Abstract
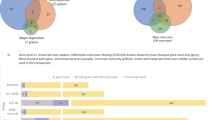
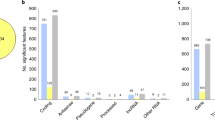
RNA-sequencing (RNA-seq) is a powerful technique to investigate the complexity of gene expression in the human brain. We used RNA-seq to survey the brain transcriptome in high-quality postmortem dorsolateral prefrontal cortex from 11 individuals diagnosed with bipolar disorder (BD) and from 11 age- and gender-matched controls. Deep sequencing was performed, with over 350 million reads per specimen. At a false discovery rate of <5%, we detected five differentially expressed (DE) genes and 12 DE transcripts, most of which have not been previously implicated in BD. Among these, Prominin 1/CD133 and ATP-binding cassette-sub-family G-member2 (ABCG2) have important roles in neuroplasticity. We also show for the first time differential expression of long noncoding RNAs (lncRNAs) in BD. DE transcripts include those of serine/arginine-rich splicing factor 5 (SRSF5) and regulatory factor X4 (RFX4), which along with lncRNAs have a role in mammalian circadian rhythms. The DE genes were significantly enriched for several Gene Ontology categories. Of these, genes involved with GTPase binding were also enriched for BD-associated SNPs from previous genome-wide association studies, suggesting that differential expression of these genes is not simply a consequence of BD or its treatment. Many of these findings were replicated by microarray in an independent sample of 60 cases and controls. These results highlight common pathways for inherited and non-inherited influences on disease risk that may constitute good targets for novel therapies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Kim S, Webster MJ . The Stanley Neuropathology Consortium integrative database: a novel, web-based tool for exploring neuropathological markers in psychiatric disorders and the biological processes associated with abnormalities of those markers. Neuropsychopharmacology 2010; 35: 473–482.
Ryan MM, Lockstone HE, Huffaker SJ, Wayland MT, Webster MJ, Bahn S . Gene expression analysis of bipolar disorder reveals downregulation of the ubiquitin cycle and alterations in synaptic genes. Mol Psychiatry 2006; 11: 965–978.
Konradi C, Eaton M, MacDonald ML, Walsh J, Benes FM, Heckers S . Molecular evidence for mitochondrial dysfunction in bipolar disorder. Arch Gen Psychiatry 2004; 61: 300–308.
Nakatani N, Hattori E, Ohnishi T, Dean B, Iwayama Y, Matsumoto I et al Genome-wide expression analysis detects eight genes with robust alterations specific to bipolar I disorder: relevance to neuronal network perturbation. Hum Mol Genet 2006; 15: 1949–1962.
Elashoff M, Higgs BW, Yolken RH, Knable MB, Weis S, Webster MJ et al Meta-analysis of 12 genomic studies in bipolar disorder. J Mol Neurosci 2007; 31: 221–243.
Wang Z, Gerstein M, Snyder M . RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 2009; 10: 57–63.
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ et al Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 2010; 28: 511–515.
Lipska BK, Deep-Soboslay A, Weickert CS, Hyde TM, Martin CE, Herman MM et al Critical factors in gene expression in postmortem human brain: focus on studies in schizophrenia. Biol Psychiatry 2006; 60: 650–658.
Trapnell C, Pachter L, Salzberg SL . TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 2009; 25: 1105–1111.
Langmead B, Trapnell C, Pop M, Salzberg SL . Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 2009; 10: R25.
Roberts A, Pachter L . Streaming fragment assignment for real-time analysis of sequencing experiments. Nat Methods 2013; 10: 71–73.
Anders S, Huber W . Differential expression analysis for sequence count data. Genome Biol 2010; 11: R106.
Liptak T . On the combination of independent tests. Magyar Tud. Akad. Mat. Kutato Int. Kozl 1958; 3: 171–197.
Huang da W, Sherman BT, Lempicki RA . Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009; 4: 44–57.
Huang da W, Sherman BT, Lempicki RA . Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 2009; 37: 1–13.
Wang K, Li M, Bucan M . Pathway-based approaches for analysis of genomewide association studies. Am J Hum Genet 2007; 81: 1278–1283.
Voineagu I, Wang X, Johnston P, Lowe JK, Tian Y, Horvath S et al Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature 2011; 474: 380–384.
Chen DT, Jiang X, Akula N, Shugart YY, Wendland JR, Steele CJ et al Genome-wide association study meta-analysis of European and Asian-ancestry samples identifies three novel loci associated with bipolar disorder. Mol Psychiatry 2013; 18: 195–205.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D et al PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007; 81: 559–575.
Veyrieras JB, Kudaravalli S, Kim SY, Dermitzakis ET, Gilad Y, Stephens M et al High-resolution mapping of expression-QTLs yields insight into human gene regulation. PLoS Genet 2008; 4: e1000214.
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D et al Circos: an information aesthetic for comparative genomics. Genome Res 2009; 19: 1639–1645.
Goldstein I, Lerer E, Laiba E, Mallet J, Mujaheed M, Laurent C et al Association between sodium- and potassium-activated adenosine triphosphatase alpha isoforms and bipolar disorders. Biol Psychiatry 2009; 65: 985–991.
Chetcuti A, Adams LJ, Mitchell PB, Schofield PR . Microarray gene expression profiling of mouse brain mRNA in a model of lithium treatment. Psychiatr Genet 2008; 18: 64–72.
Grimes CA, Jope RS . Cholinergic stimulation of early growth response-1 DNA binding activity requires protein kinase C and mitogen-activated protein kinase kinase activation and is inhibited by sodium valproate in SH-SY5Y cells. J Neurochem 1999; 73: 1384–1392.
Kim S, Choi KH, Baykiz AF, Gershenfeld HK . Suicide candidate genes associated with bipolar disorder and schizophrenia: an exploratory gene expression profiling analysis of post-mortem prefrontal cortex. BMC Genomics 2007; 8: 413.
Matigian N, Windus L, Smith H, Filippich C, Pantelis C, McGrath J et al Expression profiling in monozygotic twins discordant for bipolar disorder reveals dysregulation of the WNT signalling pathway. Mol Psychiatry 2007; 12: 815–825.
Perez-Santiago J, Diez-Alarcia R, Callado LF, Zhang JX, Chana G, White CH et al A combined analysis of microarray gene expression studies of the human prefrontal cortex identifies genes implicated in schizophrenia. J Psychiatr Res 2012; 46: 1464–1474.
LeBlanc M, Kulle B, Sundet K, Agartz I, Melle I, Djurovic S et al Genome-wide study identifies PTPRO and WDR72 and FOXQ1-SUMO1P1 interaction associated with neurocognitive function. J Psychiatr Res 2012; 46: 271–278.
Soria V, Martinez-Amoros E, Escaramis G, Valero J, Perez-Egea R, Garcia C et al Differential association of circadian genes with mood disorders: CRY1 and NPAS2 are associated with unipolar major depression and CLOCK and VIP with bipolar disorder. Neuropsychopharmacology 2010; 35: 1279–1289.
Vacic V, McCarthy S, Malhotra D, Murray F, Chou HH, Peoples A et al Duplications of the neuropeptide receptor gene VIPR2 confer significant risk for schizophrenia. Nature 2011; 471: 499–503.
Florek M, Haase M, Marzesco AM, Freund D, Ehninger G, Huttner WB et al Prominin-1/CD133, a neural and hematopoietic stem cell marker, is expressed in adult human differentiated cells and certain types of kidney cancer. Cell Tissue Res 2005; 319: 15–26.
Islam MO, Kanemura Y, Tajria J, Mori H, Kobayashi S, Hara M et al Functional expression of ABCG2 transporter in human neural stem/progenitor cells. Neurosci Res 2005; 52: 75–82.
Coles EG, Lawlor ER, Bronner-Fraser M . EWS-FLI1 causes neuroepithelial defects and abrogates emigration of neural crest stem cells. Stem Cells 2008; 26: 2237–2244.
Coon SL, Munson PJ, Cherukuri PF, Sugden D, Rath MF, Moller M et al Circadian changes in long noncoding RNAs in the pineal gland. Proc Natl Acad Sci USA 2012; 109: 13319–13324.
Millar JK, James R, Brandon NJ, Thomson PA . DISC1 and DISC2: discovering and dissecting molecular mechanisms underlying psychiatric illness. Ann Med 2004; 36: 367–378.
Williams JM, Beck TF, Pearson DM, Proud MB, Cheung SW, Scott DA . A 1q42 deletion involving DISC1, DISC2, and TSNAX in an autism spectrum disorder. Am J Med Genet A 2009; 149A: 1758–1762.
Qureshi IA, Mattick JS, Mehler MF . Long non-coding RNAs in nervous system function and disease. Brain Res 2010; 1338: 20–35.
Lin M, Pedrosa E, Shah A, Hrabovsky A, Maqbool S, Zheng D et al RNA-Seq of human neurons derived from iPS cells reveals candidate long non-coding RNAs involved in neurogenesis and neuropsychiatric disorders. PLoS ONE 2011; 6: e23356.
Wapinski O, Chang HY . Long noncoding RNAs and human disease. Trends Cell Biol 2011; 21: 354–361.
Wang H, Liu Y, Briesemann M, Yan J . Computational analysis of gene regulation in animal sleep deprivation. Physiol Genomics 2010; 42: 427–436.
Yang S, Wang K, Valladares O, Hannenhalli S, Bucan M . Genome-wide expression profiling and bioinformatics analysis of diurnally regulated genes in the mouse prefrontal cortex. Genome Biol 2007; 8: R247.
Glatt SJ, Everall IP, Kremen WS, Corbeil J, Sasik R, Khanlou N et al Comparative gene expression analysis of blood and brain provides concurrent validation of SELENBP1 up-regulation in schizophrenia. Proc Natl Acad Sci USA 2005; 102: 15533–15538.
Lips ES, Cornelisse LN, Toonen RF, Min JL, Hultman CM, Holmans PA et al Functional gene group analysis identifies synaptic gene groups as risk factor for schizophrenia. Mol Psychiatry 2012; 17: 996–1006.
McGlincy NJ, Valomon A, Chesham JE, Maywood ES, Hastings MH, Ule J . Regulation of alternative splicing by the circadian clock and food related cues. Genome Biol 2012; 13: R54.
Araki R, Takahashi H, Fukumura R, Sun F, Umeda N, Sujino M et al Restricted expression and photic induction of a novel mouse regulatory factor X4 transcript in the suprachiasmatic nucleus. J Biol Chem 2004; 279: 10237–10242.
Glaser B, Kirov G, Bray NJ, Green E, O'Donovan MC, Craddock N et al Identification of a potential bipolar risk haplotype in the gene encoding the winged-helix transcription factor RFX4. Mol Psychiatry 2005; 10: 920–927.
Fogel BL, Wexler E, Wahnich A, Friedrich T, Vijayendran C, Gao F et al RBFOX1 regulates both splicing and transcriptional networks in human neuronal development. Hum Mol Genet 2012; 21: 4171–4186.
Szatmari P, Paterson AD, Zwaigenbaum L, Roberts W, Brian J, Liu XQ et al Mapping autism risk loci using genetic linkage and chromosomal rearrangements. Nat Genet 2007; 39: 319–328.
Walsh T, McClellan JM, McCarthy SE, Addington AM, Pierce SB, Cooper GM et al Rare structural variants disrupt multiple genes in neurodevelopmental pathways in schizophrenia. Science 2008; 320: 539–543.
Baum AE, Akula N, Cabanero M, Cardona I, Corona W, Klemens B et al A genome-wide association study implicates diacylglycerol kinase eta (DGKH) and several other genes in the etiology of bipolar disorder. Mol Psychiatry 2008; 13: 197–207.
Zhang C, Zhang Z, Castle J, Sun S, Johnson J, Krainer AR et al Defining the regulatory network of the tissue-specific splicing factors Fox-1 and Fox-2. Genes Dev 2008; 22: 2550–2563.
Niciu MJ, Ionescu DF, Mathews DC, Richards EM, Zarate CA . Second messenger/signal transduction pathways in major mood disorders: moving from membrane to mechanism of action, part II: bipolar disorder. CNS Spectr 2013; 11: 1–10.
Oshlack A, Robinson MD, Young MD . From RNA-seq reads to differential expression results. Genome Biol 2010; 11: 220.
McCullumsmith RE, Meador-Woodruff JH . Novel approaches to the study of postmortem brain in psychiatric illness: old limitations and new challenges. Biol Psychiatry 2011; 69: 127–133.
Price JL, Drevets WC . Neurocircuitry of mood disorders. Neuropsychopharmacology 2010; 35: 192–216.
Acknowledgements
This study was supported by the Intramural Research Program of the NIMH (grant no. ZIAMH002810 and K99MH085098). RNA from postmortem brain tissue was donated by The Stanley Medical Research Institute’s brain collection courtesy of Drs Michael B Knable, E Fuller Torrey, Maree J Webster, Serge Weis and Robert H Yolken. Data analysis was done on the high-performance computational Biowulf cluster at NIH.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
Rights and permissions
About this article
Cite this article
Akula, N., Barb, J., Jiang, X. et al. RNA-sequencing of the brain transcriptome implicates dysregulation of neuroplasticity, circadian rhythms and GTPase binding in bipolar disorder. Mol Psychiatry 19, 1179–1185 (2014). https://doi.org/10.1038/mp.2013.170
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2013.170
Keywords
This article is cited by
-
A resource of induced pluripotent stem cell (iPSC) lines including clinical, genomic, and cellular data from genetically isolated families with mood and psychotic disorders
Translational Psychiatry (2023)
-
Differential expression of gene co-expression networks related to the mTOR signaling pathway in bipolar disorder
Translational Psychiatry (2022)
-
Amygdala and anterior cingulate transcriptomes from individuals with bipolar disorder reveal downregulated neuroimmune and synaptic pathways
Nature Neuroscience (2022)
-
A Novel Long-Noncoding RNA LncZFAS1 Prevents MPP+-Induced Neuroinflammation Through MIB1 Activation
Molecular Neurobiology (2022)
-
Exact transcript quantification over splice graphs
Algorithms for Molecular Biology (2021)