Abstract
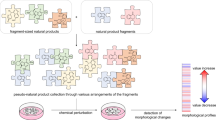
Fragment-based ligand and drug discovery predominantly employs sp2-rich compounds covering well-explored regions of chemical space. Despite the ease with which such fragments can be coupled, this focus on flat compounds is widely cited as contributing to the attrition rate of the drug discovery process. In contrast, biologically validated natural products are rich in stereogenic centres and populate areas of chemical space not occupied by average synthetic molecules. Here, we have analysed more than 180,000 natural product structures to arrive at 2,000 clusters of natural-product-derived fragments with high structural diversity, which resemble natural scaffolds and are rich in sp3-configured centres. The structures of the cluster centres differ from previously explored fragment libraries, but for nearly half of the clusters representative members are commercially available. We validate their usefulness for the discovery of novel ligand and inhibitor types by means of protein X-ray crystallography and the identification of novel stabilizers of inactive conformations of p38α MAP kinase and of inhibitors of several phosphatases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Murray, C. W. & Rees, D. C. The rise of fragment-based drug discovery. Nature Chem. 1, 187–192 (2009).
Roughley, S. D. & Hubbard, R. E. How well can fragments explore accessed chemical space? A case study from heat shock protein 90. J. Med. Chem. 54, 3989–4005 (2011).
Chessari, G. & Woodhead, A. J. From fragment to clinical candidate—a historical perspective. Drug Discov. Today 14, 668–675 (2009).
Babaoglu, K. & Shoichet, B. K. Deconstructing fragment-based inhibitor discovery. Nature Chem. Biol. 2, 720–723 (2006).
Hubbard, R. E., Chen, I. & Davis, B. Informatics and modeling challenges in fragment-based drug discovery. Curr. Opin. Drug Discov. Dev. 10, 289–297 (2007).
Siegel, M. G. & Vieth, M. Drugs in other drugs: a new look at drugs as fragments. Drug Discov. Today 12, 71–79 (2007).
Nicholls, A. et al. Molecular shape and medicinal chemistry: a perspective. J. Med. Chem. 53, 3862–3886 (2010).
Ritchie, T. J., Macdonald, S. J. F., Young, R. J. & Pickett, S. D. The impact of aromatic ring count on compound developability: further insights by examining carbo- and hetero-aromatic and -aliphatic ring types. Drug Discov. Today 16, 164–171 (2011).
Lovering, F., Bikker, J. & Humblet, C. Escape from flatland: increasing saturation as an approach to improving clinical success. J. Med. Chem. 52, 6752–6756 (2009).
Hung, A. W. et al. Route to three-dimensional fragments using diversity-oriented synthesis. Proc. Natl Acad. Sci. USA 108, 6799–6804 (2011).
Newman, D. J. & Cragg, G. M. Natural products as sources of new drugs over the 30 years from 1981 to 2010. J. Nat. Prod. 75, 311–335 (2012).
Wetzel, S., Schuffenhauer, A., Roggo, S., Ertl, P. & Waldmann, H. Cheminformatic analysis of natural products and their chemical space. Chimia Int. J. Chem. 61, 355–360 (2007).
Grabowski, K., Baringhaus, K-H. & Schneider, G. Scaffold diversity of natural products: inspiration for combinatorial library design. Nat. Prod. Rep. 25, 892–904 (2008).
Congreve, M., Carr, R., Murray, C. & Jhoti, H. A ‘Rule of Three’ for fragment-based lead discovery? Drug Discov. Today 8, 876–877 (2003).
Köster, H. et al. A small nonrule of 3 compatible fragment library provides high hit rate of endothiapepsin crystal structures with various fragment chemotypes. J. Med. Chem. 54, 7784–7796 (2011).
Wetzel, S., Bon, R. S., Kumar, K. & Waldmann, H. Biology-oriented synthesis. Angew. Chem. Int. Ed. 50, 10800–10826 (2011).
Bon, R. S. & Waldmann, H. Bioactivity-guided navigation of chemical space. Acc. Chem. Res. 43, 1103–1114 (2010).
Kumar, K. & Waldmann, H. Synthesis of natural product inspired compound collections. Angew. Chem. Int. Ed. 48, 3224–3242 (2009).
Schuffenhauer, A. et al. The scaffold tree; visualization of the scaffold universe by hierarchical scaffold classification. J. Chem. Inf. Model. 47, 47–58 (2007).
Koch, M. A. et al. Charting biologically relevant chemical space: a structural classification of natural products (SCONP). Proc. Natl Acad. Sci. USA 102, 17272–17277 (2005).
Dictionary of Natural Products Version 18.2, 2009 (Chapman & Hall/CRC Press, 2009).
Bemis, G. W. & Murcko, M. A. The properties of known drugs. 1. Molecular frameworks. J. Med. Chem. 39, 2887–2893 (1996).
Irwin, J. J., Sterling, T., Mysinger, M. M., Bolstad, E. S. & Coleman, R. G. ZINC: a free tool to discover chemistry for biology. J. Chem. Inf. Model. 52, 1757–1768 (2012).
Butina, D. Unsupervised data base clustering based on daylight's fingerprint and Tanimoto similarity: a fast and automated way to cluster small and large data sets. J. Chem. Inf. Comput. Sci. 39, 747–750 (1999).
Gill, A. L. et al. Identification of novel p38α MAP kinase inhibitors using fragment-based lead generation. J. Med. Chem. 48, 414–426 (2004).
Pollack, S. et al. A comparative study of fragment screening methods on the p38α kinase: new methods, new insights. J. Comput. Aided Molec. Des. 25, 677–687 (2011).
Goettert, M., Schattel, V., Koch, P., Merfort, I. & Laufer, S. Biological evaluation and structural determinants of p38α mitogen-activated-protein kinase and c-Jun-N-terminal kinase 3 inhibition by flavonoids. ChemBioChem 11, 2579–2588 (2010).
Dearden, M. J., McGrath, M. J. & O'Brien, P. Evaluation of (+)-sparteine-like diamines for asymmetric synthesis. J. Org. Chem. 69, 5789–5792 (2004).
Simard, J. R. et al. A new screening assay for allosteric inhibitors of cSrc. Nature Chem. Biol. 5, 394–396 (2009).
Rabiller, M. et al. Proteus in the world of proteins: conformational changes in protein kinases. Arch. Pharm. (Weinheim) 343, 193–206 (2010).
Ahn, Y. M. et al. Switch control pocket inhibitors of p38–MAP kinase. Durable type II inhibitors that do not require binding into the canonical ATP hinge region. Bioorg. Med. Chem. Lett. 20, 5793–5798 (2010).
Swann, S. L. et al. Biochemical and biophysical characterization of unique switch pocket inhibitors of p38α. Bioorg. Med. Chem. Lett. 20, 5787–5792 (2010).
Getlik, M. et al. Structure-based design, synthesis and biological evaluation of N-pyrazole, N′-thiazole urea inhibitors of MAP kinase p38α. Eur. J. Med. Chem. 48, 1–15 (2012).
Kaiser, M., Wetzel, S., Kumar, K. & Waldmann, H. Biology-inspired synthesis of compound libraries. Cell. Mol. Life Sci. 65, 1186–1201 (2008).
Vintonyak, V. V., Antonchick, A. P., Rauh, D. & Waldmann, H. The therapeutic potential of phosphatase inhibitors. Curr. Opin. Chem. Biol. 13, 272–283 (2009).
Vintonyak, V. V., Waldmann, H. & Rauh, D. Using small molecules to target protein phosphatases. Bioorg. Med. Chem. 19, 2145–2155 (2011).
Bialy, L. & Waldmann, H. Inhibitors of protein tyrosine phosphatases: next-generation drugs? Angew. Chem. Int. Ed. 44, 3814–3839 (2005).
Nören-Müller, A. et al. Discovery of protein phosphatase inhibitor classes by biology-oriented synthesis. Proc. Natl Acad. Sci. USA 103, 10606–10611 (2006).
Renner, S. et al. Bioactivity-guided mapping and navigation of chemical space. Nature Chem. Biol. 5, 585–592 (2009).
Hert, J., Irwin, J. J., Laggner, C., Keiser, M. J. & Shoichet, B. K. Quantifying biogenic bias in screening libraries. Nature Chem. Biol. 5, 479–483 (2009).
Hessler, G. & Baringhaus, K-H. The scaffold hopping potential of pharmacophores. Drug Discov. Today: Technologies 7, e263–e269 (2010).
Nisius, B. & Rester, U. Fragment shuffling: an automated workflow for three-dimensional fragment-based ligand design. J. Chem. Inf. Model. 49, 1211–1222 (2009).
Bamborough, P., Brown, M. J., Christopher, J. A., Chung, C-W. & Mellor, G. W. Selectivity of kinase inhibitor fragments. J. Med. Chem. 54, 5131–5143 (2011).
Varin, T., Schuffenhauer, A., Ertl, P. & Renner, S. Mining for bioactive scaffolds with scaffold networks: improved compound set enrichment from primary screening data. J. Chem. Inf. Model. 51, 1528–1538 (2011).
Wetzel, S. et al. Interactive exploration of chemical space with scaffold hunter. Nature Chem. Biol. 5, 581–583 (2009).
Steinbeck C. et al. Recent developments of the chemistry development kit (CDK)—an open-source Java library for chemo- and bioinformatics. Curr. Pharmaceut. Des. 1217, 2111–2120 (2006).
Acknowledgements
The research leading to these results was supported by funding from the European Research Council (ERC) under the European Union's Seventh Framework Program (FP7/ 2007-2013) and ERC grant agreement no. 268309, from the German Federal Ministry for Education and Research through the German National Genome Research Network-Plus (NGFN-Plus) (grant no. BMBF 01GS08104 to H.W. and D.R.), as well as from the Fonds der Chemischen Industrie.
Author information
Authors and Affiliations
Contributions
B.O. and Y.N. performed computational experiments and syntheses. S.W. and S.R. performed computational experiments. B.O. performed biochemical experiments. B.O. and C.G. determined crystal structure analyses. B.O., S.W., D.R. and H.W. designed experiments. D.R. and H.W. supervised the research. B.O., S.W., D.R. and H.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 4921 kb)
Supplementary information
Representative natural product fragment library (PDF 6271 kb)
Supplementary information
Structure data file for representative natural product fragment library (SDF 2006 kb)
Supplementary information
Supplementary Table S1 - Summary of commercially available fragments (PDF 11016 kb)
Supplementary information
Structure data file for Supplementary Table S1 (SDF 1656 kb)
Rights and permissions
About this article
Cite this article
Over, B., Wetzel, S., Grütter, C. et al. Natural-product-derived fragments for fragment-based ligand discovery. Nature Chem 5, 21–28 (2013). https://doi.org/10.1038/nchem.1506
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.1506
This article is cited by
-
Biocatalytic strategy for the construction of sp3-rich polycyclic compounds from directed evolution and computational modelling
Nature Chemistry (2024)
-
Exploring protein hotspots by optimized fragment pharmacophores
Nature Communications (2021)
-
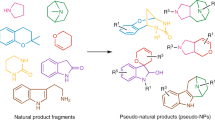
Natural product fragment combination to performance-diverse pseudo-natural products
Nature Communications (2021)
-
Marine furanocembranoids-inspired macrocycles enabled by Pd-catalyzed unactivated C(sp3)-H olefination mediated by donor/donor carbenes
Nature Communications (2021)
-
Protective effect of (+)–catechin against lipopolysaccharide-induced inflammatory response in RAW 264.7 cells through downregulation of NF-κB and p38 MAPK
Inflammopharmacology (2021)