Abstract
A single injection of N-methyl-D-aspartate receptor (NMDAR) antagonists produces a rapid antidepressant response. Lasting changes in the synapse structure and composition underlie the effectiveness of these drugs. We recently discovered that rapid antidepressants cause a shift in the γ-aminobutyric acid receptor (GABABR) signaling pathway, such that GABABR activation shifts from opening inwardly rectifiying potassium channels (Kir/GIRK) to increasing resting dendritic calcium signal and mammalian Target of Rapamycin activity. However, little is known about the molecular and biochemical mechanisms that initiate this shift. Herein, we show that GABABR signaling to Kir3 (GIRK) channels decreases with NMDAR blockade. Blocking NMDAR signaling stabilizes the adaptor protein 14-3-3η, which decouples GABABR signaling from Kir3 and is required for the rapid antidepressant efficacy. Consistent with these results, we find that key proteins involved in GABABR signaling bidirectionally change in a depression model and with rapid antidepressants. In socially defeated rodents, a model for depression, GABABR and 14-3-3η levels decrease in the hippocampus. The NMDAR antagonists AP5 and Ro-25-6981, acting as rapid antidepressants, increase GABABR and 14-3-3η expression and decrease Kir3.2. Taken together, these data suggest that the shift in GABABR function requires a loss of GABABR-Kir3 channel activity mediated by 14-3-3η. Our findings support a central role for 14-3-3η in the efficacy of rapid antidepressants and define a critical molecular mechanism for activity-dependent alterations in GABABR signaling.
Similar content being viewed by others
Introduction
Major depressive disorder (MDD) is the second largest contributing factor to disability in developed countries.1 In the United States, ~20% of the population will experience an MDD episode in their lifetimes.2 Serotonergic- and adrenergic-based antidepressants, typically prescribed to treat MDD, require 2–4 weeks of use before relief of symptoms occur.3 This lag time, unfortunately, presents a vulnerability to suicide.4 Furthermore, these medications are not effective in ~20% of patients suffering from MDD.5, 6, 7, 8, 9, 10 Thus developing new antidepressants that act rapidly and provide relief to individuals with treatment-resistant depression is vital. Clinical studies show that ketamine acts rapidly (within days or hours) and is efficacious in treatment-resistant depression.11, 12, 13 However, ketamine has the potential for abuse, tempering enthusiasm for its clinical use. Research both in animal models and clinical studies suggests that the antidepressant efficacy of ketamine arises from its blockade of N-methyl-D-aspartate receptors (NMDARs).14, 15, 16 Given the clinical promise of NMDAR antagonists, characterizing the molecular mechanisms that underlie the effectiveness of NMDAR blockers is a timely and critical step toward the development of safer and more effective antidepressants.
γ-Aminobutyric acid (GABA) is the major inhibitory neurotransmitter in the brain. Metabotropic GABAB receptors (GABABRs) are G-protein coupled receptors that mediate slow synaptic inhibition by increasing outward potassium current through inwardly rectifying potassium channels (GIRK/Kir3.X).17 The role of GABABR signaling in MDD is beginning to emerge.18 We recently demonstrated that GABABRs are necessary for the rapid antidepressant efficacy of NMDAR antagonists.19 Contrary to GABABRs’ established function, NMDAR blockade promotes a shift in GABABR signaling such that activation of the receptor increases resting dendritic calcium in a manner that requires L-type Ca2+ channels. The rise in dendritic calcium activates hotspots of mammalian Target of Rapamycin (mTOR) kinase activity in the dendrites. mTOR activity promotes the syntheses of plasticity-related proteins, a necessary step for the initiation and sustainment of rapid antidepressant efficacy of NMDAR antagonists (herein referred to as the GABABR shift in function).20 The present study examines the molecular events upstream of the shift in GABABR function by asking: (1) are dynamic changes in dendritic protein expression required for the shift in GABABR signaling, and (2) how is the inhibitory component of GABABR signaling changed with NMDAR antagonists?
Herein, we describe a mechanism by which NMDAR antagonists cause GABABRs to decouple from their primary inhibitory target, Kir3 channels. Intraperitoneal (i.p.) injection of an NMDAR antagonist, D-(-)-2-amino-5-phosphonopentanoic acid (AP5) or Ro-25-6981, induces rapid changes in protein expression in vivo to favor the shift in GABABR signaling such that the protein levels of Kir3.2 (GIRK) decrease and GABABR2 increase in the hippocampus through mRNA translation. 14-3-3η is an adaptor protein that regulates GABABR coupling to Kir3.X channels21, 22 and is implicated in diseases in which NMDAR signaling is dysregulated.23 We show that 14-3-3η expression increases with acute NMDAR antagonism and decreases in socially defeated rodents, a model of depression. NMDAR blockade unexpectedly slows both protein synthesis and degradation of 14-3-3η, elevating total levels of 14-3-3η. Blocking the functional interaction between 14-3-3η with a dominant-negative form of 14-3-3η (DN-14-3-3 η) restores GABABR-mediated inhibition in the presence of NMDAR blockade. Moreover, stereotaxic injection of a virus coding for DN-14-3-3η prevents the rapid antidepressant efficacy of AP5 and Ro-25-6981. Our findings altogether suggest that removing GABABR-Kir3 activity may be the initial step required for the shift in GABABR function that is necessary for the efficacy of rapid antidepressants.
Materials and methods
Isolation of synaptoneurosomes (SNs) and neuronal lysates
SNs were isolated from day in vitro (DIV )14–18 cortical cultures or from rat or mouse prefrontal cortex and hippocampus by a modified method previously described.24 Briefly, neurons were harvested in buffer B (20 mM HEPES (4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid), pH 7.4; 5 mM EDTA, pH 8.0; protease inhibitor cocktail (Complete, Roche, Indianapolis, IN, USA); phosphatase inhibitor) and homogenized. Homogenate was filtered first through a sterile 100-μm nylon filter followed by a 5-μm filter. SNs were pelleted at 14 000 g for 20 min at 4 °C. SN pellet was solubilized with radio-immunoprecipitation assay buffer (150 mM NaCl; 10 mM Tris, pH 7.4; 0.1% SDS (sodium dodecyl sulfate); 1% Triton X-100; 1% deoxycholate; 5 mM EDTA; protease inhibitor cocktail tablet and phosphatase inhibitor) for 20 min on ice. The insoluble fraction was then pelleted at 14 000 g for 20 min at 4 °C, and the supernatant was used for immunoblot analysis.
Western blottings
Equal amounts of SN (25 μg) sample in radio-immunoprecipitation assay (RIPA) buffer and SDS loading buffer were run on a 10% SDS-polyacrylamide gel and transferred to a nitrocellulose membrane for 1 h in 20% transfer buffer (25 mM Tris, 192 mM glycine, 20% methanol). Membranes were then blocked for 1 h at room temperature (RT) while shaking in TBST (TBS and 0.1% Tween-20) with 5% nonfat milk. Membranes were incubated overnight (O/N) at 4 °C in primary antibody and for 45 min at RT in secondary antibody. After each incubation step, membranes were washed for 6 × 10 min in TBST. Membranes were imaged with the Odyssey immunoblot software (Lincoln, NE, USA). For 14-3-3η blots, membranes were probed by enhanced chemiluminescence. ImageJ (Bethesda, MD, USA) was used for densitometry analysis. The antibodies used were as follows: GABABR1 (AB55051, Abcam, Cambridge, UK), GABABR2 (Neuromab, Davis, CA, USA, N81/2), Kir3.2 (Alomone, Jerusalem, Isreal, APC-006), 14-3-3η (sc-17286, Santa Cruz, Dallas, TX, USA), and α-tubulin (ab15246, Abcam, Cambridge, UK).
Cloning of 14-3-3η
14-3-3η was cloned by RT-PCR from rat brain cDNA primers 5′-ATGGGGGACCGAGAGCAGCTG-3′ and 5′-GCGCACATCATAATTTAAAGAGGCACAGC-3′ amplifying nucleotides 176–1446 (Genbank accession no.: D17445). The remaining 3′ untranslated region (UTR) was cloned by sequential ligations of annealed oligonucleotides (1447–1689). The DN construct was made by site-directed mutagenesis replacing the CGA with GCA at amino acid 56 and the AGG with a GCG at amino acid 60 to replace the arginine to an alanine.25 Primers used for recombinant adeno-associated virus (rAAV) insert are:
Forward primer: 5′-GAGCTCGCCACCATGGGGGACCGAGAGCAGCTGCTCCAGCGGGCGCGAC-3′ and
Reverse primer: 5′-CTGCAGTCAGTTGCCTTCTCCGGCTTCTTCATCC-3′
Kaede was initially cloned into the EGFP-N1 vector (Clontech, Mountain View, CA, USA) by replacing the EGFP cDNA inserted at the BamH1 and Not I (blunt) sites. Kaede-14-3-3η was generated by PCR using a primer specific for the EGFP-N1 vector in the forward direction (5′-AGGCGTGTACGGTGGGAGGTCTATATA-3′) and a primer that contained the last 15 nucleotides of the cDNA before the stop codon, of Kaede, with the addition of a Nhe I site at the 3′ end (Kaede: 5′-CCCGGCCGCTAGCCTTGACGTTGTCC-3′) in the reverse direction. The PCR product was digested with Nhe I and inserted into a Nhe I site of the14-3-3η cDNA at the start codon that was previously engineered into the sequence by the Quick Change Site Directed Mutagenesis Kit (Stratagene, La Jolla, CA, USA). Kaede-14-3-3η 3′UTR cDNA was then subcloned into the StuI site of the SinRep5 virus vector (Invitrogen, Carlsbad, CA, USA), and pseudovirions were produced according to the manufacturer’s directions.
Adeno-associated viral vectors
The DN-14-3-3η and tdTomato proteins26 were cloned into separate adeno-associated viral vectors containing a mouse synapsin promoter, a woodchuck posttranscriptional regulatory element and SV40 poly-adenylation sequence between flanking AAV2 inverted terminal repeats.27 rAAVs were assembled using a modified helper-free system (Stratagene) as serotype 2/1 (rep/cap genes) viruses and harvested and purified over sequential cesium chloride gradients as previously described.28 Viral titers were >1 × 109 infectious particles per microliter.
Neuronal cultures and transgene expression
Primary neurons were prepared as previously described by Ma et al.29 For DN-14-3-3η and tdTomato co-infections, rAAV(DN-14-3-3η):rAAV(tdTomato) ratio was 4:1. For tdTomato controls, rAAV(tdTomato) was diluted 1:5 with phosphate-buffered saline (PBS).14-3-3η-Kaede in Sindbis was infected 24–36 h prior to live imaging. For imaging experiments, hippocampal neurons were plated at 50–75 000 cells/12 mm coverslip and used at DIV 14–18. Transfections were performed on DIV 12–14 hippocampal neurons using 0.4 μg DNA according to the manufacturer’s instructions (Invitrogen), but in Neurobasal media (Invitrogen), and returned to conditioned media 4–6 h posttransfection. Kaede and Mermaid cDNA was a kind gift from Dr Atsushi Miyawaki at the Riken Brain Institute, Japan. rAAV infections were performed using 1 μl viral mix per coverslip.
Pharmacology
Drug treatment was done in media. All live-imaging experiments were done in HEPES-based artificial cerebral spinal fluid (aCSF (in mM): 100 NaCl, 10 HEPES (pH 7.4), 3 KCl, 2 CaCl2, 1 MgCl2, 10 glucose) adjusted to match the osmolarity of cell culture media. For in vitro manipulations, neurons were pretreated with the NMDAR antagonist AP5 (50 mM, D-AP5; aCSF+AP5) for 90 min. For GABABR activation, neurons were treated with baclofen (50 mM, 5 min). To block Kir3 channels, tertiapin-Q (50 nM, 2 min) was applied. KCl (25 mM) was used to depolarize neurons. All drugs were purchased from Tocris (Bristol, UK).
Microscopy
Images were acquired with a Leica SP5 disk confocal microscope (Wetzlar, Germany) using an oil-immersion × 63 lens or a × 63 water immersion lens for live imaging. For co-localizaton assays, a 2048 × 2048 frame was sampled at the Nyquist rate (~50 nm per pixel), 400 Hz for 2-dimensional co-localization analysis. For surface expression, bioorthogonal noncanonical amino acid tagging and proximity ligation assay (BONCAT–PLA) new translation imaging and immunostaining, a 6-μm Z-stack of 1024 × 1024 pixels was obtained using a scan rate of 400 Hz. Max projections of the stack were used for analysis.
Immunofluorescence
For cultured neurons, cells were fixed in 4% paraformaldehyde with 4% sucrose for 20 min at RT, permeabilized with 0.25% Triton X-100 in PBS (PBS) and then blocked in PBS with 8% goat serum for at least 30 min at RT. Primary and secondary antibody incubations were done in blocking buffer O/N at 4 °C and 2 h at RT, respectively. Between incubation steps, neurons were washed 3 × 10 min with PBS. For surface staining, neurons were fixed in 4% paraformaldehyde with 4% sucrose for 20 min on ice and blocked in PBS with 8% goat serum for at least 30 min at RT. Surface primary antibody incubation was completed in blocking buffer O/N at 4 °C. After washing 3 × 10 min with PBS, the cells were permeabilized, and total antibody was probed as above. The following antibodies were used: surface GABABR1 (AB55051, Abcam), and total GABABR1 (SC-14006, Santa Cruz), GABABR2 (Neuromab, N81/2), and Kir3.2 (Alomone APC-006).
Live labeling of surface GABABRs
Live labeling was performed using a modified method from Hayashi et al.30 DIV 14–20 hippocampal neurons were incubated in 3% bovine serum albumin aCSF with surface GABABR1 antibody for 1 h. Following two rinses with bovine serum albumin aCSF, neurons were returned to conditioned media and incubated with AP5 (50 μM) or vehicle (H2O) for 90 min. Neurons were then fixed in 4% paraformaldehyde and stained as above. The following antibodies were used: surface GABABR1 (AB55051, Abcam), and Kir3.2 (Alomone APC-006).
BONCAT–PLA for protein-specific new translation
BONCAT was performed using a method similar to that described by Dieterich et al.31, 32, 33 and using the Click-iT Metabolic Labeling AHA and Reaction Buffer Kit (Life Technologies, Carlsbad, CA, USA). Coverslips were moved to a methionine free media, aCSF, for 30 min at 37 °C. Drug (AP5, 50 μM or vehicle, H2O) and azide-linked non-canonical amino acid, azidohomoalanine (AHA) were then spiked into the media, and cells were incubated for 1 h at 37 °C. Following this, Click-iT Metabolic Labeling was performed according to the manufacturer’s instructions using a biotin-alkyne to tag the newly synthesized proteins. Coverslips were then blocked and incubated in primary antibody O/N at 4 °C. The following day, the PLA secondary staining was performed using the Duolink Kit according to the manufacturer’s instructions (Duolink, Sigma). Antibodies used include: GABABR1 (Santa Cruz, sc-14006), GABABR2 (Neuromab, N81/2), 14-3-3η (sc-17286, Santa Cruz), biotin/α-rabbit (Sigma, St. Louis, MO, USA, SAB3700857) and biotin/α-mouse (Abcam, ab79111), and MAP2 (Aves, Tigard, OR, USA, MAP).
Live imaging
Live-imaging experiments were done in aCSF in a 35-mm culture dish. Single plane images of 512 × 512 pixels were collected at a rate of 400 Hz for 1 min during each acquisition period for mermaid imaging. For imaging of protein stability using Kaede, a Z-stack of 1024 × 1024 pixels was obtained using a scan rate of 400 Hz stacks for the full extent of each neuron used (12–15 μm). Laser intensity, exposure time and image size were held constant within each experiment to allow for comparison.
Live mermaid (voltage) imaging
All live-imaging experiments were done as previously described.19 Mermaid-transfected neurons were incubated with vehicle (H2O) or AP5 for 90 min and washed three times in aCSF or aCSF+AP5 before image acquisition. Measurements are reported as a ratio (R) of the acceptor (red) to donor (green) fluorescence, where an increase in the ratio indicates depolarization and a decrease indicates hyperpolarization.34 The baseline ratio (R0) was established by taking one image every 5 s for 1 min. All drugs were allowed to equilibrate for 2–5 minutes prior to measuring the fluorescence during the next 1-min imaging period (RDrug). Cumulative data are expressed as ΔR/R0. Kir3 channels were blocked with tertiapin-Q (50 nM, 2 min). All drugs were purchased from Tocris.
Live imaging of protein stability
Live imaging of 14-3-3η-Kaede was performed similar to Raab-Graham et al.36 and Sosanya et al.35 Coverslips were infected with a Sindbis virus coding for 14-3-3η-Kaede. 24 to 36 h later, coverslips were washed three times with aCSF and moved to the imaging chamber. An initial Z-stack of the full volume of the neuron (12–18 μm) was acquired (pre-ultraviolet (pre-UV)). Kaede protein was then converted from green to red using UV light for 1 min. Another Z-stack was acquired for the t=0 time point (F0), after which AP5 (50 μM) or Vehicle (H2O) was applied to the bath. A Z-stack was acquired every 15 min for 1 h (F15, F30, F45, F60) after the addition of AP5 or H2O.
Image analysis
For co-localization analysis, the full visible extent of each region of interest (ROI) within a primary dendrite was boxed in a 4 × 4 μm2 area. These ROIs were then analyzed using the JaCOP plugin to determine the Pearson coefficient.37 Thresholds were set according to the computer-generated value for each ROI.
For intensity and punctal analyses, ROIs were demarcated manually by tracing the full visible extent of dendrites that extend out at least 60 μm from the cell body. Background was subtracted by determining the signal in a region close to the dendrite but void of all processes. For dendritic intensity, the average intensity value for the full extent of each dendrite is reported. For surface staining, the average ratio of surface GABABR1 intensity to total GABABR1 intensity is reported. For BONCAT–PLA analysis, the PLA intensity values for each protein are reported as a ratio of the average PLA intensity to map2 intensity. For punctal analysis, puncta were defined as pixel values within each dendritic ROI where the signal intensity for Kir3.2 exceeded the mean signal intensity plus two standard deviations for untreated, control neurons. Non-zero values were then totaled to determine the total number of hotspots and plotted as a function of distance from the soma to visualize areas of increased Kir3.2 signal.
Live-imaging analysis
For live imaging of protein stability using 14-3-3η-Kaede, the average intensity for each fluorophore was calculated for 25-μm ROIs that began 10 μm from the cell body and continued consecutively for the full extent of the dendrite for each time point. The change in intensity was then calculated as a percentage of change from t=0 time point (ΔF=(F60−F0)/F0) for both old/red protein and new/green protein.
For Mermaid voltage imaging, the fluorescence resonance energy transfer (FRET) signal was analyzed by defining an ROI that started ~5 μm from the cell body and extended to the first visible branch point. The ROI included the entire visible portion of the dendrite. Neurons were background subtracted, and intensity (red/acceptor and green/donor) values for each fluorophore at each time point were determined using ImageJ. The acceptor:donor ratio (red/acceptor:green/donor intensity) for each time point in the first imaging period were averaged as the baseline (R0). The ROI ratio values obtained for each time point after the baclofen incubation were averaged (R or RBAC in Figure 3). The ROI ratio values obtained for each time point after the Tertiapin-Q/GIRK blocker incubation were averaged (R or RBACTQ). To calculate the change in signal, the following equations were used: (1) for application of one drug ΔR/R=((R−R0)/R0) was used and (2) to calculate the change in fluorescence following Tertiapin-Q/Kir 3 blocker after application of baclofen, ΔR/R=((RBACTQ−RBAC)/RBAC) was used. The acceptor:donor ratio values at each time point was also averaged and plotted as a percentage of the baseline.
Stereotaxic virus injections
All procedures were performed in accordance with the National Institutes of Health’s Guide for the Care and Use of Laboratory Animals and approved by The University of Texas at Austin Institutional Animal Care and Use Committee using sterile technique. The animals were anesthetized using inhaled isoflurane in O2 and fixed in a stereotaxic frame using non-perforating ear bars. The surgical site was prepared by clipping the hair on the scalp and sterilizing the scalp with 70% ethanol. A 5–10-mm incision was made to expose the skull. A 1-mm hole was then drilled at predetermined coordinates using an F/G dental drill bit. A Nanoject II microinjector (Drummond Scientific, Broomall, PA, USA) fitted with a pulled-glass injection pipette (5–10-μm tip diameter) containing the viral suspension was inserted into the brain using a stereotaxic manipulator (Sutter Instruments, Novato, CA, USA). Injection coordinates for the dorsal hippocampal CA1 (from bregma) were −2.2 mm A/P, +/− 1.5 mm M/L; −2.5 mm A/P, and +/− 1.6 mm M/L. Thirty nanoliters of virus was injected at depths of −1.2, −1.1 and −1.0 mm from pia at each site.
In vivo injections
I.p. injections of AP5 (2.5 mg kg−1) or Ro-25-6981 (10 mg kg−1) were administered to 20–30-day-old Sprague-Dawley male rats and 6–10-week-old mice for behavioral analyses (forced swim test (FST), tail suspension test (TST) and splash test) and biochemistry. Hippocampi were harvested 45 min postinjection for immunoblot analysis. Behavioral testing was completed in uninjected naive and rAAV (tdTomato+vector or tdTomato+DN-14-3-3η-Flag)-injected 8–10-week-old mice.
Behavioral testing
FST was performed using a method similar to Porsolt et al.38 Mice were injected with drug or vehicle and 45 min later subjected to the FST. Immobility was scored blindly the last 4 of 6 min of the session. Immobility was defined as no discernible movement past the movement necessary to keep its head above water. A training session was conducted for mice injected with Ro-25-6891 2 h prior to injections.
TST was completed 24 h postinjection as described in Can et al.39 Mice were scored for the struggling the entire 6 min of the test. Mice were filmed for the entire session and scored blindly for mobility. Mobility was defined as any movement of the mouse but not movement due to oscillations or swing resulting from earlier movement.
Splash test was assessed approximately 30 min after the TST. Grooming frequency was assessed for 5 min after 200 μl of 10% sucrose was squirted on the dorsal coat of the mouse.40, 41, 42 Grooming was defined as any behavior in which a mouse licked its fur, groomed with its forepaws and scratched with any limb.
Social defeat
Vasectomized 12-week-old male Sprague-Dawley rats (Harlan, Houston, TX, USA) were selected for aggressive behavior by assessing those males that displayed aggressive grooming, pinning or biting towards the probe rat within 1 min of placement into the resident cage. These animals were used for multiple rounds of social defeat. Female cage mates were rotated among residents. Five-week-old male Sprague-Dawleys (Harlan) were housed in groups of 2–3 and were utilized as the intruders for the stress paradigm.
Residents and intruders were taken to a behavioral procedure room in their home cages at the end of the dark cycle (0700 hours). The room was lit with red light to encourage activity. Females were removed from residents’ cages to a holding cage. Intruders were introduced to residents’ home cages, and the latency to display of aggressive behavior by the resident (aggressive grooming, pinning, or biting) was recorded. After 5 min of direct contact, a perforated Plexiglass barrier was inserted to physically separate animals while maintaining sensory contact for an additional 25 min. After the defeat session, all animals were returned to their home cages and taken back to the colony, where residents and intruders were examined for injuries. For repeated social defeat, one defeat session occurred daily for 5 days, and intruders were exposed to a different resident each session.
Statistical analysis
All statistical analyses were performed using the Graphpad Prism software (La Jolla, CA, USA). For all experiments: significance was calculated at α=0.05 level. For both one-way and two-way analysis of variance, significance is represented by: *P<0.05, **P<0.01, ***P<0.001; and NS indicates no significant difference. For Figure 2c, significance (#) was calculated with a one-sample T-test where μ=0. Error bars represent s.e.m.
Results
Rapid antidepressants are NMDAR antagonists that increase synaptic efficacy by affecting key regulators of protein synthesis.19, 20, 43, 44 Rapid antidepressant efficacy requires GABABR activation of mTOR-dependent protein synthesis in dendrites.19 It is unclear if the events prior to the shift in GABABR require new protein synthesis. Therefore we independently tested two NMDAR antagonists, AP5 and Ro-25-6981. First, we determined whether these NMDAR antagonists produced a behavioral phenotype consistent with antidepressant efficacy. AP-7, an NMDAR antagonist similar in structure, selectivity and potency to AP5, has demonstrated rapid antidepressant properties.45 Ro-25-6981, which blocks GluN2B-containing NMDARs, produces rapid antidepressant effects in animal models and is being tested clinically for MDD.15 Few studies suggest that AP5 may not cross the blood–brain barrier;46 however, many report its effectiveness in the central nervous system following peripheral administration.47, 48, 49, 50, 51, 52, 53, 54 To compare the two NMDAR antagonists, we i.p. injected mice with either AP5 or Ro-25-6981 and subsequently measured their behavior during the FST. FST is a well-established measure of antidepressant efficacy, requiring rodents to swim without escape and eliciting an immobile response.55, 56 Treatment with antidepressants reduces the length of immobility induced by FST (that is, the animals swim longer).55 Similar to Ro-25-6981, AP5 reduced immobility time by ~40 s at 45 min and ~50 s at 24 h post-i.p. injection (Figures 1a and b, Supplementary Figure S1A; 45 min: Control, 140±6 s; AP5, 97±4 s, P<0.005; Figure 1b; Control, 146±13 s; Ro-25-6981, 76±18 s, P<0.02). These results demonstrate that a single i.p. injection of AP5 produces antidepressant behavioral effects, similar to other NMDAR antagonists.19, 20, 45
Bidirectional regulation of the γ-aminobutyric acid receptor (GABABR) pathway by N-methyl-D-aspartate receptor (NMDAR) antagonists, D-(-)-2-amino-5-phosphonopentanoic acid (AP5) and Ro-25-6981. Compared with control, (a) AP5 (2.5 mg kg−1, intraperitoneal (i.p.)) or (b) Ro-25-6981 (10 mg kg−1, i.p.) reduced immobility time by ~40 s at 45 min postinjection. N=3–4 animals. Representative western blottings and quantification of (c and f) Kir3.2, (d and g) GABABR1 and (e and h) GABABR2 show that NMDAR antagonists decrease Kir3 expression, whereas increasing GABABR2 expression in hippocampal synaptoneurosomes (SN) of rats i.p. injected with (c–e) AP5 or (f–h) Ro-25-6981 compared with vehicle. Tissue harvested 45 min postinjection. N=4 animals. (i–l) AP5 does not change (i and j) GABABR1 levels, but increases translation of (k and l) R2 using bioorthogonal noncanonical amino acid tagging and proximity ligation assay. N=10 neurons. Scale bar=25 μm. (m–o) Social defeat does not change (m) Kir3.2 levels but significantly decreases expression of (n) GABABR1 and (o) GABABR2 in hippocampal SN harvested at 36 h after last defeat session. N=8 animals. Significance assessed by Student’s T-test.
NMDAR antagonists reduce Kir3.2 and correspondingly increase GABABR2 levels
We next asked whether GABABR coupling to Kir3 channels changes with NMDAR antagonism. We have previously determined that NMDAR antagonists increase the surface expression of GABABR1; however, it is unclear whether this elevated surface expression requires an increase in total protein. GABABR is an obligate heteromultimer, requiring the assembly of GABABR1 and R2 to make a functional receptor.57 Activation of the GABABR releases the βγ subunits of its associated trimeric G protein, consequently opening Kir3 channels.58 The extent of GABABR-Kir3 coupling could vary with the expression levels of Kir3.2, GABABR1 and R2. Thus we examined the protein levels of Kir3.2, GABABR1 and R2 at 45 min postinjection in the hippocampal synaptoneurosomes (SN, presynaptic and postsynaptic nerve endings) of Control (Vehicle, saline) and AP5 (2.5 mg kg−1) or Ro-25-6981 (10 mg kg−1) mice. Western blotting analysis revealed that AP5 reduced total Kir3.2 expression by ~65% compared with vehicle (Figure 1c; Kir3.2: 0.35±0.04, P<0.02, relative to Control), complementing the report that NMDAR activity stabilizes Kir3.2 surface expression.59 Though GABABR1 expression did not increase at 45 min (Figure 1d), AP5 increased GABABR2 levels by 20% (Figure 1e; R2: 1.25±0.04, P<0.01, relative to Control). These findings were consistent with Ro-25-6981 injection (Figures 1f–h; Kir3.2 0.69±0.05, P<0.01; R1: 1.13±0.19; R2: 1.47±.0.1, P<0.01; relative to Control). GABABR1 and GABABR2 levels were elevated at 2 h postinjection of Ro-25-6981 (Supplementary Figures S1B and C; R1: 1.6±0.20, P<0.03; R2: 1.43±0.05, P<0.04 relative to Control). Taken together, NMDAR antagonists alter protein expression that may favor the shift in GABABR function.
NMDAR antagonism increases GABABR2 expression through new protein synthesis
An increase in GABABR1 and R2 expression with NMDAR blockade may occur through new protein synthesis. We measured new translation of GABABR1 or GABABR2 during NMDAR blockade using BONCAT in cultured hippocampal neurons.31, 32, 33 BONCAT incorporates an AHA-tagged non-canonical amino acid by utilizing an alkyne-azide reaction to label newly synthesized proteins with biotin. To detect new GABABR1 or GABABR2 protein specifically, we subsequently performed a proximity ligation assay (PLA) on BONCAT-treated neurons. PLA emits a fluorescent signal only when two antibodies (anti-biotin and anti-GABABR1 or -GABABR2) are within 30–40 nm of each other. The combination of these methods stringently identifies newly translated GABABR1 or GABABR2. NMDAR blockade by AP5 significantly increased newly synthesized dendritic GABABR2 protein by ~60–70% but not GABABR1 (Figures 1i–l; (0–100 μm: R1: 0.97±0.27; R2: 1.70±0.28, P<0.01); (100–150 μm: R1: 1.04±0.34; R2: 1.63±0.32, P<0.01); relative to Control). These results demonstrate that NMDAR inhibition can induce rapid protein synthesis and that the elevated GABABR2 levels observed during NMDAR blockade may be through new mRNA translation.
GABABR levels are reduced in a rat model of depression
Depression models seek to mimic the physical or emotional stresses that may alter normal molecular signaling and lead to depression or other mood disorders.60, 61 Therefore, we hypothesized that social defeat, a depression model, downregulates GABABR and upregulates Kir3 channel expression, consistent with a bidirectional regulation of the pathway.3, 62 In the social defeat model of depression, experimental rats are placed in the home cage of a novel, aggressive rat for 30 min day−1, including 5 min of direct contact for 5 days.63, 64, 65 To assess the effect of social defeat on the GABABR pathway, we isolated hippocampi from socially defeated and control rats 36 h after their last encounter with the aggressor.64 Western blotting analysis showed that rats exposed to social defeat have significantly lower levels of GABABR1 and GABABR2 relative to controls, while Kir3.2 levels remained unchanged in hippocampal SNs (Figures 1m–o, Supplementary Figures S1D–F; Kir3.2: 0.99±0.10; R1: 0.79±0.04, P<0.009; R2: 0.75±0.04, P<0.004; relative to Control). These data suggest that rapidly altering GABABR2 and Kir3.2 expression may favor the rapid shift in GABABR function, as these changes coincide with the earliest time point assayed for rapid antidepressant efficacy. In contrast, an increase in GABABR1 may be required for long-lasting effects with NMDAR antagonists.
Blocking NMDARs reduces GABABR–Kir3.2 co-localization
As NMDAR antagonism reduces total Kir3.2 expression but increases GABABR2 expression, we asked whether co-localization between GABABR and Kir3.2 decreases. In contrast to the increase in new GABABR2 protein and consistent with what we observe in vivo (Figure 1), Kir3.2 expression levels measured by puncta number and intensity decrease with acute AP5 treatment (Supplementary Figures S2A and B). We previously demonstrated that the surface expression of GABABR1 increases with NMDAR blockade.19 Thus a reduction in co-localization between GABABRs and Kir3 channels may occur because of (1) an increase in the number of surface GABABRs or (2) a decoupling of GABABR and Kir3 channels (Figure 2a). To examine whether NMDAR blockade affects GABABR coupling to Kir3 channels in dendrites, we used a live-labeling protocol that allows us to specifically mark the population of GABAB surface receptors present on the membrane prior to AP5 exposure (Figure 2a, old receptors coupled to Kir3 channels are pink).30 After antibody feeding to live cells, we applied AP5 for 90 min and fixed the cells for immunocytochemistry to detect Kir3.2 co-localization with the labeled, surface GABABRs (pink receptors, Figure 2a). In AP5-treated neurons, GABABR co-localization with Kir3.2 decreased by ~34% relative to vehicle-treated neurons (Figures 2b and c; 0.66±0.05; relative to Control). These results demonstrate that NMDAR antagonism reduces GABABR coupling to Kir3 channels.
Blockade of N-methyl-D-aspartate receptors (NMDARs) reduces co-localization between γ-aminobutyric acid receptor (GABABR) and Kir3.2 and GABABR activation of Kir3. (a) Model showing how D-(-)-2-amino-5-phosphonopentanoic acid (AP5) may affect GABABR co-localization with Kir3.2. (Left) In control condition, GABABR activation increases Kir3 activity. (Right, top panel) (1): Possible mechanism if co-localization does not change with NMDAR blockade. (Right, bottom panel) (2): Possible mechanism if NMDAR blockade reduces the co-localization of GABABRs and Kir3 channels. (b) Representative images and (c) quantification demonstrating that AP5 significantly decreases the percentage of GABABR co-localized with Kir3.2 relative to vehicle-treated neurons. White dots indicate co-localized points. Scale=10 μm. N=10–12 neurons. (d) Averaged traces and (e) histogram show baclofen negligibly affects membrane potential in AP5-treated neurons. (f) Averaged summary trace shows NMDAR-blocked neurons exhibit no detectable Kir3/GIRK response to its specific channel blocker tertiapin-Q (tert-Q), unlike control neurons that exhibit a small but significant depolarization. (g) Summary graph shows relative response to baclofen and baclofen+tert-Q in control- and AP5-treated neurons. N=3 independent cultures. Pearson’s correlation coefficient used to assess the percentage of co-localization in panel (c). # Indicates significance from zero. * Indicates significance between treatments using Tukey’s one-way analysis of variance (ANOVA) in panel (e), Student’s T-test in panel (c) and Bonferroni’s two-way ANOVA in panel (g).
Blocking NMDARs decreases GABABR-induced hyperpolarization mediated by Kir3/GIRK in the dendrites
As AP5 reduced GABABR–Kir3.2 co-localization, we suspected that GABABR-mediated hyperpolarization decreases in the dendrites. Unlike in vivo, NMDAR antagonists in culture require exogenous GABABR activation to increase resting dendritic calcium and mTOR kinase activity.19, 20 This difference allows us to separate activation from downstream signaling, an event that is indistinguishable in vivo due to endogenous GABA.19 Using Mermaid, a voltage-sensitive genetic FRET sensor, we measured the relative changes in membrane potential within the primary dendrite, where the shift in GABABR function is localized.19, 34 Although fluorescent sensors are slow,66 the shift in GABABR function with NMDAR blockade occurs 3–5 min after AP5 application, increases resting dendritic calcium signal that lasts at least 5 min and requires L-type calcium channels.19 Accordingly, Mermaid is an appropriate tool to visualize site-specific changes in dendritic membrane potential.66 An increase in FRET signal indicates depolarization (Supplementary Figures S3C and D, Supplementary Methods), whereas a decrease indicates hyperpolarization relative to the starting membrane potential.34
Using cultured hippocampal neurons expressing Mermaid, we determined the effect of GABABR activation in the presence or absence of AP5 (Figures 2d–g). As expected, application of GABABR agonist, baclofen, significantly hyperpolarized the control dendrites as indicated by a decrease in FRET ratio (Figures 2d and e; ΔR/R=−0.09±0.03, P⩽0.02; one-way T-test). However, when NMDARs are blocked, baclofen negligibly changed the membrane potential of the primary dendrite (Figures 2d and e; ΔR/R=0.003±0.01) without altering the somatic response (Supplementary Figures S2E and F). These results suggest that inhibiting NMDAR activity abrogates GABABR-induced hyperpolarization in the dendrites.
To assess the contribution of Kir3 activity to GABABR-induced hyperpolarization during NMDAR blockade, we treated Mermaid-transfected cells with baclofen in the presence or absence of AP5. After baclofen treatment, we measured the FRET signal after blocking Kir3 channels with tertiapin Q (tert-Q, 50 nM; ΔR/R=RBACTQ−RBAC/RBAC). As expected, baclofen hyperpolarized the dendritic membrane (Figure 2f, middle panel unfilled circles; Figure 2g, white bar). Addition of tert-Q to vehicle-treated neurons increased the FRET signal, indicating a significant depolarization (Figure 2f, right panel; Figure 2g, hatched bar ΔR/R=3±1%). In contrast, AP5-treated neurons showed no observable change in FRET signal with the Kir3 blocker (Figure 2f, filled circles right panel; Figure 2g, hatched bar). These data suggest that blocking NMDARs reduces GABABR-coupled Kir3 activity.
NMDAR antagonism increases the stability of 14-3-3η, an adaptor protein that decouples GABABR and Kir3.2
Several reports link 14-3-3η to the regulation of GABABR signaling.21, 22 Thus 14-3-3η could potentially regulate GABABR response to NMDAR blockade. First, we determined whether NMDAR antagonism altered 14-3-3η expression. Western blotting analysis of hippocampal SNs indicated that NMDAR blockade (AP5 or Ro-25-6981) increased 14-3-3η expression relative to control (Figures 3a and b; AP5: 1.26±0.09, P<0.03; Ro-25-6981: 1.46±0.13, P<0.02). As seen with GABABR, social defeat significantly decreased 14-3-3η (Figure 3c; 0.68±0.04, P<0.001; relative to Control).
N-methyl-D-aspartate receptor antagonism slows both degradation and new protein synthesis of 14-3-3η. (a–c) Representative western blotting of hippocampal synaptoneurosomes show 14-3-3η protein significantly increases 45 min postinjection of (a) D-(-)-2-amino-5-phosphonopentanoic acid (AP5) or (b) Ro-25-6891 and (c) significantly decreases with social defeat. N=4 animals in panels (a and b). N=8 animals in panel (c). (d and e) AP5 decreases new translation of 14-3-3η in dendrites. N=10 neurons. Scale bar=25 μm. (f–h) AP5 slows the rate of degradation of 14-3-3η. (f) Representative images of 14-3-3η-Kaede expressing neurons before and after ultraviolet conversion (from green to red) following 60-min incubation with AP5 or vehicle (H2O, control). (g and h) Summary graphs show that AP5 significantly increases old (red) protein present in the dendrite and significantly decreases new (green) protein in distal dendrites after 1 h (see also Supplementary Figure S3). N=10 neurons. Significance assessed by Student’s T-test.
To determine whether the increase in 14-3-3η expression was due to new protein synthesis, as observed with GABABR2 (Figure 1), we used BONCAT–PLA to detect new 14-3-3η protein. Unexpectedly, NMDAR antagonism reduced the detectable levels of new, biotinylated 14-3-3η protein by ~56% (Figures 3d and e; 0–100 μm: 0.44±0.04; 100–150 μm: 0.38±0.04, P<0.005; relative to Control). These results were puzzling as the overall 14-3-3η expression levels increased with NMDAR antagonists (Figures 3a and b).
We hypothesized that NMDAR blockade increases the stability of the 14-3-3η rather than its mRNA translation. To test this prediction, we expressed the photoconvertible fluorescent protein Kaede fused to 14-3-3η in cultured hippocampal neurons. UV light converts the fluorescence Kaede from green to red irreversibly.67 After a brief UV exposure, detection of the green signal indicates a ‘new’ protein synthesis, which can be differentiated from the ‘old’ red protein. Thus Kaede can distinguish changes in protein synthesis from changes in protein stability (that is, increase in detectable green protein indicates new translation versus no change in red protein indicates protein stability).35, 36, 68, 69 Changes in ‘new’ green and ‘old’ red protein are measured by ΔF/F (ΔF/F=F0−F/F0, where F0 is the signal intensity at time point 0 after the initial photoconversion, and F is the signal intensity 60 min later). Similar to the BONCAT–PLA assay (Figures 3d and e), the increase in new green 14-3-3η was smaller in neurons where NMDARs were blocked (Figures 3f–h; green bars; 0–100 μm: Control=16±3%; AP5=10±3%; Supplementary Figure S3B). Albeit, the decrease in ‘new’ green protein was not as dramatic, as detected by BONCAT–PLA, in dendritic regions close to the cell body. However, at distances >100 μm from the soma, there was significantly less ‘new’ green protein, indicating that AP5 may significantly slow new translation of 14-3-3η in more distal dendrites (Figures 3f–h; green bars; 100–150 μm:,Control=21±4%; AP5=6±2%; P<0.003). In contrast, NMDAR blockade completely prevented the decrease in ‘old’ red protein both proximally and at distances >100 μm from the cell body, 60 min postconversion (Figures 3f–h; red bars; 0–100 μm: Control=−17±4%; AP5 4±7%; P<0.009; 100–150 μm: Control=−18±2%; AP5=3±8%; P<0.03). These results suggest that during basal NMDAR activity, 14-3-3η is rapidly synthesized and degraded. NMDAR blockade slows down both 14-3-3η synthesis and protein degradation. Thus the enhanced protein stability outweighs the reduced protein synthesis, resulting in a net increase in 14-3-3η expression levels.
14-3-3η reduces the dendritic GABABR and Kir3.2 co-localization and the surface GABABR expression in NMDAR-blocked neurons
To determine whether the increase in 14-3-3η protein with NMDAR blockade is critical to the decoupling of GABABR and Kir3 channels, we constructed a DN form of 14-3-3η (DN-14-3-3η),25 in which two arginines (R56 and R60) are mutated to alanines at the interface between the two subunits. These mutations significantly reduce dimerization of 14-3-3 proteins and hence interaction with its substrates.25 We expressed DN-14-3-3η in cultured hippocampal neurons using a rAAV. A second rAAV was used to mark the infected neurons with tdTomato red fluorescent protein. To ensure that all red neurons also expressed DN-14-3-3η, the DN-14-3-3η:tdTomato ratio was set at 4:1. We then performed the live antibody-feeding assay described above to assess the co-localization of GABABR and Kir3.2 after the addition of AP5. As previously observed (Figures 2b and c), NMDAR blockade reduced the co-localization of GABABR with Kir3.2 in neurons expressing tdTomato alone (Figures 4a and b). In DN-14-3-3η-expressing cells, AP5 did not reduce GABABR–Kir3.2 co-localization (Figures 4a and b; DN+AP5, 0.74±0.06; AP5: 1.08±0.08; both relative to Control). These data suggest that 14-3-3η may decouple GABABRs from Kir3 channels upon NMDAR antagonism.
14-3-3η is required for the reduction in γ-aminobutyric acid receptor (GABABR) co-localization with Kir3.2 and regulates surface GABABR with N-methyl-D-aspartate receptor (NMDAR) blockade. (a and b) Co-localization of GABABR with Kir3.2 is restored in NMDAR-blocked neurons expressing DN-14-3-3η. (a) Representative images of Kir3.2, surface GABABR1 and co-localized points for control, D-(-)-2-amino-5-phosphonopentanoic acid (AP5)-treated neurons expressing tdTomato and empty vector or DN-14-3-3η, scale=25 μm. (b) Summary graph of normalized Pearson’s correlation coefficient (PCC).37 N=6–8 neurons. (c and d) Blocking 14-3-3η interaction significantly enhances surface GABABR with AP5. (c) Representative neurons expressing dsRed and empty vector (pc) or DN-14-3-3η and immunostained for GABABR surface and total. (d) Summary graph of surface/total GABABR1 expression. Scale=25 μm. N=8–10 neurons. Significance (*) determined by Newman–Keuls one-way analysis of variance.
How does 14-3-3η promote the reduced GABABR–Kir3 co-localization with NMDAR blockade? 14-3-3η prevents the interaction between the C-terminal domains of GABABR1 and GABABR2,21 thereby disrupting the heterodimer.22 Thus a reduction in GABABR–Kir3 co-localization in AP5 might occur due to reduced stability of surface GABABRs present on the membrane prior to NMDAR inhibition. Because it is unknown whether heterodimerization is required for GABABR stabilization on the membrane, we next asked whether preventing 14-3-3η interaction with GABABRs affects dendritic surface expression. After co-expressing DN-14-3-3η and dsRed in hippocampal neurons, we measured surface GABABR1 in non-permeabilized neurons. As previously observed, AP5 increased surface GABABRs.19 Expression of the DN-14-3-3η enhanced the AP5-induced increase on surface GABABR1 by ~55% above AP5-treated, control neurons (Figures 4c and d; DN+AP5: 1.96±0.26; AP5: 1.40±0.16; both relative to Control; reported as the percentage of Surface/Total). These results argue that 14-3-3η reduces the membrane stability of GABABRs, which may govern the reduction in GABABR-Kir 3 channel co-localization with NMDAR blockade.
Blocking the functional interaction of 14-3-3η with GABABRs restores GABABR-mediated inhibition in neurons with reduced NMDAR activity
GABABR–Kir3 activity decreases in the presence of NMDAR antagonists (Figure 2). Moreover, 14-3-3η reduces surface GABABRs and GABABR–Kir3 co-localization (Figures 4a–d). Thus we hypothesized that blocking 14-3-3η with its DN would restore the baclofen-induced hyperpolarization during NMDAR blockade. As previously observed using the voltage-sensor Mermaid (Figure 2), AP5 significantly reduced the hyperpolarization induced by baclofen compared with vehicle-treated cells. Expression of DN-14-3-3η completely restored GABABR-mediated inhibition in AP5-treated neurons (Figures 5a and b; control: −3±0.5%; AP5: 0±0.1%; DN+AP5: −3±0.8%). These data indicate that, during NMDAR blockade, 14-3-3η is necessary to decouple GABABR from Kir3.
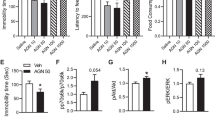
Blocking 14-3-3η function restores GABABR (γ-aminobutyric acid receptor)-mediated hyperpolarization in D-(-)-2-amino-5-phosphonopentanoic acid (AP5)-treated neurons and prevents rapid antidepressant effect of N-methyl-D-aspartate receptor antagonists in vivo. (a and b) Averaged traces of fluorescence resonance energy transfer ratio and summary graph show that the DN 14-3-3η restores baclofen-induced hyperpolarization. N=3 independent cultures. (c) Injection of DN-14-3-3η into CA1 of hippocampus prevented AP5 (2.5 mg kg−1; intraperitoneal (i.p.)) or Ro-25-6981 (Ro, 10 mg kg−1; i.p.) induced decrease in immobility in the forced swim test 45 min postinjection. (d and e) Twenty-four hours postinjection, DN14-3-3η stereotaxic injection prevented Ro-25-9681-induced increase in mobility in tail suspension test and grooming frequency in the splash test. N=3–4 animals (AP5) or N=6–8 animals (Ro-25-6891).
In vivo expression of DN-14-3-3η in hippocampal CA1 pyramidal neurons prevents the rapid antidepressant efficacy of NMDAR antagonists
We have shown in neuronal cultures that during NMDAR inhibition GABABR function shifts from opening Kir3.X channels to facilitating an increase in dendritic calcium. We have also demonstrated that the GABABR shift in function requires 14-3-3η. In vivo, we have determined that blocking GABABRs prevents the positive effects mediated by rapid antidepressants.19 Hence, we sought to examine the requirement of 14-3-3η in mediating the antidepressant effects of NMDAR antagonists. We assessed the rapid antidepressant efficacy of AP5 and Ro-25-6981 in animals expressing DN14-3-3η-Flag+tdTomato or empty vector+GFP. The expression of construct was confirmed by western blotting analysis of isolated hippocampi (Supplementary Figure S5A). Stereotaxically injected controls performed similarly to non-injected animals on the FST for saline (black), AP5 (2.5 mg kg−1; red) and Ro-25-6981 (10 mg kg−1, pink) treated mice (Figures 1a and b, Figure 5c). AP5 reduced immobility time by ~40 s at 45 min post-injection and ~45 s at 24 h postinjection. Similar to AP5, Ro-25-6981-injected mice reduced immobility by ~60 s at 45 min postinjection. However, mice expressing DN-14-3-3η (blue) were unresponsive to AP5 or Ro-25-6981 injection and had similar immobility times to saline-injected mice (Figure 5c; Control=137±4 s, DN+AP5=134±11 s; Control=136±15 s, DN+Ro-25-6981, 134±11 s; Supplementary Figure S4A).
To further validate the role of 14-3-3η in rapid antidepressant efficacy, we subjected stereotaxically injected mice to two additional behavioral readouts, the TST and the splash test, for grooming behavior. TST is commonly used to screen antidepressants,70 and grooming frequency in the splash test assesses self-care and motivational behavior.41 Consistent with the FST, Ro-25-6981 significantly increased the time spent struggling of vector-injected, control mice but not of DN-14-3-3η-Flag-injected animals (Figure 5d; Control=125±7 s, Ro-25-6981=176±10 s, DN+Ro-25-6981=118±11 s). Following the TST, animals were assessed for grooming behavior using the splash test (30–40 min after TST). We predicted that Ro-25-6981-treated, vector-injected mice will display increased rate of grooming over vehicle-treated (10% dimethyl sulfoxide), vector-injected mice. Furthermore, expression of DN-14-3-3η will prevent Ro-25-6981-induced increase in grooming. A 10% sucrose solution was squirted on the dorsal coat of the mouse, and grooming frequency was measured.42 Indeed, Ro-25-6981-treated, control mice displayed augmented grooming behavior relative to vehicle-treated, control mice (Figure 5e; Control=4±0.4, Ro-25-6981=9±1). Importantly, Ro-25-6981 in mice expressing the DN-14-3-3η did not increase the grooming frequency compared with control (Figure 5e; DN+Ro-25-6981=5±1). These data altogether suggest that 14-3-3η is necessary for the rapid antidepressant effect of NMDAR antagonists in vivo.
Discussion
Model for molecular changes mediating rapid antidepressant efficacy
We describe a mechanism that addresses how rapid antidepressants affect GABABR-mediated inhibition. We provide evidence that 14-3-3η facilitates the decoupling of GABABR from Kir3 channels. By decoupling GABABR and Kir3, GABABR function shifts from opening potassium channels to increasing resting dendritic calcium, a mechanism required to activate mTOR-dependent protein synthesis and mediate the rapid antidepressant effects of NMDAR antagonists (Supplementary Figure S5).19
Intriguingly, our results reveal an overall remodeling of dendritic proteins with rapid antidepressants. In a rat model of depression, GABABR1, GABABR2 and 14-3-3η levels are decreased. In contrast, treatments with rapid antidepressants reduce Kir3.2 levels, whereas 14-3-3η and GABABR2 concurrently increase. As R1 total protein does not initially change (Figure 1) and requires R2 to traffic to the membrane,57 the increase in R2 may be necessary for new receptors to assemble and target the membrane. An increase in new surface GABABRs that facilitate L-type Ca+2 channel activity,19 in combination with removal of GABABR-Kir 3 channels via 14-3-3η, favors the rapid shift in GABABR function to mediate calcium entry (Supplementary Figure S5). Furthermore, the drop in Kir3.2 expression in the presence of NMDAR antagonists likely ensures that GABABR activation supports an immediate increase in resting dendritic calcium, as hippocampal Kir3.2 levels do not change between socially defeated and control rats.
14-3-3η is linked to diseases whereby NMDA-dependent changes in protein expression are dysregulated, such as schizophrenia, bipolar disorder and now rodent models of depression.71, 72, 73 Our results indicate that NMDAR signaling increases the rate of 14-3-3η mRNA translation and protein degradation, arguing that precise synthesis and degradation of 14-3-3η are imperative for normal neuronal function. Consistent with this idea, a seven-base pair repeat and a single-nucleotide polymorphism within the 5′ and 3′ UTRs of 14-3-3η are associated with schizophrenia and bipolar disorder, respectively.73, 74 As most translational regulation through RNA-binding proteins and microRNAs occur at sites within the UTRs, it will be medically informative to determine whether these polymorphisms result in aberrant 14-3-3η protein levels and GABABR signaling. Interestingly, 14-3-3η is detectable in cerebrospinal fluid of patients with Alzheimer’s disease and is being considered as an early biomarker.75, 76 Our results provide a mechanistic insight into how 14-3-3η reduces inhibition that is beneficial for the treatment of depression, but if left unchecked may lead to excitotoxicity and neurodegeneration. Future work on how NMDAR signaling balances the mRNA translation and protein degradation of 14-3-3η will expand our understanding on the nature of mental health disorders.
In summary, we have identified 14-3-3η as a critical player in the removal of GABABR-Kir3 signaling that is required for the efficacy of rapid antidepressants. Notably, molecular changes induced by NMDAR antagonists in animals subjected to depression paradigms parallel those observed in naive animals.20, 77, 78, 79, 80 Consistent with these reports, we have shown that the protein levels of GABABR1, GABABR2 and 14-3-3η drop in a rat model of depression. However, a single i.p. injection of a NMDAR antagonist results in elevated expression of these proteins. NMDAR antagonists exert their effects by altering the rate of protein synthesis and degradation of GABABR2 and 14-3-3η, respectively. The exact signaling pathways that facilitate protein synthesis and degradation of these proteins are yet to be defined. Be that as it may, through careful dissection of the molecular changes that mediate the shift in GABABR signaling with rapid antidepressants, we have identified 14-3-3η as a promising target for the treatment of MDD.
Accession codes
References
Ferrari AJ, Charlson FJ, Norman RE, Patten SB, Freedman G, Murray CJ et al. Burden of depressive disorders by country, sex, age, and year: findings from the global burden of disease study 2010. PLoS Med 2013; 10: e1001547.
Kessler RC, Bromet EJ . The epidemiology of depression across cultures. Annu Rev Public Health 2013; 34: 119–138.
Levinstein MR, Samuels BA . Mechanisms underlying the antidepressant response and treatment resistance. Front Behav Neurosci 2014; 8: 208.
Duman RS, Li N, Liu RJ, Duric V, Aghajanian G . Signaling pathways underlying the rapid antidepressant actions of ketamine. Neuropharmacology 2012; 62: 35–41.
Samuels BA, Leonardo ED, Gadient R, Williams A, Zhou J, David DJ et al. Modeling treatment-resistant depression. Neuropharmacology 2011; 61: 408–413.
Pochwat B, Palucha-Poniewiera A, Szewczyk B, Pilc A, Nowak G . NMDA antagonists under investigation for the treatment of major depressive disorder. Expert Opin Invest Drugs 2014; 23: 1181–1192.
Rush AJ, Trivedi MH, Wisniewski SR, Nierenberg AA, Stewart JW, Warden D et al. Acute and longer-term outcomes in depressed outpatients requiring one or several treatment steps: a STAR*D report. Am J Psychiatry 2006; 163: 1905–1917.
Rush AJ, Trivedi MH, Wisniewski SR, Stewart JW, Nierenberg AA, Thase ME et al. Bupropion-SR, sertraline, or venlafaxine-XR after failure of SSRIs for depression. N Engl J Med 2006; 354: 1231–1242.
Holtzheimer PE, Mayberg HS . Deep brain stimulation for psychiatric disorders. Annu Rev Neurosci 2011; 34: 289–307.
Holtzheimer PE, Mayberg HS . Stuck in a rut: rethinking depression and its treatment. Trends Neurosci 2011; 34: 1–9.
Berman RM, Cappiello A, Anand A, Oren DA, Heninger GR, Charney DS et al. Antidepressant effects of ketamine in depressed patients. Biol Psychiatry 2000; 47: 351–354.
Zarate CA Jr., Singh JB, Carlson PJ, Brutsche NE, Ameli R, Luckenbaugh DA et al. A randomized trial of an N-methyl-D-aspartate antagonist in treatment-resistant major depression. Arch Gen Psychiatry 2006; 63: 856–864.
aan het Rot M, Collins KA, Murrough JW, Perez AM, Reich DL, Charney DS et al. Safety and efficacy of repeated-dose intravenous ketamine for treatment-resistant depression. Biol Psychiatry 2010; 67: 139–145.
Papp M, Moryl E . Rewarding properties of non-competitive and competitive NMDA antagonists as measured by place preference conditioning in rats. Pol J Pharmacol 1994; 46: 79–81.
Skolnick P, Popik P, Trullas R . Glutamate-based antidepressants: 20 years on. Trends Pharmacol Sci 2009; 30: 563–569.
Pozzi L, Dorocic IP, Wang X, Carlen M, Meletis K . Mice lacking NMDA receptors in parvalbumin neurons display normal depression-related behavior and response to antidepressant action of NMDAR antagonists. PLoS One 2014; 9: e83879.
Dutar P, Nicoll RA . A physiological role for GABAB receptors in the central nervous system. Nature 1988; 332: 156–158.
Cryan JF, Slattery DA . GABAB receptors and depression. Current status. Adv Pharmacol 2010; 58: 427–451.
Workman ER, Niere F, Raab-Graham KF . mTORC1-dependent protein synthesis underlying rapid antidepressant effect requires GABABR signaling. Neuropharmacology 2013; 73: 192–203.
Li N, Lee B, Liu RJ, Banasr M, Dwyer JM, Iwata M et al. mTOR-dependent synapse formation underlies the rapid antidepressant effects of NMDA antagonists. Science 2010; 329: 959–964.
Couve A, Kittler JT, Uren JM, Calver AR, Pangalos MN, Walsh FS et al. Association of GABA(B) receptors and members of the 14-3-3 family of signaling proteins. Mol Cell Neurosci 2001; 17: 317–328.
Laffray S, Bouali-Benazzouz R, Papon MA, Favereaux A, Jiang Y, Holm T et al. Impairment of GABAB receptor dimer by endogenous 14-3-3zeta in chronic pain conditions. EMBO J 2012; 31: 3239–3251.
Foote M, Zhou Y . 14-3-3 proteins in neurological disorders. Int J Biochem Mol Biol 2012; 3: 152–164.
Quinlan EM, Olstein DH, Bear MF . Bidirectional, experience-dependent regulation of N-methyl-D-aspartate receptor subunit composition in the rat visual cortex during postnatal development. Proc Natl Acad Sci USA 1999; 96: 12876–12880.
Thorson JA, Yu LW, Hsu AL, Shih NY, Graves PR, Tanner JW et al. 14-3-3 proteins are required for maintenance of Raf-1 phosphorylation and kinase activity. Mol Cell Biol 1998; 18: 5229–5238.
Shaner NC, Campbell RE, Steinbach PA, Giepmans BN, Palmer AE, Tsien RY . Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat Biotechnol 2004; 22: 1567–1572.
Varga V, Losonczy A, Zemelman BV, Borhegyi Z, Nyiri G, Domonkos A et al. Fast synaptic subcortical control of hippocampal circuits. Science 2009; 326: 449–453.
Grieger JC, Choi VW, Samulski RJ . Production and characterization of adeno-associated viral vectors. Nat Protoc 2006; 1: 1412–1428.
Ma T, Hoeffer CA, Capetillo-Zarate E, Yu F, Wong H, Lin MT et al. Dysregulation of the mTOR pathway mediates impairment of synaptic plasticity in a mouse model of Alzheimer's disease. PLoS One 2010; 5: e12845.
Hayashi T, Huganir RL . Tyrosine phosphorylation and regulation of the AMPA receptor by SRC family tyrosine kinases. J Neurosci 2004; 24: 6152–6160.
Dieterich DC, Hodas JJ, Gouzer G, Shadrin IY, Ngo JT, Triller A et al. In situ visualization and dynamics of newly synthesized proteins in rat hippocampal neurons. Nat Neurosci 2010; 13: 897–905.
Dieterich DC, Lee JJ, Link AJ, Graumann J, Tirrell DA, Schuman EM . Labeling, detection and identification of newly synthesized proteomes with bioorthogonal non-canonical amino-acid tagging. Nat Protoc 2007; 2: 532–540.
Dieterich DC, Link AJ, Graumann J, Tirrell DA, Schuman EM . Selective identification of newly synthesized proteins in mammalian cells using bioorthogonal noncanonical amino acid tagging (BONCAT). Proc Natl Acad Sci USA 2006; 103: 9482–9487.
Tsutsui H, Karasawa S, Okamura Y, Miyawaki A . Improving membrane voltage measurements using FRET with new fluorescent proteins. Nat Methods 2008; 5: 683–685.
Sosanya NM, Huang PP, Cacheaux LP, Chen CJ, Nguyen K, Perrone-Bizzozero NI et al. Degradation of high affinity HuD targets releases Kv1.1 mRNA from miR-129 repression by mTORC1. J Cell Biol 2013; 202: 53–69.
Raab-Graham KF, Haddick PC, Jan YN, Jan LY . Activity- and mTOR-dependent suppression of Kv1.1 channel mRNA translation in dendrites. Science 2006; 314: 144–148.
Bolte S, Cordelieres FP . A guided tour into subcellular colocalization analysis in light microscopy. J Microsc 2006; 224: 213–232.
Porsolt RD, Le Pichon M, Jalfre M . Depression: a new animal model sensitive to antidepressant treatments. Nature 1977; 266: 730–732.
Can A, Dao DT, Terrillion CE, Piantadosi SC, Bhat S, Gould TD . The tail suspension test. J Vis Exp 2012; 59: e3769.
Santarelli L, Saxe M, Gross C, Surget A, Battaglia F, Dulawa S et al. Requirement of hippocampal neurogenesis for the behavioral effects of antidepressants. Science 2003; 301: 805–809.
Surget A, Saxe M, Leman S, Ibarguen-Vargas Y, Chalon S, Griebel G et al. Drug-dependent requirement of hippocampal neurogenesis in a model of depression and of antidepressant reversal. Biol Psychiatry 2008; 64: 293–301.
Yalcin I, Belzung C, Surget A . Mouse strain differences in the unpredictable chronic mild stress: a four-antidepressant survey. Behav Brain Res 2008; 193: 140–143.
Nosyreva E, Szabla K, Autry AE, Ryazanov AG, Monteggia LM, Kavalali ET . Acute suppression of spontaneous neurotransmission drives synaptic potentiation. J Neurosci 2013; 33: 6990–7002.
Duman RS, Aghajanian GK . Synaptic dysfunction in depression: potential therapeutic targets. Science 2012; 338: 68–72.
Trullas R, Folio T, Young A, Miller R, Boje K, Skolnick P . 1-aminocyclopropanecarboxylates exhibit antidepressant and anxiolytic actions in animal models. Eur J Pharmacol 1991; 203: 379–385.
Boast C . Neuroprotection after brain ischemia: role of competitive NMDA antagonists. Neuol Neurobiol 1988; 46: 8.
Loscher W, Nolting B, Honack D . Evaluation of CPP, a selective NMDA antagonist, in various rodent models of epilepsy. Comparison with other NMDA antagonists, and with diazepam and phenobarbital. Eur J Pharmacol 1988; 152: 9–17.
Schoepp DD, Gamble AY, Salhoff CR, Johnson BG, Ornstein PL . Excitatory amino acid-induced convulsions in neonatal rats mediated by distinct receptor subtypes. Eur J Pharmacol 1990; 182: 421–427.
Kovacs Z, Czurko A, Kekesi KA, Juhasz G . Intracerebroventricularly administered lipopolysaccharide enhances spike-wave discharges in freely moving WAG/Rij rats. Brain Res Bull 2011; 85: 410–416.
Chapman AG, Graham JL, Patel S, Meldrum BS . Anticonvulsant activity of two orally active competitive N-methyl-D-aspartate antagonists, CGP 37849 and CGP 39551, against sound-induced seizures in DBA/2 mice and photically induced myoclonus in Papio papio. Epilepsia 1991; 32: 578–587.
Guerrini L, Molteni A, Wirth T, Kistler B, Blasi F . Glutamate-dependent activation of NF-kappaB during mouse cerebellum development. J Neurosci 1997; 17: 6057–6063.
Loeb C, Patrone A, Besio G, Balestrino M, Mainardi P . The antiepileptic effect of low-dose amino-phosphono-valeric acid (APV) is not enhanced by phosphatidylserine association. Seizure 1993; 2: 309–310.
Loeb C, Patrone A, Besio G, Balestrino M, Mainardi P . The excitatory amino acid antagonist amino-phosphono-valeric acid (APV) provides protection against penicillin-induced epileptic activity in the rat. Epilepsy Res 1990; 6: 249–251.
Leung LS, Shen B . N-methyl-D-aspartate receptor antagonists are less effective in blocking long-term potentiation at apical than basal dendrites in hippocampal CA1 of awake rats. Hippocampus 1999; 9: 617–630.
Porsolt RD . Animal model of depression. Biomedicine 1979; 30: 139–140.
Porsolt RD, Bertin A, Blavet N, Deniel M, Jalfre M . Immobility induced by forced swimming in rats: effects of agents which modify central catecholamine and serotonin activity. Eur J Pharmacol 1979; 57: 201–210.
Margeta-Mitrovic M, Jan YN, Jan LY . A trafficking checkpoint controls GABA(B) receptor heterodimerization. Neuron 2000; 27: 97–106.
Padgett CL, Slesinger PA . GABAB receptor coupling to G-proteins and ion channels. Adv Pharmacol 2010; 58: 123–147.
Chung HJ, Qian X, Ehlers M, Jan YN, Jan LY . Neuronal activity regulates phosphorylation-dependent surface delivery of G protein-activated inwardly rectifying potassium channels. Proc Natl Acad Sci USA 2009; 106: 629–634.
McClung CA, Nestler EJ . Neuroplasticity mediated by altered gene expression. Neuropsychopharmacology 2008; 33: 3–17.
Nestler EJ, Gould E, Manji H, Buncan M, Duman RS, Greshenfeld HK et al. Preclinical models: status of basic research in depression. Biol Psychiatry 2002; 52: 503–528.
Berton O, McClung CA, Dileone RJ, Krishnan V, Renthal W, Russo SJ et al. Essential role of BDNF in the mesolimbic dopamine pathway in social defeat stress. Science 2006; 311: 864–868.
Duclot F, Kabbaj M . Individual differences in novelty seeking predict subsequent vulnerability to social defeat through a differential epigenetic regulation of brain-derived neurotrophic factor expression. J Neurosci 2013; 33: 11048–11060.
Kabbaj M, Norton CS, Kollack-Walker S, Watson SJ, Robinson TE, Akil H . Social defeat alters the acquisition of cocaine self-administration in rats: role of individual differences in cocaine-taking behavior. Psychopharmacology 2001; 158: 382–387.
Costa-Nunes J, Zubareva O, Araujo-Correia M, Valenca A, Schroeter CA, Pawluski JL et al. Altered emotionality, hippocampus-dependent performance and expression of NMDA receptor subunit mRNAs in chronically stressed mice. Stress 2014; 17: 108–116.
Perron A, Mutoh H, Akemann W, Gautam SG, Dimitrov D, Iwamoto Y et al. Second and third generation voltage-sensitive fluorescent proteins for monitoring membrane potential. Front Mol Neurosci 2009; 2: 5.
Mizuno H, Mal TK, Tong KI, Ando R, Furuta T, Ikura M et al. Photo-induced peptide cleavage in the green-to-red conversion of a fluorescent protein. Mol Cell 2003; 12: 1051–1058.
Banerjee S, Neveu P, Kosik KS . A coordinated local translational control point at the synapse involving relief from silencing and MOV10 degradation. Neuron 2009; 64: 871–884.
Wang DO, Kim SM, Zhao Y, Hwang H, Miura SK, Sossin WS et al. Synapse- and stimulus-specific local translation during long-term neuronal plasticity. Science 2009; 324: 1536–1540.
Steru L, Chermat R, Thierry B, Simon P . The tail suspension test: a new method for screening antidepressants in mice. Psychopharmacology 1985; 85: 367–370.
Duan S, Gao R, Xing Q, Du J, Liu Z, Chen Q et al. A family-based association study of schizophrenia with polymorphisms at three candidate genes. Neurosci Lett 2005; 379: 32–36.
Grover D, Verma R, Goes FS, Mahon PL, Gershon ES, McMahon FJ et al. Family-based association of YWHAH in psychotic bipolar disorder. Am J Med Genet B Neuropsychiatr Genet 2009; 150B: 977–983.
Toyooka K, Muratake T, Tanaka T, Igarashi S, Watanabe H, Takeuchi H et al. 14-3-3 protein eta chain gene (YWHAH) polymorphism and its genetic association with schizophrenia. Am J Med Genet 1999; 88: 164–167.
Wong AH, Macciardi F, Klempan T, Kawczynski W, Barr CL, Lakatoo S et al. Identification of candidate genes for psychosis in rat models, and possible association between schizophrenia and the 14-3-3eta gene. Mol Psychiatry 2003; 8: 156–166.
Wiltfang J, Otto M, Baxter HC, Bodemer M, Steinacker P, Bahn E et al. Isoform pattern of 14-3-3 proteins in the cerebrospinal fluid of patients with Creutzfeldt-Jakob disease. J Neurochem 1999; 73: 2485–2490.
Li XH, Noguchi A, Nishida T, Takahashi H, Zheng Y, Yang XH et al. Cytoplasmic expression of p33ING1b is correlated with tumorigenesis and progression of head and neck squamous cell carcinoma. Histol Histopathol 2011; 26: 597–607.
Papp M, Moryl E . Antidepressant activity of non-competitive and competitive NMDA receptor antagonists in a chronic mild stress model of depression. Eur J Pharmacol 1994; 263: 1–7.
Li N, Liu RJ, Dwyer JM, Banasr M, Lee B, Son H et al. Glutamate N-methyl-D-aspartate receptor antagonists rapidly reverse behavioral and synaptic deficits caused by chronic stress exposure. Biol Psychiatry 2011; 69: 754–761.
Autry AE, Adachi M, Nosyreva E, Na ES, Los MF, Cheng PF et al. NMDA receptor blockade at rest triggers rapid behavioural antidepressant responses. Nature 2011; 475: 91–95.
Donahue RJ, Muschamp JW, Russo SJ, Nestler EJ, Carlezon WA Jr . Effects of striatal δFosB overexpression and ketamine on social defeat stress-induced anhedonia in mice. Biol Psychiatry 2014; 76: 550–558.
Acknowledgements
This work was supported by the NSF Grants IOS-1026527 and IOS-1355158, Department of Defense USAMRMC Award W81XWH-14-10061 and a University of Texas Research Grant to KFR-G. FN is supported by NSF PRFB 1306528. We thank Dr Atsushi Miyawaki and Dr Hidekazu Tsutsui for the Kaede and Mermaid cDNA, Dr Hitoshi Morikawa and Claire Stelly for helping with socially defeated rat experiments, Dr Michael Drew and Kylie Huckleberry for their expert advice and assistance with the experimental design for antidepressant efficacy, Dr Daniel Johnston and Dr Michael Drew for their critical reading of this manuscript and Dr Lily and Dr Yuh Nung Jan for their support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
PCGH is employed by Genentech/Roche and may own shares in the company. This does not alter the authors’ adherence to all the journal’s policies on sharing data and materials. The remaining authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Workman, E., Haddick, P., Bush, K. et al. Rapid antidepressants stimulate the decoupling of GABAB receptors from GIRK/Kir3 channels through increased protein stability of 14-3-3η. Mol Psychiatry 20, 298–310 (2015). https://doi.org/10.1038/mp.2014.165
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2014.165