Abstract
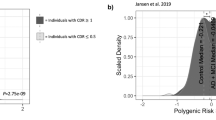
Scientists often interpret P-values as measures of the relative strength of statistical findings. This is common practice in large-scale genomic studies where P-values are used to choose which of numerous hypothesis test results should be pursued in subsequent research. In this study, we examine P-value variability to assess the degree of certainty P-values provide. We develop prediction intervals for the P-value in a replication study given the P-value observed in an initial study. The intervals depend on the initial value of P and the ratio of sample sizes between the initial and replication studies, but not on the underlying effect size or initial sample size. The intervals are valid for most large-sample statistical tests in any context, and can be used in the presence of single or multiple tests. While P-values are highly variable, future P-value variability can be explicitly predicted based on a P-value from an initial study. The relative size of the replication and initial study is an important predictor of the P-value in a subsequent replication study. We provide a handy calculator implementing these results and apply them to a study of Alzheimer’s disease and recent findings of the Cross-Disorder Group of the Psychiatric Genomics Consortium. This study suggests that overinterpretation of very significant, but highly variable, P-values is an important factor contributing to the unexpectedly high incidence of non-replication. Formal prediction intervals can also provide realistic interpretations and comparisons of P-values associated with different estimated effect sizes and sample sizes.
Similar content being viewed by others
Introduction
Genome-wide association studies (GWAS) and other studies that use modern high-throughput technologies, include very large numbers of hypothesis tests. Current genotyping arrays test up to 2.5M different single nucleotide polymorphisms (SNPs) on a single chip. While few, or sometimes no, SNPs may reach strict genome-wide statistical significance (e.g. P < 10−8), literally thousands of SNPs may appear worthy of further investigation. In addition to using P-values to make hypothesis test decisions, investigators often rely on P-value size to decide which findings to pursue in further research. Biological experiments, replication studies and custom designed arrays can usually target only the most promising SNPs from an initial study.1 Policies requiring the reporting of P-values for all SNPs investigated in a given study,2 are further evidence that P-values are seen as an important metric.
By definition, a P-value is the probability of seeing results as supportive of a genetic or other association as the observed results if, in fact, no such association exists. However, P-values are also data-dependent statistics that vary from sample to sample, even when the underlying effects, population and sampling strategy are the same.
A number of authors have shown that the sampling variability of P-values is high and that they have little predictive value for subsequent replication studies.3, 4, 5, 6 Specifically, Goodman3 reported numerical examples based on a Bayesian posterior distribution to show that power based on P-values or ‘replication probabilities … are much lower than what most people would expect’. Senn4 extended these calculations for ‘repetition probabilities’ in unequal-sized samples, again focusing on power. Cumming5 addressed the problem from a frequentist perspective, for a same-sized replication study, concluding that ‘P-values predict the future only vaguely’. However, Goodman, Senn and Cumming all specifically limited their results to a single statistical setting: a two-group comparison of Normal population data. In contrast, Boos and Stefanski6 took an operational approach to predicting P-values, applying a bootstrap prediction interval procedure developed by Mojirsheibani and Tibshirani.7 The flexibility of the bootstrap allows predictions in a variety of settings beyond the simple two-group comparisons described by Goodman, Senn and Cumming. As in the current project, application of bootstrap intervals implicitly assumes that the P-values are derived from test statistics with underlying large-sample Normal distributions. However, the bootstrap approach, which relies on a data-dependent operation, additionally requires access to the complete data used in the original analysis for implementation. Thus, bootstrap prediction intervals do not generalize to other data sets or settings and this operational strategy does not reveal general patterns.
In this paper, we show that P-values are not only highly variable, but that the degree of variability is predictable and numerically consistent across most types of statistical studies so that identical prediction intervals apply without extensive computation or access to raw data. These intervals can be computed using only the P-value and the relative sample size of the replication study to the sample size used to generate the original finding. Relative sample size, independent of absolute sample size, turns out to be an important predictor of P-value variability when retesting the same hypothesis. Our results reveal general patterns and can be used to provide explicit interpretations and comparisons of P-values in the literature without specific information about study designs or statistical analyses. We illustrate the prediction intervals using results from a recent genome-wide association study of magnetic resonance imaging (MRI) atrophy measures in Alzheimer’s disease8 and findings of the Cross-Disorder Group of the Psychiatric Genomics Consortium (PGC).9 We conclude that P-value variation may explain many failures to replicate apparently strong initial findings of genetic association.
Materials and methods
We obtained prediction intervals for p2, the P-value in a replication study. The prediction interval coverage rate, C%, is the percentage of replication P-values covered by the interval in repeated sampling, when both initial and replication studies are repeated many times.
We first considered prediction intervals for P-values obtained from one-sided tests. Consider an initial study 1 of size n1 and an independent replication study 2 of size n2, each testing the population parameter or effect θ against the null hypothesis θ = 0. For example, θ might be the log odds ratio (OR) in logistic regression. We adopt a frequentist framework and assume that large-sample Normal theory applies as it does for most statistical tests. That is, θ is fixed and unknown. Furthermore, the test statistic can be written as a Z-statistic,  , in large samples, where ni is the sample size,
, in large samples, where ni is the sample size,  is an estimator of θ and σ is a constant.
is an estimator of θ and σ is a constant.
Under standard large-sample theory, Z1 from the original study and Z2 from the replication study are independent Normal random variables with variance 1 and means  . Let Z(p) be the inverse cumulative distribution function (cdf) of the standard Normal distribution. Given an observed initial P-value, p1, we compute the initial Z-statistic as Z1 = Z(p1). (Without loss of generality, we use Z < 0.) To obtain a prediction interval for p2, we first construct a prediction interval for the difference
. Let Z(p) be the inverse cumulative distribution function (cdf) of the standard Normal distribution. Given an observed initial P-value, p1, we compute the initial Z-statistic as Z1 = Z(p1). (Without loss of generality, we use Z < 0.) To obtain a prediction interval for p2, we first construct a prediction interval for the difference  . In large samples, R is Normal with expected value
. In large samples, R is Normal with expected value  and variance 1+n2/n1. An exact C% prediction interval for R is given by
and variance 1+n2/n1. An exact C% prediction interval for R is given by

where β = 1−C/100 is the non-coverage probability. (For a 95% prediction interval, β = 0.05 and Z(0.975) = 1.96.) We rearrange the interval for R to obtain an equivalent prediction interval for Z2,

The former interval covers R if and only if the latter interval covers Z2 and their coverage rates must be the same. The C% prediction interval for p2 is calculated by applying the standard Normal cdf to the endpoints of the interval for Z2. The strategy used here to derive the prediction intervals is the same strategy used to predict new observations under an estimated linear regression model.10,11 An important consequence of this approach is that the resulting intervals do not depend on the unknown effect size or the absolute sample sizes, providing a general formula requiring only the value of p1 observed in the initial study and the ratio of sample sizes between the two studies.
See Supplementary Methods for an alternative Bayesian derivation giving identical results and a user-friendly Excel calculator.
Two-sided P-values
If the P-values are two-sided, dividing the observed value of p1 by 2 yields the more significant of the two, corresponding, one-sided P-values. The prediction interval for this one-sided P-value is then calculated and converted back to a two-sided P-value using the equation ptwo-sided = min (1, 2 × pone-sided). Thus, the formula for one-sided P-values can be used to derive intervals for two-sided P-values.
Selection bias correction for the winner’s curse
If prediction interval(s) are constructed only for the most significant P-value(s) in a large study comprised of many tests, selection bias or the ‘winner’s curse’ will cause the above uncorrected intervals to be invalid. Previous work on P-value variability has not accounted for selection bias. To account for selection bias due to multiple testing in an initial study comprised of H tests, we applied a Bonferroni-style bias correction. Specifically, we use a corrected P-value, min(Hp1, 1), in place of p1. Bonferroni corrections are known to be conservative.12 In this case, the conservatism applies to the interval endpoints, both of which may be larger than necessary resulting in possible over-coverage of larger values and, at the same time, under-coverage of smaller values. For larger P-values, the selection bias correction is very conservative, especially when p1⩾1/H and the corrected P-value equals one. The correction is designed to avoid the effects of the winner’s curse. However, random P-value variability will always result in both ‘winners’ (overly small estimated P-values) and ‘losers’ (overly large estimated P-values). If K P-value prediction intervals are considered simultaneously and family-wise coverage of the intervals is desired, the non-coverage rate β should be replaced by β/K.
Results
To illustrate the random nature of P-values, we simulated values of −log p for 1000 independent realizations of a two-sided test of a SNP with a true OR of 1.74 in a sample of 2000 individuals. (All logs are base 10.) The P-values range from 0.015 to < 10−20 with a median of 10−8. Ten percent of P-values are < 10−11, while another 10% are >10−5. In other scenarios, the distribution’s shape and spread would differ depending on the true effect size, sample size and other factors. In a real study, we observe only one P-value per SNP without knowing where the rest of the distribution lies relative to it.
We applied our prediction interval formula to the simulated data above. For each of the 1000 P-values in Figure 1, we computed a separate 95% prediction interval (equal samples sizes, no selection bias correction) and calculated how many of the other 999 P-values it covered. The estimated coverage rate for our prediction interval procedure was 94.9%, consistent with theoretical results. By definition, the coverage rate is an average across the distribution of p1. Accordingly, the percentage of replication P-values covered varied, ranging from 17.3% for P = 0.015 to 99.6% for P = 10−8 (i.e. at the median).
Sampling distribution of −log p. Simulated independent values of −log p from 1000 identical studies of the same SNP in the same population using a one-sided test. Within each study, sample size = 2000, OR = 1.74, and risk to major homozygote = 9.5%. Genotype frequencies based on minor allele frequency of 30% under Hardy–Weinberg equilibrium. Dashed vertical line corresponds to P = 0.05. Overlaid curve is the theoretical exponential density under the null hypothesis that the OR = 1. Upper range of the density, which reaches 2.3 when −log P = 0, is not shown. SNP, single nucleotide polymorphism.
We next explored the properties of P-value variability in general, as revealed by the proposed prediction intervals. In Figure 2a, the initial and replication sample sizes are equal. Given an initial, observed P-value on the x axis, the prediction interval is the vertical line segment from the bottom to the top of the inner funnel. For an initial P-value of 10−5, the prediction interval (10−12, 0.05) barely excludes values of P>0.05. As the significance of the observed P-value increases, the prediction interval for the replication expands indicating increasing uncertainty on the log scale. However, intervals for larger P-values are also wide. For a P-value of 0.5, the prediction interval is (0.003, 0.997). The prediction intervals reveal both a lack of precision with regard to future P-values and a lack of resolution for comparing P-values from different hypothesis tests. For example, intervals for p1 = 10−3 vs 10−8 overlap substantially. This helps explain why genome-wide nonsignificant SNPs sometimes represent a substantial portion of potentially replicable, genuine associations.13
Ninety-five percent (95%) prediction intervals for replication P-values. A vertical line extending from the bottom to the top of the appropriate funnel gives the prediction interval given the original −log p on the x axis. P-values based on one-sided tests. Dashed lines at P = 0.05. Black arrows correspond to the two most significant SNP P-values observed by Furney et al. in an analysis of 939 individuals. Blue arrows correspond to the same P-values after applying a selection bias correction for the 488 911 SNPs tested in that study. (a) Prediction intervals with equal study sizes. Left (red) funnel: prediction intervals for a single test. Right (partially hidden, yellow funnel): prediction intervals with Bonferroni selection bias correction for picking the best of 1M tests in the initial study, Furney et al.8 (b) Prediction intervals with unequal study sizes and no selection bias correction. Bottom funnel: prediction intervals when replication study size is one-fourth the size of the original study. Top funnel: prediction intervals when replication study is four times the size of the original study. SNP, single nucleotide polymorphism.
If the P-value is one of many, a selection bias correction should be used to avoid the effects of the winner’s curse. The funnel on the right incorporates a correction for selecting the best SNP from 1M tests in an initial GWAS. The corrected funnel is identical to the one on the left, except that it has been shifted log(1M) = 6 units to the right. After correction for 1M tests, P must be < 10−6 for the prediction interval to exclude P = 1.
Figure 2b depicts replication studies with different sample sizes than the original study. If the replication study is smaller, the P-value distribution is shifted towards larger, less significant values. If the replication study is larger, the P-value distribution is shifted towards greater significance, even if the initial P-value was not, itself, statistically significant. Specifically, when the replication is one-fourth the size of the original study (bottom funnel), the prediction interval for a P-value of 10−14 is (10−9, 0.050). When the replication is four times the size of the original (top funnel), an initial P-value of 10−3 yields a prediction interval of (10−25, 0.050) which excludes values >0.05. When sample sizes differ, the initial P-value may lie outside the replication prediction interval.
Suppose a replication study with sample size n2 = 1000 seeks to replicate two previously published findings. One finding is a P-value of 10−13 from a study of size n1 = 4000. The other is a P-value of 0.001 from a study of size n1 = 250. The prediction intervals are (2.24 × 10−9, 0.069) and (2.21 × 10−27, 0.036), respectively. Despite appearances, the larger P-value from the smaller sample is a more promising target for replication. This suggests a problem for large discovery studies with several thousand subjects. Such studies are more likely to identify genome-wide significant P-values, even when the underlying effects have minimal clinical impact. What has previously been unappreciated is the difficulty of replicating results from such large studies, as subsequent replication studies are likely to have a much smaller sample.
To illustrate the interpretation of the intervals, we considered a recent GWAS on MRI atrophy measures and Alzheimer’s disease.8 Furney et al. reported two SNPs with associations reaching or approaching genome-wide significance in analyses of 488 911 SNPs and 939 individuals with complete data passing quality control. Specifically, they reported a disease-specific association of rs1925690 (nearest gene ZNF292 on chromosome 6) with entorhinal cortical volume (P = 2.56 × 10−8) and an association of rs11129640 (nearest gene ARPP-21 on chromosome 3) with entorhinal cortical thickness (P = 5.57 × 10−8). Two black arrows denote these P-values in Figure 2a. The prediction intervals for these two P-values are given by vertical lines crossing the upper red funnel at these two points. For an equal-sized replication study, the uncorrected intervals are (1.03 × 10−16, 3.73 × 10−3) and (3.27 × 10−16, 5.62 × 10−3), respectively. Two blue arrows point to the bias-corrected P-values for H = 488 911 SNPs. The upper red funnel is also used to determine the bias-corrected intervals, which are (2.68 × 10−7, 7.02 × 10−1) and (1.33 × 10−6, 8.02 × 10−1), respectively. The predicted medians are 0.013 and 0.027 after bias correction. Note that shifting a P-value to the left by log(H) achieves exactly the same bias correction as shifting the prediction funnel by log(H) to the right. In Figure 2b, the matching arrows point to the uncorrected and bias-corrected intervals for two possible alternative replication sample sizes. Figure 3 addresses the replication sample size question more completely, illustrating prediction intervals without and with selection bias correction for the ZNF292 SNP for sample sizes from 100–3000.
Ninety-five percent (95%) prediction intervals for the most significant P-value reported by Furney et al.8 as a function of replication sample size. Above funnel in red outline shows intervals without selection bias correction. Below funnel in yellow outline, which is partly hidden by the upper funnel, shows intervals with selection bias correction. Dashed line at P = 0.05.
We also examined recent findings of the PGC9 for attention deficit-hyperactivity disorder (ADHD), autism spectrum disorder (ASD), bipolar disorder (BPD), major depressive disorder (MDD) and schizophrenia. Table 1 shows the reported two-sided P-values and case-control counts for four SNPs with significant cross-disorder effects in Figure 2 of the PGC paper. We used the online calculator (Supplementary Information) to compute two sets of prediction intervals. The first set is for an identical replication of each study at its original sample size. For all SNPs, the individual disorder intervals overlap suggesting that a second, identical implementation of the original PGC design could have resulted in a different ranking of results due to chance alone, either for disorders within each SNP or for SNPs within each disorder. However, ADHD and ASD would be unlikely to yield any P-values < 10−8. In contrast, schizophrenia could reach such values for all SNPs. The second set of intervals is calibrated to remove effects of different sample sizes and case-control ratios. To do this, we calculated the effective sample size (ESS), which is the sample size needed to achieve equivalent power with a case:control ratio of 1. We then calculated intervals for replications with ESS = 16 957, the same as for schizophrenia in the largest original study. With sample size effects removed, the combined study is predicted to perform worse than some disorders and better than others. For example for rs2535629, the combined study is predicted to do less well than BPD, MDD and schizophrenia, but better than ADHD and ASD, suggesting that this SNP has greater associations with later onset disorders. Increased sample size makes it more likely that ADHD will achieve significant results, but the intervals widen and nonsignificant values cannot be excluded. Most notably for rs2799573, ADHD had only the third most significant P-value, but has the highest prediction of replication success for equal sample sizes. On the other hand, ASD is unlikely to yield P-values < 10−8 for these SNPs even in a study as big as the schizophrenia study.
Discussion
This study has presented prediction intervals for replication P-values, demonstrating the large scale of P-value variability. The scale of P-value variability cannot be attributed to additional, potentially addressable, factors that also decrease a study’s reliability such as poor study design, bias and/or uncertainties in the underlying assumptions, inadequate phenotype measurements, inappropriate statistical models or methods, population heterogeneity, variable penetrance, publication bias and the ‘winner’s curse’.14, 15, 16, 17, 18, 19, 20 Although the intervals require no assumptions about unknown effect sizes or the initial sample size, they are strongly affected by the relative size of the replication and initial studies. While large P-value variability has been previously recognized, it is often disregarded in practice. GWAS and other studies rely on a single sample of data, yielding only one observed P-value per test out of a large range of possible P-values. Furthermore, the more significant a P-value is, the less the precision that can be attached to it. While uncertainty is high for a P-value from a single test, P-values obtained from GWAS, or other multiplexed studies requiring multiple testing corrections, provide almost no information with which to make future predictions.
The degree of statistical uncertainty associated with P-values is not specific to the P-value itself, which is just a convenient measure of the evidence provided by the data against the null hypothesis. Rather, uncertainty is an intrinsic property of statistical sampling. Noting the low precision provided by P-values, Cumming recommended relying more on effect size confidence intervals.5 We agree that confidence intervals give more information about effect sizes than P-values and should always be reported when possible. It should be noted, however, that whenever a 95% confidence interval excludes the null value of a parameter, there is a corresponding P-value less than 0.05. Thus, there is a one-to-one correspondence between decision rules based on confidence intervals and decision rules based on P-values. Additionally, confidence intervals do not provide a one-dimensional summary of a statistical result and, therefore, are more difficult to compare.
The present results may appear to contradict our previous findings on the consequences of the extreme P-values needed for genome-wide significance.21 That study showed that the increased stringency of the significance level associated with increasing the number of hypothesis tests requires relatively modest increases in sample size or effect size. In contrast, the present study shows that, after a study is complete, any observed P-value, no matter how small, is a very imprecise predictor of future performance. Thus, studies can have good power to reject the null hypothesis of no association, while providing little information with respect to the reproducibility or relative strength of the true association.
The findings of this study support proposals by others to combine multiple lines of biological evidence in deciding which results to investigate further, rather than depending on association P-values alone.22, 23, 24, 25, 26, 27 Our findings may also lead to improved multi-stage SNP/subject selection strategies potentially using small, low-cost studies to screen large numbers of genetic variants, followed by larger, targeted replication studies to reach firmer conclusions. Similar strategies have been applied successfully in phase I-IV clinical trials for drug development.28 More immediately, the proposed prediction intervals provide practical information for designing replication studies about the relative sample size needed to expect successful confirmation of a previously reported P-value. Most importantly, the present findings caution against unrealistic expectations for replication based on P-value size, as the apparent numerical precision can be misleading. These results may better enable scientific investigators and consumers to evaluate P-values in their own studies and those reported by others.
References
Pearson T, Manolio T . How to interpret a genome-wide association study. JAMA 2008; 299: 1335–1344.
Nature genetics editorial. Asking for more. Nat Genet 2012; 44: 733.
Goodman S . A comment on replication, P-values and evidence. Stat Med 1992; 11: 875–879.
Senn S . Letter to the editor: a comment on replication, P-values and evidence. Stat Med 2002; 21: 2437–2444.
Cumming G . Replication and P intervals: P values predict the future only vaguely, but confidence intervals do much better. Perspect Psychol Sci 2008; 3: 286–300.
Boos D, Stefanski S . P-value precision and reproducibility. Am Stat 2012; 65: 213–221.
Mojirsheibani M, Tibshirani R . Some results on bootstrap prediction intervals. Can J Stat 1996; 24: 549–568.
Furney SJ, Simmons A, Breen G, Pedroso I, Lunnon K, Proitsi P et al. Genome-wide association with MRI atrophy measures as a quantitative trait locus for Alzheimer’s disease. Mol Psychiatry 2011; 16: 1130–1138.
Cross-Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 2013; 381: 1371–1379.
Cox D Hinkley D . Theoretical Statistics. Chapman & Hall: London, UK, 1974, pp 242–244.
Whitmore G . Prediction limits for a univariate normal observation. Am Stat 1986; 40: 141–143.
Lazzeroni L, Lange K . A conditional inference framework for extending the transmission/disequilibrium test. Hum Hered 1998; 48: 67–81.
Panagiotou O, Ioannidis J . What should the genome-wide significance threshold be? Empirical replication of borderline genetic associations. Int J Epidemiol 2011; 41: 273–286.
Dickersin K . The existence of publication bias and risk factors for its occurrence. JAMA 1990; 263: 1385–1389.
Easterbrook P, Berlin J, Gopalan R, Matthews D . Publication bias in clinical research. Lancet 1991; 337: 867–872.
Ioannidis J . Why most published research findings are false. PLoS Med 2005; 2: e124.
Zollner S, Pritchard J . Overcoming the winner’s curse: estimating penetrance parameters from case-control data. Am J Hum Genet 2007; 80: 605–615.
Moonesinghe R, Khoury M, Liu T, Ioannidis J . Required sample size and nonreplicability thresholds for heterogeneous genetic associations. Proc Natl Acad Sci USA 2008; 105: 617–622.
Lazzeroni LC, Ray A . A generalized DeFries-Fulker regression framework for the analysis of twin data. Behav Genet 2013; 43: 85–96.
Lazzeroni LC . Evaluating the evidence of replication of genetic associations with schizophrenia. JAMA Psych 2014; 71: 94–95.
Lazzeroni L, Ray A . The cost of large number of hypothesis tests on power, effect size and sample size. Mol Psychiatry 2012; 17: 108–114.
Wang K, Li M, Bucan M . Pathway-based approaches for analysis of genome-wide association studies. Am J Hum Genet 2007; 81: 1278–1283.
Huang D, Sherman B, Lempicki R . Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 2008; 37: 1–13.
Elbers C, van Eijk K, Franke L, Mulder F, van der Schouw YT, Wijmenga C et al. Using genome-wide pathway analysis to unravel the etiology of complex diseases. Genet Epidemiol 2009; 33: 419–431.
Medina I, Montaner D, Bonifaci N, Pujana MA, Carbonell J, Tarraga J et al. Gene set-based analysis of polymorphisms: finding pathways or biological processes associated with traits in genome-wide association studies. Nucleic Acids Res 2009; 37: W340–W344.
Cantor R, Lange K, Sinsheimer J . Prioritizing GWAS results: a review of statistical methods and recommendations for their applcation. Am J Hum Genet 2010; 86: 6–22.
Moreau Y, Tranchevent L-C . Computational tools for prioritazing candidate genes: boosting disease gene discovery. Nat Genet Rev 2012; 13: 523–526.
Harrington D . The randomized clinical trial. J Am Stat Assoc 2000; 95: 312–315.
Acknowledgements
This work was supported by the Clinical Science Research & Development Service of the Department of Veterans Affairs (A Cooperative Studies Program-Wide DNA Bank, CSP#478), and the National Institutes of Health (R01 MH086135) and the Consortium on the Genetics of Schizophrenia (COGS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Lazzeroni, L., Lu, Y. & Belitskaya-Lévy, I. P-values in genomics: Apparent precision masks high uncertainty. Mol Psychiatry 19, 1336–1340 (2014). https://doi.org/10.1038/mp.2013.184
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2013.184
Keywords
This article is cited by
-
Enhancing genomic prediction with genome-wide association studies in multiparental maize populations
Heredity (2017)
-
Bayesian prediction intervals for assessing P-value variability in prospective replication studies
Translational Psychiatry (2017)
-
A genetic factor associated with low final bone mineral density in children after a long-term glucocorticoids treatment
The Pharmacogenomics Journal (2017)
-
Solutions for quantifying P-value uncertainty and replication power
Nature Methods (2016)
-
Cross-Disorder Psychiatric Genomics
Current Behavioral Neuroscience Reports (2016)