Abstract
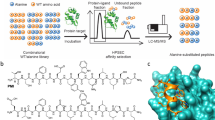
Ligands that can bind selectively to proteins with single amino-acid point mutations offer the potential to detect or treat an abnormal protein in the presence of the wild type (WT). However, it is difficult to develop a selective ligand if the point mutation is not associated with an addressable location, such as a binding pocket. Here we report an all-chemical synthetic epitope-targeting strategy that we used to discover a 5-mer peptide with selectivity for the E17K-transforming point mutation in the pleckstrin homology domain of the Akt1 oncoprotein. A fragment of Akt1 that contained the E17K mutation and an I19[propargylglycine] substitution was synthesized to form an addressable synthetic epitope. Azide-presenting peptides that clicked covalently onto this alkyne-presenting epitope were selected from a library using in situ screening. One peptide exhibits a 10:1 in vitro selectivity for the oncoprotein relative to the WT, with a similar selectivity in cells. This 5-mer peptide was expanded into a larger ligand that selectively blocks the E17K Akt1 interaction with its PIP3 (phosphatidylinositol (3,4,5)-trisphosphate) substrate.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Rusling, J. F., Kumar, C. V., Gutkind, J. S. & Patel, V. Measurement of biomarker proteins for point-of-care early detection and monitoring of cancer. Analyst 135, 2496–2511 (2010).
Chong, C. R. & Janne, P. A. The quest to overcome resistance to EGFR-targeted therapies in cancer. Nature Med. 19, 1389–1400 (2013).
Tjin Tham Sjin, R. et al. In vitro and in vivo characterization of irreversible mutant-selective EGFR inhibitors that are wild-type sparing. Mol. Cancer Ther. 13, 1468–1479 (2014).
Capper, D., Zentgraf, H., Balss, J., Hartmann, C. & von Deimling, A. Monoclonal antibody specific for IDH1 R132H mutation. Acta Neuropathol. 118, 599–601 (2009).
Yu, J. et al. Mutation-specific antibodies for the detection of EGFR mutations in non-small-cell lung cancer. Clin. Cancer Res. 15, 3023–3028 (2009).
Capper, D. et al. Assessment of BRAF V600E mutation status by immunohistochemistry with a mutation-specific monoclonal antibody. Acta Neuropathol. 122, 11–19 (2011).
Marschall, A. L. J., Frenzel, A., Schirrmann, T., Schüngel, M. & Dubel, S. Targeting antibodies to the cytoplasm. mAbs 3, 3–16 (2011).
Rondon, I. J., & Marasco, A. W. Intracellular antibodies (intrabodies) for gene therapy of infectious diseases. Ann. Rev. Microbiol. 51, 257–283 (1997).
Kodadek, T., Reddy, M. M., Olivos, H. J., Bachhawat-Sikder, K. & Alluri, P. G. Synthetic molecules as antibody replacements. Acc. Chem. Res. 37, 711–718 (2004).
Testa, J. R. & Tsichlis, P. N. AKT signaling in normal and malignant cells. Oncogene 24, 7391–7393 (2005).
Vivanco, I. & Sawyers, C. L. The phosphatidylinositol 3-kinase-AKT pathway in human cancer. Nature Rev. Cancer 2, 489–501 (2002).
Carpten, J. D. et al. A transforming mutation in the pleckstrin homology domain of AKT1 in cancer. Nature 448, 439–444 (2007).
Nag, A. et al. A chemical epitope-targeting strategy for protein capture agents: the serine 474 epitope of the kinase Akt2. Angew. Chem. Int. Ed. 52, 13975–13979 (2013).
Agnew, H. D. et al. Iterative in situ click chemistry creates antibody-like protein-capture agents. Angew. Chem. Int. Ed. 48, 4944–4948 (2009).
Farrow, B. et al. A chemically synthesized capture agent enables the selective, sensitive, and robust electrochemical detection of anthrax protective antigen. ACS Nano 7, 9452–9660 (2013).
Millward, S. W. et al. Iterative in situ click chemistry assembles a branched capture agent and allosteric inhibitor for Akt1. J. Am. Chem. Soc. 133, 18280–18288 (2011).
Pfeilsticker, J. A. et al. A cocktail of thermally stable, chemically synthesized capture agents for the efficient detection of anti-Gp41 antibodies from human sera. PLoS ONE 8, e76224 (2013).
Millward, S. W. et al. In situ click chemistry: from small molecule discovery to synthetic antibodies. Integr. Biol. 5, 87–95 (2013).
Coin, I., Beyermann, M. & Bienert, M. Solid-phase peptide synthesis: from standard procedures to the synthesis of difficult sequences. Nature Protocols 2, 3247–3256 (2007).
Pashkova, A., Moskovets, E. & Karger, B. L. Coumarin tags for improved analysis of peptides by MALDI-TOF MS and MS/MS. 1. Enhancement in MALDI MS signal intensities. Anal. Chem. 76, 4550–4557 (2004).
Tsukiji, S., Miyagawa, M., Takaoka, Y., Tamura, T. & Hamachi, I. Ligand-directed tosyl chemistry for protein labeling in vivo. Nature Chem. Biol. 5, 341–343 (2009).
Deshayes, S., Morris, M. C., Divita, G. & Heitz, F. Cell-penetrating peptides: tools for intracellular delivery of therapeutics. Cell. Mol. Life Sci. 62, 1839–1849 (2005).
Dunn, K. W., Kamocka, M. M. & McDonald, J. H. A practical guide to evaluating colocalization in biological microscopy. Am. J. Physiol. Cell Physiol. 300, C723–C742 (2011).
Mahadevan, D. et al. Discovery of a novel class of AKT pleckstrin homology domain inhibitors. Mol. Cancer Ther. 7, 2621–2632 (2008).
Hiromura, M. et al. Inhibition of Akt kinase activity by a peptide spanning the βA strand of the proto-oncogene TCL1. J. Biol. Chem. 279, 53407–53418 (2004).
Verdine, G. L. & Hilinski, G. J. Stapled peptides for intracellular drug targets. Methods Enzymol. 503, 3–33 (2012).
Lee, S. S. et al. Accurate MALDI-TOF/TOF sequencing of one-bead-one-compound peptide libraries with application to the identification of multiligand protein affinity agents using in situ click chemistry screening. Anal. Chem. 82, 672–679 (2010).
Acknowledgements
This work was supported by the Institute for Collaborative Biotechnologies through grant W911NF-09-0001 from the US Army Research Office, the National Cancer Institute through grant 5U54 CA119347, the Defense Advanced Research Projects Agency through Cooperative Agreement No. HR0011-11-2-0006 and the Jean Perkins Foundation. B.F. is supported by a Howard Hughes Medical Institute International Student Research Fellowship. We gratefully acknowledge assistance and resources from F. Rusnak, J. Zhou and the Protein and Peptide Mass Analysis Laboratory, M. Shahgholi and the Mass Spectrometry Lab, J. Vielmetter and the Protein Expression Center, and the Beckman Institute Biological Imaging Center.
Author information
Authors and Affiliations
Contributions
K.M.D., B.F. and J.R.H. designed the project and wrote the manuscript. K.M.D., B.F., Y.Q.H., J.W. and M.W. carried out the experiments. B.L. carried out the MS analysis. A.U. and S.W.M. designed and helped execute the protein expression and cell culture work. A.N. and S.D. helped develop the epitope-targeting strategies. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.R.H. is a founder and board member of Indi Molecular. Indi Molecular is seeking to commercialize the PCC-agent technology. B.L. is an employee of Indi Molecular. K.M.D. has consulted for Indi Molecular. The patent ‘Multi-ligand capture agents and related compositions, methods and systems’ (Patent WO2009155420 A1) by H. Agnew et al. was published 23 December 2009.
Supplementary information
Supplementary information
Supplementary information (PDF 7024 kb)
Rights and permissions
About this article
Cite this article
Deyle, K., Farrow, B., Qiao Hee, Y. et al. A protein-targeting strategy used to develop a selective inhibitor of the E17K point mutation in the PH domain of Akt1. Nature Chem 7, 455–462 (2015). https://doi.org/10.1038/nchem.2223
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2223
This article is cited by
-
Stereochemical engineering yields a multifunctional peptide macrocycle inhibitor of Akt2 by fine-tuning macrocycle-cell membrane interactions
Communications Chemistry (2023)
-
PH-domain-binding inhibitors of nucleotide exchange factor BRAG2 disrupt Arf GTPase signaling
Nature Chemical Biology (2019)
-
Activating Akt1 mutations alter DNA double strand break repair and radiosensitivity
Scientific Reports (2017)