Abstract
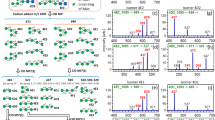
Mass spectrometry is the primary analytical technique used to characterize the complex oligosaccharides that decorate cell surfaces. Monosaccharide building blocks are often simple epimers, which when combined produce diastereomeric glycoconjugates indistinguishable by mass spectrometry. Structure elucidation frequently relies on assumptions that biosynthetic pathways are highly conserved. Here, we show that biosynthetic enzymes can display unexpected promiscuity, with human glycosyltransferase pp-α-GanT2 able to utilize both uridine diphosphate N-acetylglucosamine and uridine diphosphate N-acetylgalactosamine, leading to the synthesis of epimeric glycopeptides in vitro. Ion-mobility mass spectrometry (IM-MS) was used to separate these structures and, significantly, enabled characterization of the attached glycan based on the drift times of the monosaccharide product ions generated following collision-induced dissociation. Finally, ion-mobility mass spectrometry following fragmentation was used to determine the nature of both the reducing and non-reducing glycans of a series of epimeric disaccharides and the branched pentasaccharide Man3 glycan, demonstrating that this technique may prove useful for the sequencing of complex oligosaccharides.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
21 March 2014
After this Article went to press the authors realized that a number of the key references had been inadvertently omitted or removed before the final submission of the manuscript. The authors would therefore like to cite the following additional articles: 1. Zhu, M. L., Bendiak, B., Clowers, B. & Hill, H. H. Ion mobility-mass spectrometry analysis of isomeric carbohydrate precursor ions. Anal. Bioanal. Chem. 394, 1853–1867 (2009). Structural characterization of select isomeric oligosaccharides using atmospheric ion-mobility spectrometry for separation of linkage and branch isomers, anomeric isomers, and epimers, prior to MS3 analysis using an ion-trap mass spectrometer. 2. Williams, J. P. et al. Characterization of simple isomeric oligosaccharides and the rapid separation of glycan mixtures by ion mobility mass spectrometry. Int. J. Mass Spectrom. 298, 119–127 (2010). Using both travelling-wave ion-mobility spectrometry and drift-tube ion-mobility spectrometry, released N-glycans and isobaric glycans were separated for subsequent characterization by tandem MS. Theoretical modelling was also used to confirm experimentally determined collisional cross-section values. 3. Fenn, L. S. & McLean, J. A. Structural resolution of carbohydrate positional and structural isomers based on gas-phase ion mobilitymass spectrometry. Phys. Chem. Chem. Phys. 13, 2196–2205 (2011). Details the collisional cross-section values of ∼300 sodiated positional and structural carbohydrate isomers from MALDI IM-MS. 4. Harvey, D. J. et al. Travelling wave ion mobility and negative ion fragmentation for the structural determination of N-linked glycans. Electrophoresis 34, 2368–2378 (2013). Structural characterization of released N-glycans using negative-ion-mode collision-induced dissociation of ion-mobility-separated isomer (and conformer) precursors.
References
Hart, G. W. & Copeland, R. J. Glycomics hits the big time. Cell 143, 672–676 (2010).
Moremen, K. W., Tiemeyer, M. & Nairn, A. V. Vertebrate protein glycosylation: diversity, synthesis and function. Nature Rev. Mol. Cell Biol. 13, 448–462 (2012).
Hanisch, F-G. Top-down sequencing of O-glycoproteins by in-source decay matrix-assisted laser desorption ionisation mass spectrometry for glycosylation site analysis. Anal. Chem. 83, 4829–4837 (2011).
Kolarich, D., Lepenies, B. & Seeberger, P. H. Glycomics, glycoproteomics and the immune system. Curr. Opin. Chem. Biol. 16, 214–220 (2012).
Antonopoulos, A., North, S. J., Haslam, S. M. & Dell, A. Glycosylation of mouse and human immune cells: insights emerging from N-glycome analyses. Biochem. Soc. Trans. 39, 1334–1340 (2011).
Devakumar, A., Mechref, Y., Kang, P., Novotny, M. V. & Reilly J. P. Identification of isomeric N-glycan structures by mass spectrometry with 157 nm laser-induced photofragmentation. J. Am. Soc. Mass Spectrosc. 19, 1027–1040 (2008).
Adibekian, A. et al. Comparative bioinformatics analysis of the mammalian and bacterial glycomes. Chem. Sci. 2, 337–344 (2011).
Rini, J. M., Esko, J. D. & Varki, A. in Essentials of Glycobiology 2nd edn (eds Varki, A. et al.) Ch. 5 (Cold Spring Harbor Laboratory Press, 2009).
Gerken, T. A. et al. Emerging paradigms for the initiation of mucin-type protein O-glycosylation by the polypeptide GalNAc transferase family of glycosyltransferases. J. Biol. Chem. 286, 14493–14507 (2011).
Laurent, N. et al. SPOT synthesis of peptide arrays on self-assembled monolayers and their evaluation as enzyme substrates. ChemBioChem. 9, 2592–2596 (2008).
Laurent, N. et al. Enzymatic glycosylation of peptide arrays on gold surfaces. ChemBioChem. 9, 883–887 (2008).
Su, J. & Mrksich, M. Using mass spectrometry to characterize self-assembled monolayers presenting peptides, proteins, and carbohydrates. Angew. Chem. Int. Ed. 41, 4715–4718 (2002).
Errey, J. C., Mukhopadhyay, B., Kartha, R. & Field, R. A. Flexible enzymatic and chemo-enzymatic approaches to a broad range of uridine-diphospho-sugars. Chem. Commun. 2796–2707 (2004).
Rejzek, M. et al. Chemical synthesis of UDP-Glc-2,3-diNAcA, a key intermediate in cell surface polysaccharide biosynthesis in the human respiratory pathogens B. pertussis and P. aeruginosa. Org. Biomol. Chem. 7, 1203–1210 (2009).
Wandall, H. H. et al. Substrate specificities of three members of the human UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase family, GalNAc-T1, -T2, and -T3. J. Biol. Chem. 272, 23503–23514 (1997).
Ramakrishnan, B., Boeggeman, E. & Qasba P. K. Novel method for in vitro O-glycosylation of proteins: application for bioconjugation. Bioconjug. Chem. 18, 1912–1918 (2007).
Jung, E., Gooley, A. A., Packer N. H., Karuso P. & Williams, K. L. Rules for the addition of O-linked N-acetylglucosamine to secreted proteins in Dictyostelium discoideum—in vivo studies on glycosylation of mucin Muc 1 and Muc 2 repeats. Eur. J. Biochem. 253, 517–524 (1998).
Previato, J. O. et al. Biosynthesis of O-N-acetylglucosamine-linked glycans in Trypanosoma cruzi. J. Biol. Chem. 273, 14982–14988 (1998).
Thoden, J. B., Wohlers, T. M., Fridovich-Keil, J. L. & Holden, H. M. Human UDP-galactose 4-epimerase. Accommodation of UDP-N-acetylglucosamine within the active site. J. Biol. Chem. 276, 15131–15136 (2001).
Jansson, P. E., Kenne, L. & Widmalm, G. Computer-assisted structural analysis of polysaccharides with an extended version of CASPER using 1H- and 13C-N.M.R. data. Carbohydr. Res. 188, 169–191 (1989).
Lundborg, M. & Widmalm, G. Structural analysis of glycans by NMR chemical shift prediction. Anal. Chem. 83, 1514–1517 (2011).
Šardzík, R. et al. Chemoenzymatic synthesis of O-mannosylpeptides in solution and on solid phase. J. Am. Chem. Soc. 134, 4521–4524 (2012).
Gagneux, P. & Varki, A. Evolutionary considerations in relating oligosaccharide diversity to biological function. Glycobiology 9, 747–755 (1999).
Ercan, A. & West, C. M. Kinetic analysis of a golgi UDP-GlcNAc:polypeptide-Thr/Ser N-acetyl-α-glucosaminyltransferase from Dictyostelium. Glycobiology 15, 489–500 (2005).
Taniguchi, N., Honke, K. & Fukuda, M. (eds) in Handbook of Glycosyltransferases and Related Genes Chs 9–14 (Springer-Verlag, 2003).
Lau, K. S. et al. Complex N-glycan number and degree of branching cooperate to regulate cell proliferation and differentiation. Cell 129, 123–134 (2007).
Creese, A. J. & Cooper, H. J. Separation and identification of isomeric glycopeptides by high field asymmetric waveform ion mobility spectrometry. Anal. Chem. 84, 2597–2601 (2012).
Li, H. et al. Resolving structural isomers of monosaccharide methyl glycosides using drift tube and travelling wave ion mobility mass spectrometry. Anal. Chem. 84, 3231–3239 (2012).
Li, H., Bendiak, B., Siems, W. F., Gang, D. R. & Hill, H. H. Carbohydrate structure characterization by tandem ion mass spectroscopy (IMMS)2. Anal. Chem. 85, 2760–2769 (2013).
Pagel, K. & Harvey, D. J. Ion mobility mass spectrometry of complex carbohydrates—collision cross sections of sodiated N-linked glycans. Anal. Chem. 85, 5138–5145 (2013).
Slovin, S. F., Keding, S. J. & Ragupathi, G. Carbohydrate vaccines as immunotherapy for cancer. Immunol. Cell Biol. 83, 418–428 (2005).
Kumar, S. R., Sauter, E. R., Quinn, T. P. & Deutscher, S. L. Thomsen-Friedenreich and Tn antigens in nipple fluid: carbohydrate biomarkers for breast cancer detection. Clin. Cancer Res. 11, 6868–6871 (2005).
Hakomori, S. The glycosynapse. Proc. Natl Acad. Sci. USA 99, 225–232 (2002).
Zaia, J. Mass spectrometry of oligosaccharides. Mass Spectrom. Rev. 23, 161–227 (2004).
Borsdorf, H. & Eiceman, G. A. Ion mobility spectrometry: principles and applications. Appl. Spectrosc. Rev. 41, 323–375 (2006).
Chawner, R. et al. QconCAT standard for calibration of ion mobility-mass spectrometry systems. J. Proteome Res. 11, 5564–5572 (2012).
Wyttenbach, T., Bushnell, J. E. & Bowers, M. T. Salt bridge structures in the absence of solvent? The case for the oligoglycines. J. Am. Chem. Soc. 120, 5098–5103 (1998).
Hurtado-Guerrero, R., Dorfmueller, H. C. & van Aalten, D. M. Molecular mechanisms of O-GlcNAcylation. Curr. Opin. Struct. Biol. 18, 551–557 (2008).
Muller, R., Jenny, A. & Stanley, P. The EGF repeat-specific O-GlcNAc-transferase Eogt interacts with Notch signaling and pyrimidine metabolism pathways in Drosophila. PLoS ONE 8, e62835 (2013).
Domon, B. & Costello, C. E. Structure elucidation of glycosphingolipids and gangliosides using high-performance tandem mass spectrometry. Biochemistry 27, 1534–1543 (1988).
Acknowledgements
This research was supported by grants from the Engineering and Physical Sciences Research Council (EPSRC), the Biotechnology and Biological Sciences Research Council (BBSRC), Swiss Scheme Foundation (MA), the Royal Society (Wolfson Award to S.L.F.), the Knut and Alice Wallenberg Foundation, the Swedish Research Council and the European commission (FP7). Work at John Innes Centre (JIC) was supported by a BBSRC Institute Strategic Programme Grant (BB/J004561/1) and the John Innes Foundation.
Author information
Authors and Affiliations
Contributions
C.E.E., S.L.F., A.P.G., P.B. and C.J.G. conceived the project, designed the experiments, discussed the results and implications, and commented on the manuscript at all stages. A.P.G., C.J.G., P.B., S.L.F. and C.E.E. co-wrote the paper. P.B., A.P.G. and C.J.G. contributed equally. C.J.G. performed the IM-MS experiments and statistical analysis. P.B., J.V. and D.R. performed the molecular biology and protein purification. P.B. and A.P.G. carried out kinetic studies and bioinformatics. A.P.G. performed the glycan chemical synthesis. R.Š., J.V. and M.A. carried out peptide synthesis and biotransformations in the solid phase and in solution. C.F. and G.W. performed and reported the NMR analysis. M.R. and R.A.F. synthesized the activated sugar donors. All authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 3209 kb)
Rights and permissions
About this article
Cite this article
Both, P., Green, A., Gray, C. et al. Discrimination of epimeric glycans and glycopeptides using IM-MS and its potential for carbohydrate sequencing. Nature Chem 6, 65–74 (2014). https://doi.org/10.1038/nchem.1817
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.1817
This article is cited by
-
Glycoproteomics
Nature Reviews Methods Primers (2022)
-
Utilizing liquid chromatography, ion mobility spectrometry, and mass spectrometry to assess INLIGHT™ derivatized N-linked glycans in biological samples
Analytical and Bioanalytical Chemistry (2022)
-
Shotgun ion mobility mass spectrometry sequencing of heparan sulfate saccharides
Nature Communications (2020)
-
Comprehensive structural glycomic characterization of the glycocalyxes of cells and tissues
Nature Protocols (2020)
-
De novo structural determination of mannose oligosaccharides by using a logically derived sequence for tandem mass spectrometry
Analytical and Bioanalytical Chemistry (2019)