Abstract
Huntington’s disease (HD), a progressive neurodegenerative disease, is caused by an expanded CAG triplet repeat producing a mutant huntingtin protein (mHTT) with a polyglutamine-repeat expansion. Onset of symptoms in mutant huntingtin gene-carrying individuals remains unpredictable. We report that synthetic polyglutamine oligomers and cerebrospinal fluid (CSF) from BACHD transgenic rats and from human HD subjects can seed mutant huntingtin aggregation in a cell model and its cell lysate. Our studies demonstrate that seeding requires the mutant huntingtin template and may reflect an underlying prion-like protein propagation mechanism. Light and cryo-electron microscopy show that synthetic seeds nucleate and enhance mutant huntingtin aggregation. This seeding assay distinguishes HD subjects from healthy and non-HD dementia controls without overlap (blinded samples). Ultimately, this seeding property in HD patient CSF may form the basis of a molecular biomarker assay to monitor HD and evaluate therapies that target mHTT.
Similar content being viewed by others
Introduction
Huntington’s disease (HD) is an autosomal dominant neurodegenerative disease that results in progressive motor, cognitive and psychiatric impairment and ultimately death. The disease is caused by an abnormal CAG trinucleotide repeat expansion within exon 1 of the HD gene that produces a mutant huntingtin protein (mHTT) containing an expanded polyglutamine (poly(Q)n, n⩾36) tract.1 The length of the expansion has an inverse relationship to the age-of-onset of clinical motor symptoms.2, 3 The repeat expansion confers an increased misfolding and aggregation propensity to mHTT, leading to the build-up of soluble and insoluble mHTT aggregates and a corresponding deficit in the intracellular quality control of protein folding.4, 5, 6 Amino terminal fragments of expanded polyQ-containing mHTT protein can readily form oligomers and ultimately insoluble beta-sheet-rich amyloid-like fibrillar aggregates in vitro and in vivo in a polyQ length-dependent manner.7 The ultimate relationship of specific aggregation species to disease manifestation is unclear. Although the inability to clear mHTT may represent a major pathogenic property,8 the propensity to aggregate represents a property of mHTT that reproducibly correlates with disease pathogenesis.9
A potentially central concept has emerged recently for neurodegenerative diseases including Alzheimer’s, Parkinson’s and HD that transcellular protein aggregation and propagation may occur and reflect an underlying and associated spreading of misfolded mutant proteins.9 On the basis of previous studies demonstrating that intracellular aggregation of mHTT can be ‘seeded‘ by addition of exogenous polyglutamine peptides,10, 11 we report here that cerebrospinal fluid (CSF) from a rat model of disease and from human HD subjects contains a mHTT-dependent property that seeds aggregation. To evaluate this property, we developed a quantitative cell-based aggregation (CBA) seeding assay. This seeding assay has two measures—the percentage of cells with aggregates12 and the amount of aggregates in cell lysates. The lysates are obtained from Htt14A2.6 PC12 cells that inducibly express expanded repeat mHTT polypeptide. We demonstrate that seeding is dose dependent and that extracellular seeds are incorporated into endogenously expressed mHTT species, forming aggregated fibrils. Seeding can be immunodepleted with antibody to mHTT, demonstrating that the CBA assay is mHTT dependent. This incorporation was visualized in cell extracts by correlative cryogenic-fluorescent light microscopy (cryo-FLM) and cryo-electron tomography (cryo-ET). Using the CBA assay as a measure of pathophysiological changes in HD, CSF from an HD rat model and human HD patients was found to ‘seed‘ significantly more aggregates compared with CSF from non-HD subjects. Mutant HTT can be detected in CSF from both HD autopsy samples and living HD subjects on filter blots with antibodies, and in the CBA seeding assay. These findings support transcellular propagation and associated spreading of misfolded mHTT protein and raise the possibility of using this seeding as a sensitive molecular biomarker assay to assess mHTT levels and aggregation propensity in HD. No effective disease-modifying treatments are currently available for HD. The lack of predictive and efficient blood or CSF biomarkers that track with clinical, neuropathological and molecular measures of disease progression has been a critical barrier to the development of effective HD treatments. The findings presented raise the possibility of using the seeding property inherent in HD CSF as a molecular biomarker assay to assess mHTT levels and aggregation propensity in HD.
Materials and methods
Oligomeric seeds
PolyQ seeds were prepared from synthetic peptides of polyQ30 and polyQ40 as described previously.13 The preparation of Alexa Fluor 647-, Cy3-labeled polyQ40 seeds and HiLyte Fluor 647-labeled Aβ1–40 and Aβ1–42 are described in the Supplementary Materials.
CSF samples
Investigators were blinded to sample group membership and specific diagnoses. Postmortem CSF was obtained from the UCLA Human Brain and Spinal Fluid Resource Center. The number of CAG repeats for HD cases were genotyped by Laragen (Culver City, CA, USA) using dissected frozen brain tissues. The dementia control CSF was provided by the Easton Center for Alzheimer’s Disease Research at UCLA. PREDICT CSF samples were provided by Coriell Institute for Medical Research. BACHD rat samples were provided by CHDI Foundation. CSF was aspirated from BACHD rats as described in the Supplementary Materials. For the immunodepletion of mHTT in CSF experiment, 100 μl CSF was incubated with 20 μl protein A/G PLUS-Agrose beads (Santa Cruz Biotechnologies, Dallas, TX, USA) that were preconjugated with varying amount of MAB1574 polyQ expansion-specific antibody (EMD Millipore, Temecula, CA, USA) for 1 h at 4 °C with gentle mixing. The supernatants were saved for aggregation seeding assays.
Cell culture and cell-based seeding assay
The Htt14A2.6 PC12 line was generated and propagated as described previously.12 In brief, Htt14A2.6 was maintained in Dulbecco's modified eagle medium-GlutaMAX (Life Technologies, Grand Island, NY, USA) supplemented with horse serum (10%), heat-inactivated fetal bovine serum (5%), Pen Strip (1%), Zeocin (200 μg ml−1), and G418 (50 μg ml−1). For cell assays, cells in the log phase of growth were replated in 24-well culture plates around 50–60% confluency, grown overnight and induced for the expression of mHttex1-GFP for 8 h with 2 μM ponasterone A (Santa Cruz Biotechnologies). To treat cells, different doses of polyQ40 seeds (1.0 pM–5 μM) or CSF (2% final concentration) sample were directly diluted from stock solution with the same cell growth medium containing 2 μM ponasterone A for growth for another 16 h. Equal volume of 1% fetal bovine serum diluted in 1 × phosphate-buffered saline was used as mock control.
Cell lysate seeding aggregation assay
Htt14A2.6 PC12 cells that were grown in medium containing 2 μM ponasterone A for 8–10 h and induced expression of mHttex1-GFP but no visible intracellular aggregates were harvested, rinsed, homogenized in cold phosphate-buffered saline containing protease inhibitor cocktail, centrifuged at 16 000 g for 30 min at 4°C twice. Total protein concentrations of supernatants were calibrated using the Sigma-Aldrich (St Louis, MO, USA) total protein assay kit following vendor’s instruction. An aliquot of 100 μg total protein of the supernatant, preformed oligomeric seeds or human or BACHD CSF samples (2.5 μl) were mixed together to make a final volume of 25 μl in an Eppendorf tube for incubation at RT in dark for 16 h. Aggregates formed in the cell-free assays were then measured by filter retardation assay.
Filter retardation assay
Filter retardation was performed according to published methods with minor modifications14 (see Supplementary Materials).
Dot blot immunoassays
Dot blot immunoassays are described in the Supplementary Materials.
Cryo-FLM and cryo-ET
Cryo-FLM and cryo-ET of polyQ40 seeded aggregates are described in the Supplementary Materials.
Total internal reflection microscopy
Total internal reflection microscopy of polyQ40 or Aβ seeded aggregation methods are described in the Supplementary Materials.
SDS-AGE gel electrophoresis
The gel analysis of oligomer-mediated seeding in the cell-free seeding aggregation assay was carried out as previously described.15, 16
Statistical analysis
Mixed model regression analyses17 (SAS Proc Mixed, v9.2) examined the effects of condition (for example, 0–5000 nM), time (for example, 8, 16 and 24 h), and the condition × time interaction on percent aggregation or amount of accumulation using a within-condition repeated measurement factor. Covariances among the repeated measurement factors were modeled with the unstructured-compound symmetry (un@cs) covariance matrix and denominator degrees of freedom were estimated using the Kenward–Roger method.18 The normality of the distribution could not be determined with our sample size, therefore we also performed nonparametric Friedman’s χ2-tests for repeated measures for which the equal variance and normality assumptions of parametric tests are not applicable. The nonparametric tests produced similar results (Supplementary Table S5). We report the analysis of variance results given that the variables are continuous and the reported means are considered more relevant and interpretable to the field than rank order values.
Results
Mutant HTT aggregation can be selectively seeded by polyQ oligomers
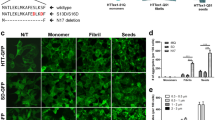
Exogenous applications of preformed oligomers have been shown to induce protein aggregation in vitro and in vivo for several proteins involved in neurodegeneration.19, 20, 21 In the case of polyglutamine repeats, nucleation and seeding of aggregation for polyglutamine repeats appears to be sequence specific.10 We used an inducible Htt14A2.6 cell model that has previously been used to assay agents that modulate formation of mHTT aggregates12, 16 as an assay system to evaluate whether exogenous application of preformed oligomers could enhance aggregation of intracellular mHTT.12, 16 This cell inducibly expresses a fragment of mHTT (truncated exon 1) epitope tagged with enhanced green fluorescence protein (mHttex1-GFP) in the presence of ponasterone A. We used synthetic polyQ (KKQ30KK and KKQ40KK) oligomers as seeds to induce mHTT aggregation in Htt14A2.6 cells. The assay was optimized to establish a limiting concentration of ponasterone A such that aggregation was minimal and any enhancement could be quantified. Oligomer concentration and time course of aggregation seeding post ponasterone A induction were also optimized. The percentage of cells with visible GFP-tagged aggregates under fluorescence and differential interference contrast microscopy was correlated to the concentration of KKQ40KK (polyQ40) seeds (10 nM–5 μM) and the incubation time (8–24 h; Figure 1a). In control experiments, cells without added polyQ40 show basal levels of aggregation (Figure 1b). We established a second assay that could detect a broader range of aggregated species using cell lysates (supernatants) prepared from induced Htt14A2.6 cells (Figure 2a). Notably, this assay provided greater sensitivity in detecting seeding by polyQ than the cell-based assay to monitor visible aggregates (Figure 2b). A control cell lysate experiment with polyQ30 displays no enhanced aggregation (Supplementary Figure S1).
Enhanced formation of mHTT aggregates in inducible Htt14A2.6 cells by preformed polyQ40 (KKQ40KK) oligomeric seeds. (a) Increased percentage of cells with mHTT aggregates (n=6) with GFP tags following oligomeric polyQ seeding at 8, 16 and 24 h post seeding. Error bars, s.e.m.; concentration: [F(6,42.4)=53.91, P<0.0001]; time: [F(2,89.7)=223.76, P<0.0001]; interaction: [F(12,52.6)=2.69, P=0.0067]. (b) Microscopic images of Htt14A2.6 cells expressing GFP-tagged mHTT seeded with preformed polyQ40 oligomeric seeds at 16 h post seeding. Bars indicate s.e.m. Each experiment had three to nine repeats. DIC, differential interference contrast. GFP, green fluorescent protein; mHTT, mutant huntingtin protein.
Enhanced formation of mHTT aggregates in cell-free lysates derived from Htt14A2.6 cells by oligomeric polyQ seeds. (a) Schematic of mHTT aggregation assay using ponasterone A-induced Htt14A2.6-derived whole-cell lysates (supernatants). (b) Dose-dependent quantification of mHTT aggregation from derived cell-free lysates on filter trap assay following oligomeric polyQ seeding. The y axis is the number of fold increases over the vehicle control. Concentration: [F(8, 3.92)=6.33, P=0.048]. (c) Correlative cryo-FLM and cryo-ET of cell lysate containing polyQ40 seeded mHttex1-GFP aggregates on electron microscope grids. Cryo-FLM identified a star-shaped aggregate that consists of oligomer seeds (red) and mHttex1-GFP (green). Black dashed box showed the area where an electron microscope (EM) image (at magnification of 4k) was taken (Black square is ice contamination adsorbed to the grid during transfer from light microscope to electron microscope). This image was correlated to the cryo-FLM image to confirm the aggregate’s identity. Yellow dashed box enclosed the region where the tomographic tilt series shown in d was collected. (d) Section and three-dimensional annotated view of polyQ40 seeded mHttex1-GFP aggregates in the cell lysate reveal the short mHTT fibril (yellow), long fibril and bundle aggregate structures (magenta), as well as cellular vesicles (blue) from the lysates. Bars indicate s.e.m. Each experiment had three to nine repeats. Cryo-ET, cryo-electron tomography; cryo-FLM, cryogenic-fluorescent light microscopy; mHTT, mutant huntingtin protein.
To probe for evidence of a direct interaction of the seeds with the induced aggregates and a potential mechanism for protein propagation, we used a correlative study of cryo-FLM and cryo-ET. Cryo-FLM identified a range of aggregates of variable sizes and morphologies (Figures 2c and d). The mixed colors of fluorescence confirmed that they are formed by the Cy3-labeled polyQ40 seeds and the inducibly expressed mHttex1-GFP. Cryo-ET of the same polyQ40 seeded aggregates as identified in Cryo-FLM revealed a range of aggregation species, including small oligomers, short and long fibrils and bundles of fibril assemblies (Figures 2c and d; supplementary movie S1).22, 23 Similar results have been observed in 14 tomograms from the corresponding cell lysate samples. These seeded aggregates are much more diverse and larger than those used for the seeding (Supplementary Figure S2b). In addition to the induced aggregation detected using filter blots and correlative cryo-FLM and cryo-ET imaging, newly formed aggregation of mHttex1 seeded by polyQ40 oligomers was also confirmed using agarose gel electrophoresis (AGE gels) followed by western blotting using a GFP-specific antibody (Supplementary Figure S1) as described previously.15
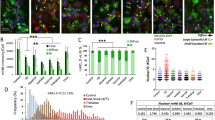
To test whether the seeding of aggregation is template specific for mHTT using the CBA seeding assays, Htt14A2.6 cells were treated with 0.5 μM of Aβ1–40 and Aβ1–42 (Alzheimer’s disease associated), and α-synuclein and A53T α-synuclein (Parkinson’s disease associated) oligomers in the same manner as polyQ40 seeds. No significant enhancement of intracellular mHttex1-GFP aggregates was detected (Figure 3). Consistent results were obtained from filter retardation-GFP immunoblotting following the cell-free aggregation assay; no notable changes in mHttex1 aggregation were found in the cell lysates seeded with up to 5 μM of Aβ1–40, Aβ1–42, α-synuclein or A53T α-synuclein oligomers, using a 50-fold concentration increase over polyQ40 seeds (Figures 3a and b). These observations were further corroborated by the total internal reflection microscopy imaging results (Supplementary Figure S3). In contrast to the polyQ40 oligomers that were mostly linked with mHttex1-GFP aggregates from total internal reflection microscopy imaging, few labeled Aβ1–42 oligomers overlapped with GFP fluorescence from the mHttex1-GFP aggregates (Supplementary Figure S3). In addition, Alexa Flour 647-conjugated polyQ40 co-aggregated with mHTT aggregates but not Aβ aggregates as determined by fluorescence correlation spectroscopy (data not shown), further supporting template specificity.
PolyQ-related template-specific seeding for mHTT aggregation in inducible Htt14A2.6 cells and derived cell-free lysates. (a) Percentage of cells with mHTT aggregates following oligomeric 0.1 μM polyQ40 and 0.5 μM of Aβ1–40 or Aβ1–42 seeding; seed condition: [F(3,15.8)= 88.95, P<0.0001]; incubation time: [F(1,27)]=7.79, P=0.0095; interaction: [F(3,15.8)=0.21, P=NS]. (b) Amount of mHTT aggregation on filter trap assay following oligomeric 0.1 μM polyQ40 seeding and 5.0 μM of Aβ1–40, Aβ1–42, α-synuclein (SNCA), and A53T α-synuclein in cell-free lysate assay. Seed condition: [F(5,4.3): 23.95, P=0.003], *P<0.05, **P<0.001. Bars indicate s.e.m. Each experiment had three to nine repeats. mHTT, mutant huntingtin protein; NS, not significant.
Autopsy HD CSF ‘seeds‘ aggregation
Given that media or lysates from induced Htt14A2.6 cells is also sufficient to seed aggregation when transferred to new naïve cell cultures (data not shown), we next determined whether the property of seeding could be extended to human HD CSF. As a first step, immunoreactivity of mHTT was evaluated in postmortem HD CSF samples (Supplementary Table S1) by spotting on filter blots and probing with HTT specific antibodies. HTT was detected in both HD and control CSF samples using a pan-HTT antibody raised against the first 17 amino-acid residues at the N terminus of HTT (N17 antibody, Sigma-Aldrich). Importantly, an antibody specific for expanded repeat HTT (MAB1574, EMD Millipore) was immunoreactive only in HD samples (Figure 4a; Supplementary Table S1), providing evidence that mHTT is indeed present in HD patient CSF and could provide a template for seeding. To next assess whether mHTT in HD CSF could seed aggregation, 10 μl of HD or control CSF samples was added into Htt14A2.6 cell culture medium (990 μl) and incubated for 16 and 24 h. Figure 4b depicts the raw percentage of cells containing visible aggregates following CSF seeding (left panel) and the relative increase in CSF seeding activity over mock control (no CSF). In each case (n=6 experiments), postmortem HD CSF samples significantly enhanced the percentage of cells with aggregates compared with non-HD control CSF.
Postmortem CSF seeding enhancement of mHTT aggregation in inducible Htt14A2.6 cells and derived cell-free lysates. (a) Dot blots detect amount of N-terminal (N17) in both autopsy HD and control CSF samples but expanded polyQ (Q>36) only in HD CSF. (b) Normalized percentage of cells with mHTT aggregates following seeding with CSF from autopsy HD and healthy control subjects (Ctl). Diagnosis: [F(1,87.6)= 120.41, P<0.0001]; incubation time: [F(1,75)=17.16, P<0.0001]; interaction: [F(1,75)=36.35, P<0.0001]. See Figure 5b for individual values. (c) Percentage of cells with mHTT aggregates following seeding with CSF from an autopsy HD subject (HD#134) that has been depleted by polyQ expansion-specific antibodies at dilutions of 1/20, 1/100 and 1/500. Antibody dilution: F(3,7.29)=45.8, P<0.0001. (d) Amount of mHTT aggregates formed in 14A2.6 cell-free lysates seeded with HD CSF of Subject HD#134 at varying dilutions (that is, final CSF concentration=10% in the cell-free lysate standard assay, 2% in ‘1/5‘ and 0.4% in ‘1/25‘ dilutions). (e) Amount of aggregate protein with GFP tag in same subjects in cell-free assay. Diagnosis: [F(1,14.7)=50.79, P<0.0001]. PolyQ: 0.1 μM polyQ40. Bars indicate s.e.m. Each experiment had three to nine repeats. CSF, cerebrospinal fluid; GFP, green fluorescent protein; HD, Huntington’s disease; mHTT, mutant huntingtin protein.
Consistent with the hypothesis that seeding is a mHTT-dependent process, the aggregation seeding potency of CSF from HD subjects was decreased when expanded polyQ-linked mHTT fragments were immunodepleted using expanded polyQ-specific antibody (MAB1574, Figure 4c). Seeding was dose dependent as serial dilutions of HD CSF produced a corresponding decrease in mHTT (Figure 4d), consistent with seeding being dependent on the presence of mHTT. Aggregation by HD CSF was also confirmed using the cell-free lysate assay (Figure 4e). Levels of insoluble mHttex1-GFP aggregates were also measured by filter retardation-GFP immunoblotting (gel under the histogram of Figures 4d and e) and showed a tight correspondence to the seeding propensity.
CSF from BACHD rat distinguishes HD from controls
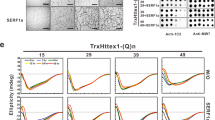
Autopsy CSF may contain substances related to postmortem changes that are not present in CSF from living subjects. To remove potential postmortem confounds, CSF was obtained from BACHD rats that express a full-length human mHTT transgene at 17 months, when rats are symptomatic.24 CSF from five BACHD and five wild-type rats was assayed (Figure 5a; Supplementary Table S2). Each HD sample was clearly distinguished from controls. Control rat CSF did not enhance aggregation above a mock control, whereas BACHD CSF samples enhanced aggregation similar to results observed for polyQ40 oligomer seeds.
Enhanced mHTT aggregation (relative to mock) in blinded CSF samples from living PREDICT subjects. (a) Amount of mHTT aggregates from cell-free lysates seeded with CSF from BACHD transgenic rats (BAC) and wild-type rats (WT). BACHD vs WT: [F(1,15.6)=117.74, P<0.0001]. (b) Relative (to mock control) levels of cells with mHTT aggregates (bars) with GFP tags following 24 h seeding with CSF from blinded PREDICT living samples (bracketed), and autopsy HD and controls (HD#, Ctl#; converted data from Figure 4b), reordered for clarity. HD subjects with clinical motor symptoms in red, CAG expansion-gene-positive subjects without clinical symptoms in yellow, healthy controls in blue. Autopsy HD subjects in light red, and autopsy controls in light blue. Group (HD, CAG expansion-gene-positive, and controls): [F(2,19.6)=37.82, P<0.0001]. HD>CAG expansion-gene-positive>Controls, all Ps<0.001. (c) Amount of aggregate protein with GFP tags following seeding with CSF from blinded PREDICT samples, with autopsy HD (HD#) and polyQ control subjects (upper) and with CSF from 10 subjects with mixed dementia (AD, Lewy body, and frontal temporal; lower; see Supplementry Figure S5A for original gels). (d) Relative amount of mHTT in cell-free lysate assay following seeding with CSF from grouped PREDICT-HD subjects with clinical symptoms (red), expansion-gene-positive subjects without clinical symptoms (yellow), and controls (blue; autopsy and PREDICT controls (CTL); and non-HD dementia controls (data from c)). **P<0.001, ***P<0.0001. Bars indicate s.e.m. Each experiment had three to nine repeats. CSF, cerebrospinal fluid; GFP, green fluorescent protein; HD, Huntington’s disease; mHTT, mutant huntingtin protein.
CSF from living HD subjects ‘seeds‘ aggregation
On the basis of the autopsy and BACHD rat CSF results, we next investigated whether aggregation could be seeded using a panel of CSF samples (blinded) obtained from living PREDICT-HD (Neurobiological Predictors of Huntington’s Disease)25 subjects (diagnosed with HD (n=2), expansion-gene-positive subjects without manifest clinical symptoms (n=7) and healthy control subjects (n=2); Figure 5b; Supplementary Table S3). The figure depicts measurements for PREDICT living subjects as well as postmortem HD and control subjects. The results for living and postmortem subjects are consistent. HD subjects diagnosed by traditional clinical criteria (that is, presence of movement disorder) were distinguished without overlap from the control subjects by the percentage of cells with aggregates and the amount of aggregates in the cell-free lysate assay (Figures 5b and d). Seeding measures from expansion-gene-positive subjects without clinical symptoms revealed a range of seeding values between HD subjects and control values in this relatively small cohort. This range could reflect time to conversion, that is, onset of HD symptoms; however, this assessment will require an additional study. We evaluated the ability of the assays to distinguish clinical stage. The mean percentage of cells with enhanced aggregation in the PREDICT-HD sample was significantly correlated with motor scores (total motor score; r= 0.79; P=0.01) and cognitive dysfunction (Symbol digit modalities test; r= −0.71, P=0.03), but not with putamen volume (r= −0.48; P=0.19). The correlation between percent aggregation and CAG-Age Product (CAP) score indexing disease burden26 was 0.54; for CAG (number of repeats) alone it was 0.34. The CAP score is a better predictor of clinical onset than the CAG expansion repeat as it takes into consideration subject age, and age is a significant predictor of symptom onset. The correlation between CAP score and total motor score is 0.67, P<0.05, and CAP score and symbol digit modalities test is −0.72, P<0.03. The corresponding correlations with the CBA cell-free lysate assay were smaller and did not reach statistical significance, although they did distinguish HD from non-HD controls (Figure 5c). The correlation between the two CBA measures, mean percentage of cells with aggregates and cell-free fold increases, was 0.58. If confirmed in longitudinal studies, these data suggest that a combination of the two CBA assay measurement approaches could ultimately provide unique benefit in the development of molecular biomarker assays for screening efficacy of drugs that reduce mHTT CSF load in HD subjects and possibly in tracking disease stage. These possibilities will be explicitly tested in future studies by testing additional CSF samples and optimized methods of quantification and assessments of therapies such as mHTT lowering treatments.
To evaluate the specificity of seeding in human samples, CSF from 10 subjects with mixed dementia was also tested in the cell-free lysate assay. Similar to the oligomer seeding studies above in which Aβ did not show seeding, none of the mixed dementia CSF showed enhancement of newly formed mHTT aggregates (Figures 5c and d), supporting template dependence and specificity for HD pathogenesis. Supplementary Table S4 provides evidence of nonseeding for non-HD subjects with a wide range of Aβ42, tau and p-tau values.
Discussion
There is an urgent need to develop effective treatments for HD. Reliable molecular biomarkers of disease in patient biofluids has been elusive for the evaluation of potential therapeutics; however, there are emerging possibilities using HD patient CSF.27 Here we provide evidence that a mHTT-specific and disease pathogenic property exists in HD CSF that can be identified using a surrogate assay (that is, CBA seeding) to measure an increased propensity to seed the number of cells having visible aggregates and the amount of aggregation in cell lysates. Importantly, seeding is template specific for mHTT. This CBA seeding activity may be highly relevant to pathogenesis of neurodegenerative diseases based on growing evidence indicating that oligomers of aggregating peptides or proteins such as Aβ, α-synuclein and tau are transmissible between cells,28, 29, 30 and can thereby spread and enhance both homologous and heterologous aggregation through nucleation seeding in a cell-to-cell protein propagation or ‘prion-like‘ manner.28, 29, 30, 31 α-Synuclein and Aβ are detected in CSF and measurements of their abundance or ratio are being considered as sensitive biomarkers for Parkinson’s and Alzheimer’s diseases, respectively.32, 33, 34, 35 More recent work in HD model systems supports cell-to-cell transfer of mHTT aggregation. For instance, mHTT exon 1 protein fused to nonfluorescent halves of a fluorescent protein, which requires dimerization-mediated biomolecular fluorescence complementation to be visualized, provided direct evidence of mHTT transfer between cells.36 Transneuronal propagation of mHTT was also observed using in vitro and in vivo approaches, demonstrating the movement of mHTT from cortical neurons to striatal medium spiny neurons37 and from HD tissue into non-HD fetal neural allografts,38 supporting a direct involvement or consequence of pathogenesis. Here we report that mHTT can be detected in HD CSF using expanded repeat-specific antibodies. This is consistent with a new study showing that soluble and nomomeric HTT can be quantitatively detected in CSF by an ultrasensitive immunoassay.27 Our study demonstrates that exogenous polyQ oligomers and CSF containing mHTT can seed aggregation in our PC12 cell-based models. This supports the idea that a prion-like protein transmission may be involved. This mechanism will be further clarified by evaluating the kinetics of aggregation in future studies. The presence of the protein is required for seeding to occur, potentially supporting a direct relationship between seeding and pathology in HD. Our data is consistent with the observations in Alzheimer’s and Parkinson’s disease that seeding with native protein is much more efficient than synthetic seeds,39 reflecting the likelihood that additional properties present in patient-derived CSF beyond the mHTT protein can also be captured as a modulator of cell-to-cell propagation, including the presence of neuroinflammatory cytokines. Indeed, we find that seeding of aggregation in inducible Htt14A2.6 cells can be further enhanced by cytokines such as interleukin-1β (unpublished results). Given literature demonstrating that aberrant immune activation is observed in HD patient CSF even prior to onset of disease symptoms,40 our finding that patient CSF has aggregation enhancing activity may reflect multiple molecular characteristics of disease.
Our studies demonstrate that a significant increase in newly formed mHTT-containing aggregates was consistently found in Htt14A2.6 cells and cell-free lysates seeded with HD CSF samples but not with CSF from healthy controls without the HD expansion. CSF from expansion-gene-positive subjects who have not yet developed clinical motor symptoms revealed a range of seeding results from HD to non-HD measures, raising the possibility that CSF seeding might provide a measure of HD progression, that is, will those with higher seeding levels be the first to convert to clinical HD? This possibility will be assessed in future studies with longitudinal patient samples from individuals who have since converted to disease. The range of findings are consistent with that seen in clinical and imaging markers showing an insidious increase in clinical symptoms in concert with the progressive loss of regional brain volume.41
The CSF seeding results also suggest that treatments may be effective by blocking the extracellular cell-to-cell transmission of mHTT. This is consistent with the propagation of HD pathology shown by Pecho-Vrieseling37 and Cicchetti,38 although the identification of mHTT aggregates exclusively within the extracellular matrix of fetal allografts in the latter study may be a consequence of alternative mechanisms involving the host immune cells. The implications from these cross-sectional data remain to be confirmed in longitudinal studies of expansion-gene-positive subjects as well as further longitudinal analysis from HD models (for example, BACHD rat model).
Determining the temporal order and the slope of change of biomarkers in the progression of gene-expansion-positive subjects to HD is important for monitoring progression and designing clinical trials that incorporate surrogate markers. The Dominantly Inherited Alzheimer’s Network (DIAN) study data provide a relevant example of having the optimal biomarker at various stages of disease. On the basis of cross-sectional data, Bateman et al.42 proposed a temporal order in which biomarkers become abnormal before the Alzheimer’s disease onset. The most sensitive for early detection (15 years before symptoms) are cerebral deposition of fibrillar amyloid on PET and increased levels of CSF tau protein. However, once clinical dementia onset begins, fluoro-deoxy-glucose PET, hippocampal volume and CSF tau may be most reflective of clinical progression. The choice of biomarkers depends on the stage of illness and the therapeutic target, for example, anti-amyloid or anti-tau. The DIAN study results are applicable to HD biomarkers such as assays that can quantitatively monitor the seeding property identified here in HD CSF. The lack of any overlap in the CBA measures between HD (n=7) and non-HD control (n=17) subjects is noteworthy. Individualized treatment depends on prediction for the individual, not a difference in group means.
Taken together, these data show that an HD-specific property exists in patient CSF that can be monitored in relatively simple cell-based assays. These finding suggest that this CSF-enhanced aggregation property could potentially form the basis for monitoring the stage and course of HD as well as serve as a molecular biomarker for therapeutic interventions that alter levels of mHTT in CSF.
References
Group THsDCR. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell 1993; 72: 971–983.
Lee JM, Ramos EM, Lee JH, Gillis T, Mysore JS, Hayden MR et al. CAG repeat expansion in Huntington disease determines age at onset in a fully dominant fashion. Neurology 2012; 78: 690–695.
Walker FO . Huntington's disease. Lancet 2007; 369: 218–228.
Hatters DM . Putting huntingtin "aggregation" in view with windows into the cellular milieu. Curr Top Med Chem 2012; 12: 2611–2622.
Li SH, Li XJ . Aggregation of N-terminal huntingtin is dependent on the length of its glutamine repeats. Hum Mol Genet 1998; 7: 777–782.
Scherzinger E, Sittler A, Schweiger K, Heiser V, Lurz R, Hasenbank R et al. Self-assembly of polyglutamine-containing huntingtin fragments into amyloid-like fibrils: implications for Huntington's disease pathology. Proc Natl Acad Sci USA 1999; 96: 4604–4609.
Legleiter J, Mitchell E, Lotz GP, Sapp E, Ng C, DiFiglia M et al. Mutant huntingtin fragments form oligomers in a polyglutamine length-dependent manner in vitro and in vivo. J Biol Chem 2010; 285: 14777–14790.
Tsvetkov AS, Arrasate M, Barmada S, Ando DM, Sharma P, Shaby BA et al. Proteostasis of polyglutamine varies among neurons and predicts neurodegeneration. Nat Chem Biol 2013; 9: 586–592.
Shao J, Diamond MI . Polyglutamine diseases: emerging concepts in pathogenesis and therapy. Hum Mol Genet 2007; 16: R115–R123.
Ren PH, Lauckner JE, Kachirskaia I, Heuser JE, Melki R, Kopito RR . Cytoplasmic penetration and persistent infection of mammalian cells by polyglutamine aggregates. Nat Cell Biol 2009; 11: 219–225.
Yang W, Dunlap JR, Andrews RB, Wetzel R . Aggregated polyglutamine peptides delivered to nuclei are toxic to mammalian cells. Hum Mol Genet 2002; 11: 2905–2917.
Apostol BL, Kazantsev A, Raffioni S, Illes K, Pallos J, Bodai L et al. A cell-based assay for aggregation inhibitors as therapeutics of polyglutamine-repeat disease and validation in Drosophila. Proc Natl Acad Sci USA 2003; 100: 5950–5955.
Chen S, Wetzel R . Solubilization and disaggregation of polyglutamine peptides. Protein Sci 2001; 10: 887–891.
Wanker EE, Scherzinger E, Heiser V, Sittler A, Eickhoff H, Lehrach H . Membrane filter assay for detection of amyloid-like polyglutamine-containing protein aggregates. Methods Enzymol 1999; 309: 375–386.
Sontag EM, Joachimiak LA, Tan Z, Tomlinson A, Housman DE, Glabe CG et al. Exogenous delivery of chaperonin subunit fragment ApiCCT1 modulates mutant Huntingtin cellular phenotypes. Proc Natl Acad Sci USA 2013; 110: 3077–3082.
Sontag EM, Lotz GP, Yang G, Sontag CJ, Cummings BJ, Glabe CG et al. Detection of mutant huntingtin aggregation conformers and modulation of SDS-soluble fibrillar oligomers by small molecules. J Huntingtons Dis 2012; 1: 127–140.
Laird NM, Ware JH . Random-effects models for longitudinal data. Biometrics 1982; 38: 963–974.
Kenward MG, Roger JH . Small sample inference for fixed effects from restricted maximum likelihood. Biometrics 1997; 53: 983–997.
Ono K, Takahashi R, Ikeda T, Yamada M . Cross-seeding effects of amyloid beta-protein and alpha-synuclein. J Neurochem 2012; 122: 883–890.
Guo JL, Covell DJ, Daniels JP, Iba M, Stieber A, Zhang B et al. Distinct alpha-synuclein strains differentially promote tau inclusions in neurons. Cell 2013; 154: 103–117.
Hinz J, Gierasch LM, Ignatova Z . Orthogonal cross-seeding: an approach to explore protein aggregates in living cells. Biochemistry 2008; 47: 4196–4200.
Miller Y, Ma B, Tsai CJ, Nussinov R . Hollow core of Alzheimer's Abeta42 amyloid observed by cryoEM is relevant at physiological pH. Proc Natl Acad Sci USA 2010; 107: 14128–14133.
Zhang R, Hu X, Khant H, Ludtke SJ, Chiu W, Schmid MF et al. Interprotofilament interactions between Alzheimer's Abeta1-42 peptides in amyloid fibrils revealed by cryoEM. Proc Natl Acad Sci USA 2009; 106: 4653–4658.
Yu-Taeger L, Petrasch-Parwez E, Osmand AP, Redensek A, Metzger S, Clemens LE et al. A novel BACHD transgenic rat exhibits characteristic neuropathological features of Huntington disease. J Neurosci 2012; 32: 15426–15438.
Paulsen JS, Long JD, Johnson HJ, Aylward EH, Ross CA, Williams JK et al. Clinical and biomarker changes in premanifest huntington disease show trial feasibility: a decade of the PREDICT-HD Study. Front Aging Neurosci 2014; 6: 78.
Zhang Y, Long JD, Mills JA, Warner JH, Lu W, Paulsen JS . Indexing disease progression at study entry with individuals at-risk for Huntington disease. Am J Med Genet B Neuropsychiatr Genet 2011; 156B: 751–763.
Wild EJ, Boggio R, Langbehn D, Robertson N, Haider S, Miller JR et al. Quantification of mutant huntingtin protein in cerebrospinal fluid from Huntington's disease patients. J Clin Invest 2015; 125: 1976–1986.
Holmes BB, Diamond MI . Prion-like properties of tau protein: the importance of extracellular tau as a therapeutic target. J Biol Chem 2014; 289: 19855–19861.
Masuda-Suzukake M, Nonaka T, Hosokawa M, Oikawa T, Arai T, Akiyama H et al. Prion-like spreading of pathological alpha-synuclein in brain. Brain 2013; 136: 1128–1138.
Stohr J, Watts JC, Mensinger ZL, Oehler A, Grillo SK, DeArmond SJ et al. Purified and synthetic Alzheimer's amyloid beta (Abeta) prions. Proc Natl Acad Sci USA 2012; 109: 11025–11030.
Jucker M, Walker LC . Self-propagation of pathogenic protein aggregates in neurodegenerative diseases. Nature 2013; 501: 45–51.
Sierks MR, Chatterjee G, McGraw C, Kasturirangan S, Schulz P, Prasad S . CSF levels of oligomeric alpha-synuclein and beta-amyloid as biomarkers for neurodegenerative disease. Integr Biol 2011; 3: 1188–1196.
Holtta M, Hansson O, Andreasson U, Hertze J, Minthon L, Nagga K et al. Evaluating amyloid-beta oligomers in cerebrospinal fluid as a biomarker for Alzheimer's disease. PLoS One 2013; 8: e66381.
Tokuda T, Qureshi MM, Ardah MT, Varghese S, Shehab SA, Kasai T et al. Detection of elevated levels of alpha-synuclein oligomers in CSF from patients with Parkinson disease. Neurology 2010; 75: 1766–1772.
Fukumoto H, Tokuda T, Kasai T, Ishigami N, Hidaka H, Kondo M et al. High-molecular-weight beta-amyloid oligomers are elevated in cerebrospinal fluid of Alzheimer patients. FASEB J 2010; 24: 2716–2726.
Herrera F, Tenreiro S, Miller-Fleming L, Outeiro TF . Visualization of cell-to-cell transmission of mutant huntingtin oligomers. PLoS Curr 2011; 3: RRN1210.
Pecho-Vrieseling E, Rieker C, Fuchs S, Bleckmann D, Esposito MS, Botta P et al. Transneuronal propagation of mutant huntingtin contributes to non-cell autonomous pathology in neurons. Nat Neurosci 2014; 17: 1064–1072.
Cicchetti F, Lacroix S, Cisbani G, Vallieres N, Saint-Pierre M, St-Amour I et al. Mutant huntingtin is present in neuronal grafts in huntington disease patients. Ann Neurol 2014; 76: 31–42.
Meyer-Luehmann M, Coomaraswamy J, Bolmont T, Kaeser S, Schaefer C, Kilger E et al. Exogenous induction of cerebral beta-amyloidogenesis is governed by agent and host. Science 2006; 313: 1781–1784.
Bjorkqvist M, Wild EJ, Thiele J, Silvestroni A, Andre R, Lahiri N et al. A novel pathogenic pathway of immune activation detectable before clinical onset in Huntington's disease. J Exp Med 2008; 205: 1869–1877.
Paulsen JS, Langbehn DR, Stout JC, Aylward E, Ross CA, Nance M et al. Detection of Huntington’s disease decades before diagnosis: the Predict-HD study. J Neurol Neurosurg Psychiatry 2008; 79: 874–880.
Bateman RJ, Xiong C, Benzinger TL, Fagan AM, Goate A, Fox NC et al. Clinical and biomarker changes in dominantly inherited Alzheimer's disease. N Engl J Med 2012; 367: 795–804.
Acknowledgements
The US National Institutes of Health supported this study through the following grants: Center for Protein Folding Machinery grant 5PN2EY016525 (WC, SGP, LMT, ZT and CG), 5U24RR021992 (SGP), the Undergraduate Research Fellowship (KTP through 5PN2EY016525), Cryo-EM Center Grant 5P41GM10383229 (WC), the PREDICT grant 5R01NS040068 (JSP), 5P50AG016570 (JMR), 5R37GM048071 (IP) and 1U01NS087825 (SGP). The University of California has filed a patent application for the technology discussed in this paper (Diagnostic and Monitoring System for Huntington's Disease, PCT App. No. PCT/US2014/037403; UC ref. UC2013-804-3; Ref. RUC002W). We thank UCLA’s Human Brain and Spinal Fluid Resource Center for the autopsy HD and the Easton Center for Alzheimer’s Disease Research for dementia control CSF samples. Doug Macdonald and Simon Noble from CHDI provided the BACHD CSF samples and corresponding data interpretation. Nguyen (University of Tuebingen) for use of the BACHD TG5 rat model and Christina Gabrysiak (Evotec, AG) for technical assistance in the collection of the BACHD TG5 CSF. We thank Karen H Gylys for overseeing the Aβ, p-tau and t-tau assays on dementia control CSF and Liana G Apostolova for oversight of the Neuroimaging and Biomarkers Core in the Easton Center. We thank Jeremy Bockholt for the bioinformatics support for the PREDICT subjects.
Data deposition: Cryo-electron tomogram of polyQ40 seeded mHttex1-GFP in cell-free lysate derived from Htt14A2.6 cell has been deposited to Electron Microscopy Data Bank (EMDB ID: EMD-2991).
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Tan, Z., Dai, W., van Erp, T. et al. Huntington’s disease cerebrospinal fluid seeds aggregation of mutant huntingtin. Mol Psychiatry 20, 1286–1293 (2015). https://doi.org/10.1038/mp.2015.81
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2015.81
This article is cited by
-
Decoding the Cellular Trafficking of Prion-like Proteins in Neurodegenerative Diseases
Neuroscience Bulletin (2024)
-
Prion-like properties of the mutant huntingtin protein in living organisms: the evidence and the relevance
Molecular Psychiatry (2022)
-
Huntington’s disease: lessons from prion disorders
Journal of Neurology (2021)
-
Disease-related Huntingtin seeding activities in cerebrospinal fluids of Huntington’s disease patients
Scientific Reports (2020)
-
Cerebrospinal fluid from Alzheimer’s disease patients promotes tau aggregation in transgenic mice
Acta Neuropathologica Communications (2019)