Ed. 49, 262–310 (2010).
Lewis, C. A. & Miller, S. J. Site-selective derivatization and remodeling of erythromycin A by using simple peptide-based chiral catalysts. Angew. Chem. Int. Ed. 45, 5616–5619 (2006).
Chen, M. S. & White, M. C. A predictably selective aliphatic C–H oxidation reaction for complex molecule synthesis. Science 318, 783–787 (2007).
Snyder, S. A., Gollner, A. & Chiriac, M. I. Regioselective reactions for programmable resveratrol oligomer synthesis. Nature 474, 461–466 (2011).
Pathak, T. P. & Miller, S. J. Site-selective bromination of vancomycin. J. Am. Chem. Soc. 134, 6120–6123 (2012).
Wender, P. A., Hilinski, M. K. & Mayweg, A. V. W. Late-stage intermolecular CH activation for lead diversification: a highly chemoselective oxyfunctionalization of the C-9 position of potent bryostatin analogues. Org. Lett. 7, 79–82 (2005).
Peddibhotla, S., Dang, Y., Liu, J. O. & Romo, D. Simultaneous arming and structure/activity studies of natural products employing O–H insertions: an expedient and versatile strategy for natural products-based chemical genetics. J. Am. Chem. Soc. 129, 12222–12231 (2007).
Saxon, E. & Bertozzi, C. R. Cell surface engineering by a modified Staudinger reaction. Science 287, 2007–2010 (2000).
Rostovtsev, V. V., Green, L. G., Fokin, V. V. & Sharpless, K. B. A stepwise Huisgen cycloaddition process: copper(I)-catalyzed regioselective ‘ligation’ of azides and terminal alkynes. Angew. Chem. Int. Ed. 41, 2596–2599 (2002).
Agard, N. J., Prescher, J. A. & Bertozzi, C. R. A strain-promoted [3+2] azide−alkyne cycloaddition for covalent modification of biomolecules in living systems. J. Am. Chem. Soc. 126, 15046–15047 (2004).
Song, W., Wang, Y., Qu, J. & Lin, Q. Selective functionalization of a genetically encoded alkene-containing protein via ‘photoclick chemistry’ in bacterial cells. J. Am. Chem. Soc. 130, 9654–9655 (2008).
Blackman, M. L., Royzen, M. & Fox, J. M. Tetrazine ligation: fast bioconjugation based on inverse-electron-demand Diels–Alder reactivity. J. Am. Chem. Soc. 130, 13518–13519 (2008).
Saxon, E., Armstrong, J. I. & Bertozzi, C. R. A ‘traceless’ Staudinger ligation for the chemoselective synthesis of amide bonds. Org. Lett. 2, 2141–2143 (2000).
Lin, C.-W. & Ting, A. Y. Transglutaminase-catalyzed site-specific conjugation of small-molecule probes to proteins in vitro and on the surface of living cells. J. Am. Chem. Soc. 128, 4542–4543 (2006).
Wollack, J. W., Silverman, J. M., Petzold, C. J., Mougous, J. D. & Distefano, M. D. A minimalist substrate for enzymatic peptide and protein conjugation. ChemBioChem 10, 2934–2943 (2009).
Yin, J. et al. Genetically encoded short peptide tag for versatile protein labeling by Sfp phosphopantetheinyl transferase. Proc. Natl Acad. Sci. USA 102, 15815–15820 (2005).
Cull, M. G. & Schatz, P. J. Biotinylation of proteins in vivo and in vitro using small peptide tags. Methods Enzymol. 326, 430–440 (2000).
Fernandez-Suarez, M. et al. Redirecting lipoic acid ligase for cell surface protein labeling with small-molecule probes. Nature Biotechnol. 25, 1483–1487 (2007).
Popp, M. W., Antos, J. M., Grotenbreg, G. M., Spooner, E. & Ploegh, H. L. Sortagging: a versatile method for protein labeling. Nature Chem. Biol. 3, 707–708 (2007).
Whitford, D. Proteins: Structure and Function (Wiley, 2005).
Walsh, C. Enabling the chemistry of life. Nature 409, 226–231 (2001).
Giles, N. M., Giles, G. I. & Jacob, C. Multiple roles of cysteine in biocatalysis. Biochem. Biophys. Res. Commun. 300, 1–4 (2003).
Weerapana, E. et al. Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature 468, 790–795 (2010).
Nathani, R. I. et al. A novel approach to the site-selective dual labelling of a protein via chemoselective cysteine modification. Chem. Sci. 4, 3455–3458 (2013).
Chalker, J. M., Bernardes, G. J. L., Lin, Y. A. & Davis, B. G. Chemical modification of proteins at cysteine: opportunities in chemistry and biology. Chem. Asian J. 4, 630–640 (2009).
Hermanson, G. T. in Bioconjugate Techniques 3rd edn (ed Hermanson, G. T.) 1–125 (Academic, 2013).
Sun, M. M. C. et al. Reduction–alkylation strategies for the modification of specific monoclonal antibody disulfides. Bioconjug. Chem. 16, 1282–1290 (2005).
Griffin, B. A., Adams, S. R. & Tsien, R. Y. Specific covalent labeling of recombinant protein molecules inside live cells. Science 281, 269–272 (1998).
Adams, S. R. et al. New biarsenical ligands and tetracysteine motifs for protein labeling in vitro and in vivo: synthesis and biological applications. J. Am. Chem. Soc. 124, 6063–6076 (2002).
Wilson, P. et al. Organic arsenicals as efficient and highly specific linkers for protein/peptide–polymer conjugation. J. Am. Chem. Soc. 137, 4215–4222 (2015).
Stroffekova, K., Proenza, C. & Beam, K. G. The protein-labeling reagent FLASH-EDT2 binds not only to CCXXCC motifs but also non-specifically to endogenous cysteine-rich proteins. Pflügers Arch. 442, 859–866 (2001).
Gautier, A. et al. An engineered protein tag for multiprotein labeling in living cells. Chem. Biol. 15, 128–136 (2008).
Rush, J. S. & Bertozzi, C. R. New aldehyde tag sequences identified by screening formylglycine generating enzymes in vitro and in vivo. J. Am. Chem. Soc. 130, 12240–12241 (2008).
Spokoyny, A. M. et al. A perfluoroaryl-cysteine SNAr chemistry approach to unprotected peptide stapling. J. Am. Chem. Soc. 135, 5946–5949 (2013).
Zhang, C., Spokoyny, A. M., Zou, Y., Simon, M. D. & Pentelute, B. L. Enzymatic ‘click’ ligation: selective cysteine modification in polypeptides enabled by promiscuous glutathione S-transferase. Angew. Chem. Int. Ed. 52, 14001–14005 (2013).
Zhang, C., Dai, P., Spokoyny, A. M. & Pentelute, B. L. Enzyme-catalyzed macrocyclization of long unprotected peptides. Org. Lett. 16, 3652–3655 (2014).
Zou, Y. et al. Convergent diversity-oriented side-chain macrocyclization scan for unprotected polypeptides. Org. Biomol. Chem. 12, 566–573 (2014).
Sievers, E. L. & Senter, P. D. Antibody–drug conjugates in cancer therapy. Annu. Rev. Med. 64, 15–29 (2013).
Junutula, J. R. et al. Site-specific conjugation of a cytotoxic drug to an antibody improves the therapeutic index. Nature Biotechnol. 26, 925–932 (2008).
Junutula, J. R. et al. Rapid identification of reactive cysteine residues for site-specific labeling of antibody-Fabs. J. Immunol. Methods 332, 41–52 (2008).
Piccart-Gebhart, M. J. et al. Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N. Engl. J. Med. 353, 1659–1672 (2005).
Spangler, J. B., Manzari, M. T., Rosalia, E. K., Chen, T. F. & Wittrup, K. D. Triepitopic antibody fusions inhibit cetuximab-resistant BRAF and KRAS mutant tumors via EGFR signal repression. J. Mol. Biol. 422, 532–544 (2012).
Perrotte, P. et al. Anti-epidermal growth factor receptor antibody C225 inhibits angiogenesis in human transitional cell carcinoma growing orthotopically in nude mice. Clin. Cancer Res. 5, 257–265 (1999).
Artamkina, G. A., Egorov, M. P. & Beletskaya, I. P. Some aspects of anionic σ complexes. Chem. Rev. 82, 427–459 (1982).
Kolb, H. C., Finn, M. G. & Sharpless, K. B. Click chemistry: diverse chemical function from a few good reactions. Angew. Chem. Int. Ed. 40, 2004–2021 (2001).
Jewett, J. C. & Bertozzi, C. R. Cu-free click cycloaddition reactions in chemical biology. Chem. Soc. Rev. 39, 1272–1279 (2010).
Lang, K. & Chin, J. W. Cellular incorporation of unnatural amino acids and bioorthogonal labeling of proteins. Chem. Rev. 114, 4764–4806 (2014).
Acknowledgements
The authors thank S. Buchwald (Massachusetts Institute of Technology, MIT) and K. Dane Wittrup (MIT) for comments and suggestions regarding the manuscript. This work was supported by an MIT start-up fund, the National Institutes of Health (NIH; R01GM110535) and the Sontag Foundation Distinguished Scientist Award (to B.L.P.). C.Z. is a recipient of the George Büchi Research Fellowship, the Koch Graduate Fellowship in Cancer Research of MIT, and the Bristol-Myers Squibb Graduate Fellowship in Synthetic Organic Chemistry. M.W. is a National Science Foundation Graduate Research Fellow. The authors acknowledge the Biological Instrument Facility of MIT for providing the Octet BioLayer Interferometry System (NIH S10 OD016326), the MIT Center for Environmental Health Sciences (CEHS) Instrument Facility for providing the flow cytometer (P30-ES002109) and R.J. Collier (Harvard) for contributing laboratory equipment used in this study.
Author information
Authors and Affiliations
Contributions
C.Z. and B.L.P. conceived the work and designed the experiments. C.Z. performed the peptide and protein labelling experiments. M.W., T.Z. and T.V.H. performed the computational studies. N.J.Y. helped with antibody expression and purification. M.S.S. and C.Z. performed the flow cytometer and cell viability assays. C.Z. and B.L.P. wrote the manuscript, with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
A patent application covering this work has been filed by MIT-TLO (US patent application no. 14/278060, International patent application no. PCT/US15/031082).
Supplementary information
Supplementary information
Supplementary information (PDF 4297 kb)
Rights and permissions
About this article
Cite this article
Zhang, C., Welborn, M., Zhu, T. et al. π-Clamp-mediated cysteine conjugation. Nature Chem 8, 120–128 (2016). https://doi.org/10.1038/nchem.2413
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2413
This article is cited by
-
Copper assisted sequence-specific chemical protein conjugation at a single backbone amide
Nature Communications (2023)
-
Site-selective chemical reactions by on-water surface sequential assembly
Nature Communications (2023)
-
Recent Advances in Bioorthogonal Ligation and Bioconjugation
Topics in Current Chemistry (2023)
-
Site-selective photocatalytic functionalization of peptides and proteins at selenocysteine
Nature Communications (2022)
-
Laboratory evolution of a sortase enzyme that modifies amyloid-β protein
Nature Chemical Biology (2021)