Abstract
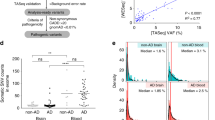
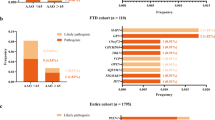
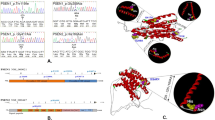
To assess the role of rare copy number variations in Alzheimer's disease (AD), we conducted a case–control study using whole-exome sequencing data from 522 early-onset cases and 584 controls. The most recurrent rearrangement was a 17q21.31 microduplication, overlapping the CRHR1, MAPT, STH and KANSL1 genes that was found in four cases, including one de novo rearrangement, and was absent in controls. The increased MAPT gene dosage led to a 1.6–1.9-fold expression of the MAPT messenger RNA. Clinical signs, neuroimaging and cerebrospinal fluid biomarker profiles were consistent with an AD diagnosis in MAPT duplication carriers. However, amyloid positon emission tomography (PET) imaging, performed in three patients, was negative. Analysis of an additional case with neuropathological examination confirmed that the MAPT duplication causes a complex tauopathy, including prominent neurofibrillary tangle pathology in the medial temporal lobe without amyloid-β deposits. 17q21.31 duplication is the genetic basis of a novel entity marked by prominent tauopathy, leading to early-onset dementia with an AD clinical phenotype. This entity could account for a proportion of probable AD cases with negative amyloid PET imaging recently identified in large clinical series.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
McKhann GM, Knopman DS, Chertkow H, Hyman BT, Jack CR, Kawas CH et al. The diagnosis of dementia due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement J Alzheimers Assoc 2011; 7: 263–269.
Ossenkoppele R, Jansen WJ, Rabinovici GD, Knol DL, van der Flier WM, van Berckel BNM et al. Prevalence of amyloid PET positivity in dementia syndromes: a meta-analysis. JAMA 2015; 313: 1939–1949.
Jack CR, Knopman DS, Chételat G, Dickson D, Fagan AM, Frisoni GB et al. Suspected non-Alzheimer disease pathophysiology—concept and controversy. Nat Rev Neurol 2016; 12: 117–124.
Jack CR, Knopman DS, Weigand SD, Wiste HJ, Vemuri P, Lowe V et al. An operational approach to National Institute on Aging-Alzheimer’s Association criteria for preclinical Alzheimer disease. Ann Neurol 2012; 71: 765–775.
Cuyvers E, De Roeck A, Van den Bossche T, Van Cauwenberghe C, Bettens K, Vermeulen S et al. Mutations in ABCA7 in a Belgian cohort of Alzheimer’s disease patients: a targeted resequencing study. Lancet Neurol 2015; 14: 814–822.
Guerreiro R, Wojtas A, Bras J, Carrasquillo M, Rogaeva E, Majounie E et al. TREM2 variants in Alzheimer’s disease. N Engl J Med 2013; 368: 117–127.
Jonsson T, Stefansson H, Steinberg S, Jonsdottir I, Jonsson PV, Snaedal J et al. Variant of TREM2 associated with the risk of Alzheimer’s disease. N Engl J Med 2013; 368: 107–116.
Le Guennec K, Nicolas G, Quenez O, Charbonnier C, Wallon D, Bellenguez C et al. ABCA7 rare variants and Alzheimer disease risk. Neurology 2016; 86: 2134–2137.
Nicolas G, Charbonnier C, Wallon D, Quenez O, Bellenguez C, Grenier-Boley B et al. SORL1 rare variants: a major risk factor for familial early-onset Alzheimer’s disease. Mol Psychiatry 2016; 21: 831–836.
Steinberg S, Stefansson H, Jonsson T, Johannsdottir H, Ingason A, Helgason H et al. Loss-of-function variants in ABCA7 confer risk of Alzheimer’s disease. Nat Genet 2015; 47: 445–447.
Hiltunen M, Helisalmi S, Mannermaa A, Alafuzoff I, Koivisto AM, Lehtovirta M et al. Identification of a novel 4.6-kb genomic deletion in presenilin-1 gene which results in exclusion of exon 9 in a Finnish early onset Alzheimer’s disease family: an Alu core sequence-stimulated recombination? Eur J Hum Genet EJHG 2000; 8: 259–266.
Prihar G, Verkkoniem A, Perez-Tur J, Crook R, Lincoln S, Houlden H et al. Alzheimer disease PS-1 exon 9 deletion defined. Nat Med 1999; 5: 1090.
Rovelet-Lecrux A, Hannequin D, Raux G, Le Meur N, Laquerrière A, Vital A et al. APP locus duplication causes autosomal dominant early-onset Alzheimer disease with cerebral amyloid angiopathy. Nat Genet 2006; 38: 24–26.
Hooli BV, Kovacs-Vajna ZM, Mullin K, Blumenthal MA, Mattheisen M, Zhang C et al. Rare autosomal copy number variations in early-onset familial Alzheimer’s disease. Mol Psychiatry 2014; 19: 676–681.
Rovelet-Lecrux A, Legallic S, Wallon D, Flaman J-M, Martinaud O, Bombois S et al. A genome-wide study reveals rare CNVs exclusive to extreme phenotypes of Alzheimer disease. Eur J Hum Genet EJHG 2012; 20: 613–617.
Brouwers N, Van Cauwenberghe C, Engelborghs S, Lambert J-C, Bettens K, Le Bastard N et al. Alzheimer risk associated with a copy number variation in the complement receptor 1 increasing C3b/C4b binding sites. Mol Psychiatry 2012; 17: 223–233.
Nicolas G, Wallon D, Charbonnier C, Quenez O, Rousseau S, Richard A-C et al. Screening of dementia genes by whole-exome sequencing in early-onset Alzheimer disease: input and lessons. Eur J Hum Genet EJHG 2016; 24: 710–716.
DeJesus-Hernandez M, Mackenzie IR, Boeve BF, Boxer AL, Baker M, Rutherford NJ et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 2011; 72: 245–256.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 2010; 20: 1297–1303.
Li H, Durbin R . Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinforma Oxf Engl 2010; 26: 589–595.
Anderson CA, Pettersson FH, Clarke GM, Cardon LR, Morris AP, Zondervan KT . Data quality control in genetic case-control association studies. Nat Protoc 2010; 5: 1564–1573.
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ . Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 2015; 4: 7.
Jun G, Flickinger M, Hetrick KN, Romm JM, Doheny KF, Abecasis GR et al. Detecting and estimating contamination of human DNA samples in sequencing and array-based genotype data. Am J Hum Genet 2012; 91: 839–848.
Backenroth D, Homsy J, Murillo LR, Glessner J, Lin E, Brueckner M et al. CANOES: detecting rare copy number variants from whole exome sequencing data. Nucleic Acids Res 2014; 42: e97.
Swaminathan S, Huentelman MJ, Corneveaux JJ, Myers AJ, Faber KM, Foroud T et al. Analysis of copy number variation in Alzheimer’s disease in a cohort of clinically characterized and neuropathologically verified individuals. PloS One 2012; 7: e50640.
Swaminathan S, Shen L, Kim S, Inlow M, West JD, Faber KM et al. Analysis of copy number variation in Alzheimer’s disease: the NIALOAD/ NCRAD Family Study. Curr Alzheimer Res 2012; 9: 801–814.
Alexander J, Kalev O, Mehrabian S, Traykov L, Raycheva M, Kanakis D et al. Familial early-onset dementia with complex neuropathologic phenotype and genomic background. Neurobiol Aging 2016; 42: 199–204.
Rovelet-Lecrux A, Hannequin D, Guillin O, Legallic S, Jurici S, Wallon D et al. Frontotemporal dementia phenotype associated with MAPT gene duplication. J Alzheimers Dis JAD 2010; 21: 897–902.
Campion D, Pottier C, Nicolas G, Le Guennec K, Rovelet-Lecrux A . Alzheimer disease: modeling an Aβ-centered biological network. Mol Psychiatry 2016; 21: 861–871.
Coe BP, Witherspoon K, Rosenfeld JA, van Bon BWM, Vulto-van Silfhout AT, Bosco P et al. Refining analyses of copy number variation identifies specific genes associated with developmental delay. Nat Genet 2014; 46: 1063–1071.
Ruderfer DM, Hamamsy T, Lek M, Karczewski KJ, Kavanagh D, Samocha KE et al. Patterns of genic intolerance of rare copy number variation in 59,898 human exomes. Nat Genet 2016; 48: 1107–1111.
Jack CR, Holtzman DM . Biomarker modeling of Alzheimer’s disease. Neuron 2013; 80: 1347–1358.
Duits FH, Teunissen CE, Bouwman FH, Visser P-J, Mattsson N, Zetterberg H et al. The cerebrospinal fluid ‘Alzheimer profile’: easily said, but what does it mean? Alzheimers Dement J Alzheimers Assoc 2014; 10: 713–723.e2.
Spillantini MG, Goedert M . Tau pathology and neurodegeneration. Lancet Neurol 2013; 12: 609–622.
Allen M, Kachadoorian M, Quicksall Z, Zou F, Chai HS, Younkin C et al. Association of MAPT haplotypes with Alzheimer’s disease risk and MAPT brain gene expression levels. Alzheimers Res Ther 2014; 6: 39.
Natacci F, Alfei E, Tararà L, D’Arrigo S, Zuffardi O, Gentilin B et al. Chromosome 17q21.31 duplication syndrome: Description of a new familiar case and further delineation of the clinical spectrum. Eur J Paediatr Neurol EJPN Off J Eur Paediatr Neurol Soc 2016; 20: 183–187.
Kitsiou-Tzeli S, Frysira H, Giannikou K, Syrmou A, Kosma K, Kakourou G et al. Microdeletion and microduplication 17q21.31 plus an additional CNV, in patients with intellectual disability, identified by array-CGH. Gene 2012; 492: 319–324.
Kirchhoff M, Bisgaard A-M, Duno M, Hansen FJ, Schwartz M . A 17q21.31 microduplication, reciprocal to the newly described 17q21.31 microdeletion, in a girl with severe psychomotor developmental delay and dysmorphic craniofacial features. Eur J Med Genet 2007; 50: 256–263.
Grisart B, Willatt L, Destrée A, Fryns J-P, Rack K, de Ravel T et al. 17q21.31 microduplication patients are characterised by behavioural problems and poor social interaction. J Med Genet 2009; 46: 524–530.
Coppola G, Chinnathambi S, Lee JJ, Dombroski BA, Baker MC, Soto-Ortolaza AI et al. Evidence for a role of the rare p.A152T variant in MAPT in increasing the risk for FTD-spectrum and Alzheimer’s diseases. Hum Mol Genet 2012; 21: 3500–3512.
Duyckaerts C, Braak H, Brion J-P, Buée L, Del Tredici K, Goedert M et al. PART is part of Alzheimer disease. Acta Neuropathol (Berl) 2015; 129: 749–756.
Jellinger KA, Alafuzoff I, Attems J, Beach TG, Cairns NJ, Crary JF et al. PART, a distinct tauopathy, different from classical sporadic Alzheimer disease. Acta Neuropathol (Berl) 2015; 129: 757–762.
Jun G, Ibrahim-Verbaas CA, Vronskaya M, Lambert J-C, Chung J, Naj AC et al. A novel Alzheimer disease locus located near the gene encoding tau protein. Mol Psychiatry 2016; 21: 108–117.
Acknowledgements
This study was funded by grants from the Clinical Research Hospital Program from the French Ministry of Health (GMAJ, PHRC 2008/067), the CNR-MAJ and the JPND PERADES (ANR-13-JPRF-0001-04). This work was supported by France Génomique, Labex GENMED ANR-10-LABX-0013, the National Foundation for Alzheimer’s disease and related disorders, the Institut Pasteur de Lille, the Centre National de Génotypage, Inserm, FRC (fondation pour la recherche sur le cerveau) and by the LABEX (laboratory of excellence program investment for the future) DISTALZ grant (Development of Innovative Strategies for a Transdisciplinary approach to Alzheimer’s disease). We are indebted to the Banque d'ADN et de cellules-Institut du Cerveau et de la Moelle épinière (ICM-Inserm U1127-UPMC P6 UMR-S 1127-CNRS UMR 7225).
Author information
Authors and Affiliations
Consortia
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
PowerPoint slides
Rights and permissions
About this article
Cite this article
Le Guennec, K., Quenez, O., Nicolas, G. et al. 17q21.31 duplication causes prominent tau-related dementia with increased MAPT expression. Mol Psychiatry 22, 1119–1125 (2017). https://doi.org/10.1038/mp.2016.226
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2016.226
This article is cited by
-
The genetic relationships between brain structure and schizophrenia
Nature Communications (2023)
-
Step by step: towards a better understanding of the genetic architecture of Alzheimer’s disease
Molecular Psychiatry (2023)
-
Brain structural abnormalities and cognitive changes in a patient with 17q21.31 microduplication and early onset dementia: a case report
Journal of Neurology (2023)
-
Dissecting the clinical heterogeneity of early-onset Alzheimer’s disease
Molecular Psychiatry (2022)
-
The contribution of CNVs to the most common aging-related neurodegenerative diseases
Aging Clinical and Experimental Research (2021)