Abstract
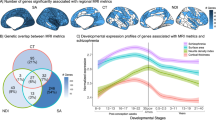
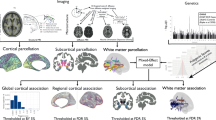
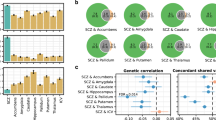
Schizophrenia is a devastating neurodevelopmental disorder with a complex genetic etiology. Widespread cortical gray matter loss has been observed in patients and prodromal samples. However, it remains unresolved whether schizophrenia-associated cortical structure variations arise due to disease etiology or secondary to the illness. Here we address this question using a partitioning-based heritability analysis of genome-wide single-nucleotide polymorphism (SNP) and neuroimaging data from 1750 healthy individuals. We find that schizophrenia-associated genetic variants explain a significantly enriched proportion of trait heritability in eight brain phenotypes (false discovery rate=10%). In particular, intracranial volume and left superior frontal gyrus thickness exhibit significant and robust associations with schizophrenia genetic risk under varying SNP selection conditions. Cross-disorder comparison suggests that the neurogenetic architecture of schizophrenia-associated brain regions is, at least in part, shared with other psychiatric disorders. Our study highlights key neuroanatomical correlates of schizophrenia genetic risk in the general population. These may provide fundamental insights into the complex pathophysiology of the illness, and a potential link to neurocognitive deficits shaping the disorder.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
21 February 2017
An Erratum to this paper has been published: https://doi.org/10.1038/mp.2017.42
References
Sullivan P, Kendler KS, Neale MC . Schizophrenia as a complex trait: evidence from a meta-analysis of twin studies. Arch Gen Psychiatry 2003; 60: 1187–1192.
van Erp T, Hibar DP, Rasmussen JM, Glahn DC, Pearlson GD, Andreassen OA et al. Subcortical brain volume abnormalities in 2028 individuals with schizophrenia and 2540 healthy controls via the ENIGMA consortium. Mol Psychiatry 2016; 21: 547–553.
Bhojraj T, Francis AN, Montrose DM, Keshavan MS . Grey matter and cognitive deficits in young relatives of schizophrenia patients. Neuroimage 2011; Suppl 1: S287–S292.
Modinos G, Vercammen A, Mechelli A, Knegtering H, McGuire PK, Aleman A . Structural covariance in the hallucinating brain: a voxel-based morphometry study. J Psychiatry Neurosci 2009; 34: 465–469.
Modinos G, Costafreda SG, van Tol MJ, McGuire PK, Aleman A, Allen P . Neuroanatomy of auditory verbal hallucinations in schizophrenia: a quantitative meta-analysis of voxel-based morphometry studies. Cortex 2013; 49: 1046–1055.
van Tol M, van der Meer L, Bruggeman R, Modinos G, Knegtering H, Aleman A . Voxel-based gray and white matter morphometry correlates of hallucinations in schizophrenia: the superior temporal gyrus does not stand alone. Neuroimage Clin 2013; 4: 249–257.
van Lutterveld R, van den Heuvel MP, Diederen KM, de Weijer AD, Begemann MJ, Brouwer RM et al. Cortical thickness in individuals with non-clinical and clinical psychotic symptoms. Brain 2014; 137 ((Pt 10)): 2664–2669.
Wible C, Anderson J, Shenton ME, Kricun A, Hirayasu Y, Tanaka S et al. Prefrontal cortex, negative symptoms, and schizophrenia: an MRI study. Psychiatry Res 2001; 108: 65–78.
Gottesman I, Gould TD . The endophenotype concept in psychiatry: etymology and strategic intentions. Am J Psychiatry 2003; 160: 636–645.
Mattai A, Chavez A, Greenstein D, Clasen L, Bakalar J, Stidd R et al. Effects of clozapine and olanzapine on cortical thickness in childhood-‐onset schizophrenia. Schizophr Res 2010; 116: 44–48.
Purcell S, Wray NR, Stone JL, Visscher PM, O'Donovan MC, Sullivan PF et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 2009; 460: 748–752.
Tesli M, Espeseth T, Bettella F, Mattingsdal M, Aas M, Melle I et al. Polygenic risk score and the psychosis continuum model. Acta Psychiatr Scand 2014; 130: 311–317.
Meyer-Lindenberg A . Imaging genetics of schizophrenia. Dialogues Clin Neurosci 2010; 12: 449–456.
Walton E, Geisler D, Lee PH, Hass J, Turner JA, Liu J et al. Prefrontal inefficiency is associated with polygenic risk for schizophrenia. Schizophr Bull 2014; 40: 1263–1271.
Esslinger C, Walter H, Kirsch P, Erk S, Schnell K, Arnold C et al. Neural mechanisms of a genomewide supported psychosis variant. Science 2009; 324: 605.
Kauppi K, Westlye LT, Tesli M, Bettella F, Brandt CL, Mattingsdal M et al. Polygenic risk for schizophrenia associated with working memory-related prefrontal brain activation in patients with schizophrenia and healthy controls. Schizophr Bull 2015; 41: 736–743.
Walton E, Turner J, Gollub RL, Manoach DS, Yendiki A, Ho BC et al. Cumulative genetic risk and prefrontal activity in patients with schizophrenia. Schizophr Bull 2013; 39: 703–711.
van Erp T, Guella I, Vawter MP, Turner J, Brown GG, McCarthy G et al. Schizophrenia miR-137 locus risk genotype is associated with dorsolateral prefrontal cortex hyperactivation. Biol Psychiatry 2014; 75: 398–405.
Oertel-Knöchel V, Lancaster TM, Knöchel C, Stäblein M, Storchak H, Reinke B et al. Schizophrenia risk variants modulate white matter volume across the psychosis spectrum: evidence from two independent cohorts. Neuroimage Clin 2015; 7: 764–770.
Terwisscha vSA, Bakker SC, van Haren NE, Derks EM, Buizer-Voskamp JE, Boos HB et al. Genetic schizophrenia risk variants jointly modulate total brain and white matter volume. Biol Psychiatry 2013; 73: 525–531.
Yang J, Lee H, Goddard ME, Visscher PM . GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet 2011; 88: 76–82.
Lee SH, Ripke S, Neale BM, Faraone SV, Purcell SM, Perlis RH et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat Genet 2013; 45: 984–994.
Davis L, Yu D, Keenan CL, Gamazon ER, Konkashbaev AI, Derks EM et al. Partitioning the heritability of Tourette syndrome and obsessive compulsive disorder reveals differences in genetic architecture. PLoS Genet 2013; 9: e1003864.
Lee S, DeCandia TR, Ripke S, Yang J et alSchizophrenia Psychiatric Genome-Wide Association Study Consortium (PGC-SCZ) International Schizophrenia Consortium (ISC). Estimating the proportion of variation in susceptibility to schizophrenia captured by common SNPs. Nat Genet 2012; 44: 247–250.
Ge T, Nichols T, Lee PH, Holmes AJ, Roffman JL, Buckner RL et al. Massively expedited genome-wide heritability analysis (MEGHA). Proc Natl Acad Sci U S A 2015; 112: 2479–2484.
Bryant C, Giovanello KS, Ibrahim JG, Chang J, Shen D, Peterson BS et al. Mapping the genetic variation of regional brain volumes as explained by all common SNPs from the ADNI study. PLoS One 2013; 8: e71723.
Gusev A, Bhatia G, Zaitlen N, Vilhjalmsson BJ, Diogo D, Stahl EA et al. Quantifying missing heritability at known GWAS loci. PLoS Genet 2013; 9: e1003993.
Gusev A, Lee SH, Trynka G, Finucane H, Vilhjálmsson BJ, Xu H et al. Regulatory and cell-type-specific variants across 11 common diseases. Am J Hum Genet 2014; 95: 535–552.
Bulik-Sullivan B, Loh PR, Finucane HK, Ripke S, Yang J et alSchizophrenia Working Group of the Psychiatric Genomics Consortium. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet 2015; 47: 291–295.
Finucane H, Bulik-Sullivan B, Gusev A, Trynka G, Reshef Y, Loh PR et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat Genet 2015; 47: 1228–1235.
de Zubicaray G, Chiang MC, McMahon KL, Shattuck DW, Toga AW, Martin NG et al. Meeting the challenges of neuroimaging genetics. Brain Imaging Behav 2008; 2: 258–263.
Holmes A, Hollinshead MO, O'Keefe TM, Petrov VI, Fariello GR, Wald LL et al. Brain Genomics Superstruct Project initial data release with structural, functional, and behavioral measures. Sci Data 2015; 2: 150031.
Holmes A, Lee PH, Hollinshead M, Bakst L, Roffman JL, Smoller JW et al. Individual differences in amygdala-medial prefrontal anatomy link negative affect, impaired social functioning, and polygenic depression risk. J Neurosci 2012; 32: 18087–18100.
Rentería M, Hansell NK, Strike LT, McMahon KL, de Zubicaray GI, Hickie IB et al. Genetic architecture of subcortical brain regions: common and region-specific genetic contributions. Genes Brain Behav 2014; 13: 821–830.
Fischl B . FreeSurfer. Neuroimage 2012; 62: 774–781.
Desikan R, Ségonne F, Fischl B, Quinn BT, Dickerson BC, Blacker D et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 2006; 31: 968–980.
Winkler A, Kochunov P, Blangero J, Almasy L, Zilles K, Fox PT et al. Cortical thickness or grey matter volume? The importance of selecting the phenotype for imaging genetics studies. Neuroimage 2010; 53: 1135–1146.
Stein J, Medland SE, Vasquez AA, Hibar DP, Senstad RE, Winkler AM et al. Identification of common variants associated with human hippocampal and intracranial volumes. Nat Genet 2012; 44: 552–561.
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D . Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 2006; 38: 904–909.
Barnes J, Ridgway GR, Bartlett J, Henley SM, Lehmann M, Hobbs N et al. Head size, age and gender adjustment in MRI studies: a necessary nuisance? Neuroimage 2010; 53: 1244–1255.
Ripke S, Neale BM, Corvin A, Walters JT, Farh KH, Holmans PA et al. Biological insights from 108 schizophrenia-associated genetic loci. Nature 2014; 511: 421–427.
Toro R, Poline JB, Huguet G, Loth E, Frouin V, Banaschewski T et al. Genomic architecture of human neuroanatomical diversity. Mol Psychiatry 2015; 20: 1011–1016.
Smoller JW, Ripke S, Lee PH, Neale B, Nurnberger JI, Santangelo S et al. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 2013; 381: 1371–1379.
Lyall A, Shi F, Geng X, Woolson S, Li G, Wang L et al. Dynamic development of regional cortical thickness and surface area in early childhood. Cereb Cortex 2015; 25: 2204–2212.
Baribeau D, Anagnostou E . A comparison of neuroimaging findings in childhood onset schizophrenia and autism spectrum disorder: a review of the literature. Front Psychiatry 2013; 4: 175.
Chen J, Nedivi E . Neuronal structural remodeling: Is it all about access? Curr Opin Neurobiol 2010; 20: 557–562.
Moyer C, Shelton MA, Sweet RA . Dendritic spine alterations in schizophrenia. Neurosci Lett 2015; 601: 46–53.
Roffman J, Weiss AP, Goff DC, Rauch SL, Weinberger DR . Neuroimaging-genetic paradigms: a new approach to investigate the pathophysiology and treatment of cognitive deficits in schizophrenia. Harv Rev Psychiatry 2006; 14: 78–91.
Baker J, Holmes AJ, Masters GA, Yeo BT, Krienen F, Buckner RL et al. Disruption of cortical association networks in schizophrenia and psychotic bipolar disorder. JAMA Psychiatry 2014; 71: 109–118.
Gong Q, Lui S, Sweeney JA . A selective review of cerebral abnormalities in patients with first-episode schizophrenia before and after treatment. Am J Psychiatry 2015; 173: 232–243.
Ordóñez A, Luscher ZI, Gogtay N . Neuroimaging findings from childhood onset schizophrenia patients and their non-psychotic siblings. Schizophr Res 2015; 173: 124–131.
Cannon T, Chung Y, He G, Sun D, Jacobson A, van Erp TG et al. Progressive reduction in cortical thickness as psychosis develops: a multisite longitudinal neuroimaging study of youth at elevated clinical risk. Biol Psychiatry 2015; 77: 147–157.
Tully L, Lincoln SH, Liyanage-Don N, Hooker CI . Impaired cognitive control mediates the relationship between cortical thickness of the superior frontal gyrus and role functioning in schizophrenia. Schizophr Res 2014; 152: 358–364.
Yeo B, Krienen FM, Sepulcre J, Sabuncu MR, Lashkari D, Hollinshead M et al. The organization of the human cerebral cortex estimated by intrinsic functional connectivity. J Neurophysiol 2011; 106: 1125–1165.
Spreng R, Sepulcre J, Turner GR, Stevens WD, Schacter DL . Intrinsic architecture underlying the relations among the default, dorsal attention, and frontoparietal control networks of the human brain. J Cogn Neurosci 2013; 25: 74–86.
Li W, Qin W, Liu H, Fan L, Wang J, Jiang T et al. Subregions of the human superior frontal gyrus and their connections. Neuroimage 2013; 78: 46–58.
Du Y, Pearlson GD, Yu Q, He H, Lin D, Sui J et al. Interaction among subsystems within default mode network diminished in schizophrenia patients: a dynamic connectivity approach. Schizophr Res 2016; 170: 55–65.
Pomarol-Clotet E, Salvador R, Sarró S, Gomar J, Vila F, Martínez A et al. Failure to deactivate in the prefrontal cortex in schizophrenia: dysfunction of the default mode network? Psychol Med 2008; 38: 1185–1193.
Chang X, Shen H, Wang L, Liu Z, Xin W, Hu D et al. Altered default mode and fronto-parietal network subsystems in patients with schizophrenia and their unaffected siblings. Brain Res 2014; 1562: 87–99.
Nekovarova T, Fajnerova I, Horacek J, Spaniel F . Bridging disparate symptoms of schizophrenia: a triple network dysfunction theory. Front Behav Neurosci 2014; 8: 171.
Deacon T Human brain evolution II. Embryology and brain allometry. In: Jerison H, Jerison I (eds). Intelligence and Evolutionary Biology. Springer-Verlag: Berlin, 1988, p 383–415.
Thompson P . Cracking the brain’s genetic code. Proc Natl Acad Sci USA 2015; 112: 15269–15270.
Narr K, Woods RP, Thompson PM, Szeszko P, Robinson D, Dimtcheva T et al. Relationships between IQ and regional cortical gray matter thickness in healthy adults. Cereb Cortex 2007; 17: 2163–2171.
Narr K, Bilder RM, Toga AW, Woods RP, Rex DE, Szeszko PR et al. Mapping cortical thickness and gray matter concentration in first episode schizophrenia. Cereb Cortex 2005; 15: 708–719.
Schultz C, Koch K, Wagner G, Roebel M, Schachtzabel C, Gaser C et al. Reduced cortical thickness in first episode schizophrenia. Schizophr Res 2010; 116: 204–209.
Greven CU, Bralten J, Mennes M, O'Dwyer L, van Hulzen KJ, Rommelse N et al. Developmentally stable whole-brain volume reductions and developmentally sensitive caudate and putamen volume alterations in those with attention-deficit/hyperactivity disorder and their unaffected siblings. JAMA Psychiatry 2015; 72: 490–499.
Kucharsky HR, Alter R, Sojoudi S, Ardekani BA, Kuzniecky R, Pardoe HR . Corpus callosum area and brain volume in autism spectrum disorder: quantitative analysis of structural MRI from the ABIDE database. J Autism Dev Disord 2015; 45: 3107–3114.
McIntosh A, Owens DC, Moorhead WJ, Whalley HC, Stanfield AC, Hall J et al. Longitudinal volume reductions in people at high genetic risk of schizophrenia as they develop psychosis. Biol Psychiatry 2011; 69: 953–958.
Lawrie S, McIntosh AM, Hall J, Owens DG, Johnstone EC . Brain structure and function changes during the development of schizophrenia: the evidence from studies of subjects at increased genetic risk. Schizophr Bull 2008; 34: 330–340.
Brans R, van Haren NE, van Baal GC, Schnack HG, Kahn RS, Hulshoff Pol HE . Heritability of changes in brain volume over time in twin pairs discordant for schizophrenia. Arch Gen Psychiatry 2008; 65: 1259–1268.
Sabuncu M, Buckner R, Smoller JW, Lee PH, Fischl B, Sperling R . The association between a polygenic alzheimer score and cortical thickness in cognitively normal subjects. Cereb Cortex 2011; 22: 2653–2661.
Voineskos A, Felsky D, Wheeler AL, Rotenberg DJ, Levesque M, Patel S et al. Limited evidence for association of genome-wide schizophrenia risk variants on cortical neuroimaging phenotypes. Schizophr Bull 2015; 42: 1027–1036.
Franke B, Stein JL, Ripke S, Anttila V, Hibar DP, van Hulzen KJ et al. Genetic influences on schizophrenia and subcortical brain volumes: large-scale proof of concept. Nat Neurosci 2016; 19: 420–431.
Lee S, Yang J, Goddard ME, Visscher PM, Wray NR . Estimation of pleiotropy between complex diseases using SNP-derived genomic relationships and restricted maximum likelihood. Bioinformatics 2012; 28: 2540–2542.
Bulik-Sullivan B, Finucane HK, Anttila V, Gusev A, Day FR, Loh PR et al. An atlas of genetic correlations across human diseases and traits. Nat Genet 2015; 47: 1236–1241.
Polderman T, Benyamin B, de Leeuw CA, Sullivan PF, van Bochoven A, Visscher PM et al. Meta-analysis of the heritability of human traits based on fifty years of twin studies. Nat Genet 2015; 47: 702–709.
Chen C, Peng Q, Schork AJ, Lo MT, Fan CC, Wang Y et al. Pediatric imaging neurocognition and genetics study; Alzheimer's disease neuroimaging initiative large-scale genomics unveil polygenic architecture of human cortical surface area. Nat Commun 2015; 6: 7549.
Morales A, Ghahremani D, Kohno M, Hellemann GS, London ED . Cigarette exposure, dependence, and craving are related to insula thickness in young adult smokers. Neuropsychopharmacology 2014; 39: 1816–1822.
Li Y, Yuan K, Cai C, Feng D, Yin J, Bi Y et al. Reduced frontal cortical thickness and increased caudate volume within fronto-striatal circuits in young adult smokers. Drug Alcohol Depend 2015; 151: 211–219.
Acknowledgements
Neuroimaging genetics data were provided by the Brain Genomics Superstruct Project (GSP) of Harvard University and MGH, with support from the Center for Brain Science Neuroinformatics Research Group, Athinoula A Martinos Center for Biomedical Imaging, Center for Human Genetic Research, and Stanley Center for Psychiatric Research. Twenty individual investigators at Harvard and MGH generously contributed data to the overall project. The QTIM study is supported by grants from NIH (R01 HD050735) and the NHMRC (389875, 486682 and 1009064). We thank the twins and siblings for their participation, Marlene Grace and Ann Eldridge for twin recruitment, Aiman Al Najjar and other radiographers for scanning, Kerrie McAloney and Daniel Park for research support, and Anjali Henders and staff for DNA sample processing and preparation. PMT, NJ and MJW were supported in part by a Consortium grant (U54 EB020403 to PMT) from the NIH Institutes contributing to the Big Data to Knowledge (BD2K) Initiative. This research was also funded in part by NIH Grants K99MH101367 (to PHL); K23MH104515 (to JTB), K01MH099232 (to AJH); K24MH094614 and R01 MH101486 (to JWS); JWS is a Tepper Family MGH Research Scholar.
Author contributions
Project conception and experiment design: PHL. Statistical analysis and interpretation of findings: PHL, JTB, J-YJ, JWS, DO and DSM. Imaging and genetics data generation and processing: AJH, PHL, RB, JR, JWS, TG, NJ, DPH, JF, KLM, GIZ, NGM, MJW and PMT. Writing of the manuscript: PHL, JTB, JWS, TG and YC. Revision of the manuscript: AJH, JTB, JWS, TG, NJ and PMT.
Web Resources
FREESURFER: http://surfer.nmr.mgh.harvard.edu
PLINK (ver. 1.9): https://www.cog-genomics.org/plink2/
EIGENSOFT (ver. 6.1): http://www.hsph.harvard.edu/alkes-price/software/
The GCTA-GREML Power Calculator: http://cnsgenomics.com/shiny/gctaPower/
Genome-wide Complex Trait Analysis (GCTA ver. 1.24.4): http://www.complextraitgenomics.com/software/gcta/
Psychiatric Genomic Consortium (PGC) SCZ, BIP, ASD, ADHD, MDD summary statistics: http://www.med.unc.edu/pgc/downloads
Crohn’s disease summary statistics: http://www.ibdgenetics.org/downloads.html
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Rights and permissions
About this article
Cite this article
Lee, P., Baker, J., Holmes, A. et al. Partitioning heritability analysis reveals a shared genetic basis of brain anatomy and schizophrenia. Mol Psychiatry 21, 1680–1689 (2016). https://doi.org/10.1038/mp.2016.164
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2016.164
This article is cited by
-
The genetic relationships between brain structure and schizophrenia
Nature Communications (2023)
-
Enrichment of self-domestication and neural crest function loci in the heritability of neurodevelopmental disorders
Human Genetics (2023)
-
Recent natural selection conferred protection against schizophrenia by non-antagonistic pleiotropy
Scientific Reports (2023)
-
A cognitive neurogenetic approach to uncovering the structure of executive functions
Nature Communications (2022)
-
Associations Between Traumatic Stress, Brain Volumes and Post-traumatic Stress Disorder Symptoms in Children: Data from the ABCD Study
Behavior Genetics (2022)