Abstract
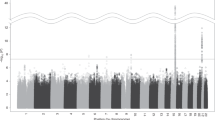
Smoking is a major risk factor for several somatic diseases and is also emerging as a causal factor for neuropsychiatric disorders. Genome-wide association (GWA) and candidate gene studies for smoking behavior and nicotine dependence (ND) have disclosed too few predisposing variants to account for the high estimated heritability. Previous large-scale GWA studies have had very limited phenotypic definitions of relevance to smoking-related behavior, which has likely impeded the discovery of genetic effects. We performed GWA analyses on 1114 adult twins ascertained for ever smoking from the population-based Finnish Twin Cohort study. The availability of 17 smoking-related phenotypes allowed us to comprehensively portray the dimensions of smoking behavior, clustered into the domains of smoking initiation, amount smoked and ND. Our results highlight a locus on 16p12.3, with several single-nucleotide polymorphisms (SNPs) in the vicinity of CLEC19A showing association (P<1 × 10−6) with smoking quantity. Interestingly, CLEC19A is located close to a previously reported attention-deficit hyperactivity disorder (ADHD) linkage locus and an evident link between ADHD and smoking has been established. Intriguing preliminary association (P<1 × 10−5) was detected between DSM-IV (Diagnostic and Statistical Manual of Mental Disorders, 4th edition) ND diagnosis and several SNPs in ERBB4, coding for a Neuregulin receptor, on 2q33. The association between ERBB4 and DSM-IV ND diagnosis was replicated in an independent Australian sample. Recently, a significant increase in ErbB4 and Neuregulin 3 (Nrg3) expression was revealed following chronic nicotine exposure and withdrawal in mice and an association between NRG3 SNPs and smoking cessation success was detected in a clinical trial. ERBB4 has previously been associated with schizophrenia; further, it is located within an established schizophrenia linkage locus and within a linkage locus for a smoker phenotype identified in this sample. In conclusion, we disclose novel tentative evidence for the involvement of ERBB4 in ND, suggesting the involvement of the Neuregulin/ErbB signalling pathway in addictions and providing a plausible link between the high co-morbidity of schizophrenia and ND.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Centers for Disease Control (CDC). Smoking-attributable mortality, years of potential life lost, and productivity losses—United States, 2000–2004. MMWR Morb Mortal Wkly Rep 2008; 57: 1226–1228.
Steuber TL, Danner F . Adolescent smoking and depression: which comes first? Addict Behav 2006; 31: 133–136.
Munafò MR, Hitsman B, Rende R, Metcalfe C, Niaura R . Effects of progression to cigarette smoking on depressed mood in adolescents: evidence from the National Longitudinal Study of Adolescent Health. Addiction 2008; 103: 162–171.
Boden JM, Fergusson DM, Horwood LJ . Cigarette smoking and depression: tests of causal linkage using a longitudinal birth cohort. Br J Psychiatry 2010; 196: 440–446.
Munafò MR, Araya R . Cigarette smoking and depression: a question of causation. Br J Psychiatry 2010; 196: 425–426.
Benowitz NL . Nicotine addiction. N Engl J Med 2010; 362: 2295–2303.
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders: DSM-IV 4th edn. American Psychiatric Association: Washington DC, USA, 1994.
Heatherton TF, Kozlowski LT, Frecker RC, Fagerström KO . The Fagerström Test for Nicotine Dependence: a revision of the Fagerström Tolerance Questionnaire. Br J Addict 1991; 86: 1119–1127.
Rose JE, Broms U, Korhonen T, Dick DM, Kaprio J . Genetics of smoking behavior. In: Kim YK (ed). Handbook of Behavior Genetics. Springer: New York, NY, USA pp 411–432 2009.
Hung RJ, McKay JD, Gaborieau V, Boffetta P, Hashibe M, Zaridze D et al. A susceptibility locus for lung cancer maps to nicotinic acetylcholine receptor subunit genes on 15q25. Nature 2008; 452: 633–637.
Amos CI, Wu X, Broderick P, Gorlov IP, Gu J, Eisen T et al. Genome-wide association scan of tag SNPs identifies a susceptibility locus for lung cancer at 15q25.1. Nat Genet 2008; 40: 616–622.
Thorgeirsson TE, Geller F, Sulem P, Rafnar T, Wiste A, Magnusson KP et al. A variant associated with nicotine dependence, lung cancer and peripheral arterial disease. Nature 2008; 452: 638–642.
Keskitalo K, Broms U, Heliövaara M, Ripatti S, Surakka I, Perola M et al. Association of serum cotinine level with a cluster of three nicotinic acetylcholine receptor genes (CHRNA3/CHRNA5/CHRNB4) on chromosome 15. Hum Mol Genet 2009; 18: 4007–4012.
Munafò MR, Timofeeva MN, Morris RW, Prieto-Merino D, Sattar N, Brennan P et al. Association between genetic variants on chromosome 15q25 locus and objective measures of tobacco exposure. J Natl Cancer Inst 2012; 104: 740–748.
Shiffman S, Waters A, Hickcox M . The Nicotine Dependence Syndrome Scale: a multidimensional measure of nicotine dependence. Nicotine Tob Res 2004; 6: 327–348.
Broms U, Wedenoja J, Largeau MR, Korhonen T, Pitkäniemi J, Keskitalo-Vuokko K et al. Analysis of detailed phenotype profiles reveals CHRNA5-CHRNA3-CHRNB4 gene cluster association with several nicotine dependence traits. Nicotine Tob Res 2012; 14: 720–733.
Uhl GR, Liu QR, Drgon T, Johnson C, Walther D, Rose JE . Molecular genetics of nicotine dependence and abstinence: whole genome association using 520,000 SNPs. BMC Genet 2007; 8: 10.
Bierut LJ, Madden PA, Breslau N, Johnson EO, Hatsukami D, Pomerleau OF et al. Novel genes identified in a high-density genome wide association study for nicotine dependence. Hum Mol Genet 2007; 16: 24–35.
Liu JZ, Tozzi F, Waterworth DM, Pillai SG, Muglia P, Middleton L et al. Meta-analysis and imputation refines the association of 15q25 with smoking quantity. Nat Genet 2010; 42: 436–440.
Tobacco and Genetics Consortium. Genome-wide meta-analyses identify multiple loci associated with smoking behavior. Nat Genet 2010; 42: 441–447.
Thorgeirsson TE, Gudbjartsson DF, Surakka I, Vink JM, Amin N, Geller F et al. Sequence variants at CHRNB3-CHRNA6 and CYP2A6 affect smoking behavior. Nat Genet 2010; 42: 448–453.
Vink JM, Smit AB, de Geus EJ, Sullivan P, Willemsen G, Hottenga JJ et al. Genome-wide association study of smoking initiation and current smoking. Am J Hum Genet 2009; 84: 367–379.
Liu YZ, Pei YF, Guo YF, Wang L, Liu XG, Yan H et al. Genome-wide association analyses suggested a novel mechanism for smoking behavior regulated by IL15. Mol Psychiatry 2009; 14: 668–680.
Lind PA, Macgregor S, Vink JM, Pergadia ML, Hansell NK, de Moor MH et al. A genomewide association study of nicotine and alcohol dependence in Australian and Dutch populations. Twin Res Hum Genet 2010; 13: 10–29.
Yoon D, Kim YJ, Cui WY, Van der Vaart A, Cho YS, Lee JY et al. Large-scale genome-wide association study of Asian population reveals genetic factors in FRMD4A and other loci influencing smoking initiation and nicotine dependence. Hum Genet 2011; 131: 1009–1021.
Drgon T, Montoya I, Johnson C, Liu QR, Walther D, Hamer D et al. Genome-wide association for nicotine dependence and smoking cessation success in NIH research volunteers. Mol Med 2009; 15: 21–27.
Caporaso N, Gu F, Chatterjee N, Sheng-Chih J, Yu K, Yeager M et al. Genome-wide and candidate gene association study of cigarette smoking behaviors. PLoS ONE 2009; 4: e4653.
Siedlinski M, Cho MH, Bakke P, Gulsvik A, Lomas DA, Anderson W et al. Genome-wide association study of smoking behaviors in patients with COPD. Thorax 2011; 66: 894–902.
Saccone NL, Culverhouse RC, Schwantes-An TH, Cannon DS, Chen X, Cichon S et al. Multiple independent loci at chromosome 15q25.1 affect smoking quantity: a meta-analysis and comparison with lung cancer and COPD. PLoS Genet 2010; 6: e1001053.
Broms U, Madden PA, Heath AC, Pergadia ML, Shiffman S, Kaprio J . The Nicotine Dependence Syndrome Scale in Finnish smokers. Drug Alcohol Depend 2007; 89: 42–51.
Loukola A, Broms U, Maunu H, Widén E, Heikkilä K, Siivola M et al. Linkage of nicotine dependence and smoking behavior on 10q, 7q and 11p in twins with homogeneous genetic background. Pharmacogenomics J 2008; 8: 209–219.
Saccone SF, Pergadia ML, Loukola A, Broms U, Montgomery GW, Wang JC et al. Genetic linkage to chromosome 22q12 for a heavy-smoking quantitative trait in two independent samples. Am J Hum Genet 2007; 80: 856–866.
Hartz SM, Short SE, Saccone NL, Culverhouse R, Chen L, Schwantes-An TH et al. Increased genetic vulnerability to smoking at CHRNA5 in early-onset smokers. Arch Gen Psychiatry 2012; 69: 854–860.
Knaapila A, Silventoinen K, Broms U, Rose RJ, Perola M, Kaprio J et al. Food neophobia in young adults: genetic architecture and relation to personality, pleasantness and use frequency of foods, and body mass index—a twin study. Behav Genet 2011; 41: 512–521.
Heath AC, Whitfield JB, Martin NG, Pergadia ML, Goate AM, Lind PA et al. A quantitative-trait genome-wide association study of alcoholism risk in the community: findings and implications. Biol Psychiatry 2011; 70: 513–518.
Bucholz KK, Cadoret R, Cloninger CR, Dinwiddie SH, Hesselbrock VM, Nurnberger JI Jr et al. A new, semi-structured psychiatric interview for use in genetic linkage studies: a report on the reliability of the SSAGA. J Stud Alcohol 1994; 55: 149–158.
Cottler LB, Robins LN, Grant BF, Blaine J, Towle LH, Wittchen HU et al. The CIDI-core substance abuse and dependence questions: cross-cultural and nosological issues. The WHO/ADAMHA Field Trial. Br J Psychiatry 1991; 159: 653–658.
Nyholt DR . A simple correction for multiple testing for single-nucleotide polymorphisms in linkage disequilibrium with each other. Am J Hum Genet 2004; 74: 765–769.
Raine A, Benishay D . The SPQ-B: a brief screening instrument for schizotypal personality disorder. J Pers Disord 1995; 9: 346–355.
Raine A, Reynolds C, Lencz T, Scerbo A, Triphon N, Kim D . Cognitive-perceptual interpersonal, and disorganized features of schizotypal personality. Schizophr Bull 1994; 20: 191–201.
Howie BN, Donnelly P, Marchini J . A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet 2009; 5: e1000529.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007; 81: 559–575.
Barrett JC, Fry B, Maller J, Daly MJ . Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 2005; 21: 263–265.
Liu JZ, McRae AF, Nyholt DR, Medland SE, Wray NR, Brown KM et al. A versatile gene-based test for genome-wide association studies. Am J Hum Genet 2010; 87: 139–145.
Bierut LJ, Stitzel JA, Wang JC, Hinrichs AL, Grucza RA, Xuei X et al. Variants in nicotinic receptors and risk for nicotine dependence. Am J Psychiatry 2008; 165: 1163–1171.
Fowler CD, Lu Q, Johnson PM, Marks MJ, Kenny PJ . Habenular α5 nicotinic receptor subunit signalling controls nicotine intake. Nature 2011; 471: 597–601.
Dani JA, Harris RA . Nicotine addiction and comorbidity with alcohol abuse and mental illness. Nat Neurosci 2005; 8: 1465–1470.
Dudbridge F, Gusnanto A . Estimation of significance thresholds for genomewide association scans. Genet Epidemiol 2008; 32: 227–234.
Pe’er I, Yelensky R, Altshuler D, Daly MJ . Estimation of the multiple testing burden for genomewide association studies of nearly all common variants. Genet Epidemiol 2008; 32: 381–385.
Sullivan P on behalf of 96 psychiatric genetics investigators. Don’t give up on GWAS. Mol Psychiatry 2012; 17: 2–3.
Visscher PM, Brown MA, McCarthy MI, Yang J . Five years of GWAS discovery. Am J Hum Genet 2012; 90: 7–24.
Browning SR, Browning BL . Haplotype phasing: existing methods and new developments. Nat Rev Genet 2011; 12: 703–714.
Moss HB, Chen CM, Yi HY . Measures of substance consumption among substance users, DSM-IV abusers, and those with DSM-IV dependence disorders in a nationally representative sample. J Stud Alcohol Drugs 2012; 73: 820–828.
Kaprio J, Koskenvuo M . Genetic and environmental factors in complex diseases: the older Finnish Twin Cohort. Twin Res 2002; 5: 358–365.
Han S, Gelernter J, Luo X, Yang BZ . Meta-analysis of 15 genome-wide linkage scans of smoking behavior. Biol Psychiatry 2010; 67: 12–19.
Romanos M, Freitag C, Jacob C, Craig DW, Dempfle A, Nguyen TT et al. Genome-wide linkage analysis of ADHD using high-density SNP arrays: novel loci at 5q13.1 and 14q12. Mol Psychiatry 2008; 13: 522–530.
Ogdie MN, Macphie IL, Minassian SL, Yang M, Fisher SE, Francks C et al. A genomewide scan for attention-deficit/hyperactivity disorder in an extended sample: suggestive linkage on 17p11. Am J Hum Genet 2003; 72: 1268–1279.
Fisher SE, Francks C, McCracken JT, McGough JJ, Marlow AJ, MacPhie IL et al. A genomewide scan for loci involved in attention-deficit/hyperactivity disorder. Am J Hum Genet 2002; 70: 1183–1196.
Lerman C, Audrain J, Tercyak K, Hawk LW Jr, Bush A, Crystal-Mansour S et al. Attention-deficit hyperactivity disorder (ADHD) symptoms and smoking patterns among participants in a smoking-cessation program. Nicotine Tob Res 2001; 3: 353–359.
Pomerleau CS, Downey KK, Snedecor SM, Mehringer AM, Marks JL, Pomerleau OF . Smoking patterns and abstinence effects in smokers with no ADHD, childhood ADHD, and adult ADHD symptomatology. Addict Behav 2003; 28: 1149–1157.
Sihvola E, Rose RJ, Dick DM, Korhonen T, Pulkkinen L, Raevuori A et al. Prospective relationships of ADHD symptoms with developing substance use in a population-derived sample. Psychol Med 2011; 20: 1–9.
Shi SH, Jan LY, Jan YN . Hippocampal neuronal polarity specified by spatially localized mPar3/mPar6 and PI 3-kinase activity. Cell 2003; 112: 63–75.
Uhl GR, Liu QR, Drgon T, Johnson C, Walther D, Rose JE et al. Molecular genetics of successful smoking cessation: convergent genome-wide association study results. Arch Gen Psychiatry 2008; 65: 683–693.
Burden S, Yarden Y . Neuregulins and their receptors: a versatile signaling module in organogenesis and oncogenesis. Neuron 1997; 18: 847–855.
Moolchan ET, Radzius A, Epstein DH, Uhl G, Gorelick DA, Cadet JL et al. The Fagerstrom Test for Nicotine Dependence and the Diagnostic Interview Schedule: do they diagnose the same smokers? Addict Behav 2002; 27: 101–113.
Piper ME, McCarthy DE, Baker TB . Assessing tobacco dependence: a guide to measure evaluation and selection. Nicotine Tob Res 2006; 8: 339–351.
Bevilacqua L, Doly S, Kaprio J, Yuan Q, Tikkanen R, Paunio T et al. A population-specific HTR2B stop codon predisposes to severe impulsivity. Nature 2010; 468: 1061–1066.
Norton N, Moskvina V, Morris DW, Bray NJ, Zammit S, Williams NM et al. Evidence that interaction between neuregulin 1 and its receptor erbB4 increases susceptibility to schizophrenia. Am J Med Genet B Neuropsychiatr Genet 2006; 141B: 96–101.
Silberberg G, Darvasi A, Pinkas-Kramarski R, Navon R . The involvement of ErbB4 with schizophrenia: association and expression studies. Am J Med Genet B Neuropsychiatr Genet 2006; 141B: 142–148.
Nicodemus KK, Luna A, Vakkalanka R, Goldberg T, Egan M, Straub RE et al. Further evidence for association between ErbB4 and schizophrenia and influence on cognitive intermediate phenotypes in healthy controls. Mol Psychiatry 2006; 11: 1062–1065.
Law AJ, Kleinman JE, Weinberger DR, Weickert CS . Disease-associated intronic variants in the ErbB4 gene are related to altered ErbB4 splice-variant expression in the brain in schizophrenia. Hum Mol Genet 2007; 16: 129–141.
Ng MY, Levinson DF, Faraone SV, Suarez BK, DeLisi LE, Arinami T et al. Meta-analysis of 32 genome-wide linkage studies of schizophrenia. Mol Psychiatry 2009; 14: 774–785.
Golub MS, Germann SL, Lloyd KC . Behavioral characteristics of a nervous system-specific erbB4 knock-out mouse. Behav Brain Res 2004; 153: 159–170.
Hahn CG, Wang HY, Cho DS, Talbot K, Gur RE, Berrettini WH et al. Altered neuregulin 1-erbB4 signaling contributes to NMDA receptor hypofunction in schizophrenia. Nat Med 2006; 12: 824–828.
Zuliani R, Moorhead TW, Bastin ME, Johnstone EC, Lawrie SM, Brambilla P et al. Genetic variants in the ErbB4 gene are associated with white matter integrity. Psychiatry Res 2011; 191: 133–137.
Paunio T, Ekelund J, Varilo T, Parker A, Hovatta I, Turunen JA et al. Genome-wide scan in a nationwide study sample of schizophrenia families in Finland reveals susceptibility loci on chromosomes 2q and 5q. Hum Mol Genet 2001; 10: 3037–3048.
Paunio T, Tuulio-Henriksson A, Hiekkalinna T, Perola M, Varilo T, Partonen T et al. Search for cognitive trait components of schizophrenia reveals a locus for verbal learning and memory on 4q and for visual working memory on 2q. Hum Mol Genet 2004; 13: 1693–1702.
Service S, DeYoung J, Karayiorgou M, Roos JL, Pretorious H, Bedoya G et al. Magnitude and distribution of linkage disequilibrium in population isolates and implications for genome-wide association studies. Nat Genet 2006; 38: 556–560.
Peltonen L, Palotie A, Lange K . Use of population isolates for mapping complex traits. Nat Rev Genet 2000; 1: 182–190.
Surakka I, Kristiansson K, Anttila V, Inouye M, Barnes C, Moutsianas L et al. Founder population-specific HapMap panel increases power in GWA studies through improved imputation accuracy and CNV tagging. Genome Res 2010; 20: 1344–1351.
Johnson C, Drgon T, Liu QR, Zhang PW, Walther D, Li CY et al. Genome wide association for substance dependence: convergent results from epidemiologic and research volunteer samples. BMC Med Genet 2008; 9: 113–122.
Drgon T, Johnson CA, Nino M, Drgonova J, Walther DM, Uhl GR . “Replicated” genome wide association for dependence on illegal substances: genomic regions identified by overlapping clusters of nominally positive SNPs. Am J Med Genet B Neuropsychiatr Genet 2011; 156: 125–138.
Uhl GR, Drgon T, Liu QR, Johnson C, Walther D, Komiyama T et al. Genome-wide association for methamphetamine dependence: convergent results from 2 samples. Arch Gen Psychiatry 2008; 65: 345–355.
Uhl GR, Drgon T, Johnson C, Walther D, David SP, Aveyard P et al. Genome-wide association for smoking cessation success: participants in the Patch in Practice trial of nicotine replacement. Pharmacogenomics 2010; 11: 357–367.
Wang KS, Liu X, Zhang Q, Pan Y, Aragam N, Zeng M . A meta-analysis of two genome-wide association studies identifies 3 new loci for alcohol dependence. J Psychiatr Res 2011; 45: 1419–1425.
Wang KS, Liu X, Zhang Q, Wu LY, Zeng M . Genome-wide association study identifies 5q21 and 9p24.1 (KDM4C) loci associated with alcohol withdrawal symptoms. J Neural Transm 2012; 199: 425–433.
Kendler KS, Kalsi G, Holmans PA, Sanders AR, Aggen SH, Dick DM et al. Genomewide association analysis of symptoms of alcohol dependence in the molecular genetics of schizophrenia (MGS2) control sample. Alcohol Clin Exp Res 2011; 35: 963–975.
Uhl GR, Drgon T, Johnson C, Ramoni MF, Behm FM, Rose JE . Genome-wide association for smoking cessation success in a trial of precessation nicotine replacement. Mol Med 2010; 16: 513–526.
Zuo L, Gelernter J, Zhang CK, Zhao H, Lu L, Kranzler HR et al. Genome-wide association study of alcohol dependence implicates KIAA0040 on chromosome 1q. Neuropsychopharmacology 2012; 37: 557–566.
Drgon T, Zhang PW, Johnson C, Walther D, Hess J, Nino M et al. Genome wide association for addiction: replicated results and comparisons of two analytic approaches. PLoS One 2010; 5: e8832.
Young P, Nie J, Wang X, McGlade CJ, Rich MM, Feng G . LNX1 is a perisynaptic Schwann cell specific E3 ubiquitin ligase that interacts with ErbB2. Mol Cell Neurosci 2005; 30: 238–248.
Acknowledgements
We warmly thank the participating twin pairs and their family members for their contribution. We would like to express our appreciation to the skilled study interviewers A-M Iivonen, K Karhu, H-M Kuha, U Kulmala-Gråhn, M Mantere, K Saanakorpi, M Saarinen, R Sipilä, L Viljanen and E Voipio. E Hämäläinen and M Sauramo are acknowledged for their skilful technical assistance. Dr E Vuoksimaa and Dr A Latvala are thanked for collaboration in FT12 traits related to cognitive functions and schizotypy. Professor A Palotie is acknowledged for his advice and expertise in whole-genome genotyping. We are ever grateful to the late Academician Leena Peltonen-Palotie for her indispensable contribution throughout the years of the study. This work was supported for data collection by Academy of Finland grants (JK) and a NIH Grant DA12854 (PAFM). Genome-wide genotyping in the Finnish sample was funded by Global Research Award for Nicotine Dependence/Pfizer Inc. (JK), and Wellcome Trust Sanger Institute, UK. Genome-wide genotyping in the Australian sample was funded by NIH Grants AA013320, AA013321, AA013326, AA011998 and AA017688. This work was further supported by the Sigrid Juselius Foundation (JK), Doctoral Programs of Public Health (UB), the Yrjö Jahnsson Foundation (UB), the Jenny and Antti Wihuri Foundation (JK), the Juho Vainio Foundation for Post-Doctoral research (UB), Finnish Cultural Foundation (TK), a NIH Grant DA019951 (MLP) and by the Academy of Finland Center of Excellence in Complex Disease Genetics (Grant numbers: 213506, 129680 to JK).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
JK has served as a consultant to Pfizer in 2008, 2011 and 2012. UB has served as a consultant to Pfizer in 2008. TK has served as a consultant to Pfizer in 2011 and 2012. The other authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
Loukola, A., Wedenoja, J., Keskitalo-Vuokko, K. et al. Genome-wide association study on detailed profiles of smoking behavior and nicotine dependence in a twin sample. Mol Psychiatry 19, 615–624 (2014). https://doi.org/10.1038/mp.2013.72
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2013.72
Keywords
This article is cited by
-
Multivariate genome-wide association study of depression, cognition, and memory phenotypes and validation analysis identify 12 cross-ethnic variants
Translational Psychiatry (2022)
-
Association of NRG3 and ERBB4 gene polymorphism with nicotine dependence in Turkish population
Molecular Biology Reports (2021)
-
Schizophrenia and pregnancy: a national register-based follow-up study among Finnish women born between 1965 and 1980
Archives of Women's Mental Health (2020)
-
Neuregulin 3 Signaling Mediates Nicotine-Dependent Synaptic Plasticity in the Orbitofrontal Cortex and Cognition
Neuropsychopharmacology (2018)
-
Human Genetics of Addiction: New Insights and Future Directions
Current Psychiatry Reports (2018)