Abstract
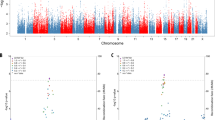
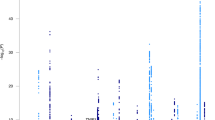
The risk of Alzheimer's disease (AD) is strongly determined by genetic factors and recent genome-wide association studies (GWAS) have identified several genes for the disease risk. In addition to the disease risk, age-at-onset (AAO) of AD has also strong genetic component with an estimated heritability of 42%. Identification of AAO genes may help to understand the biological mechanisms that regulate the onset of the disease. Here we report the first GWAS focused on identifying genes for the AAO of AD. We performed a genome-wide meta-analysis on three samples comprising a total of 2222 AD cases. A total of ∼2.5 million directly genotyped or imputed single-nucleotide polymorphisms (SNPs) were analyzed in relation to AAO of AD. As expected, the most significant associations were observed in the apolipoprotein E (APOE) region on chromosome 19 where several SNPs surpassed the conservative genome-wide significant threshold (P<5E-08). The most significant SNP outside the APOE region was located in the DCHS2 gene on chromosome 4q31.3 (rs1466662; P=4.95E-07). There were 19 additional significant SNPs in this region at P<1E-04 and the DCHS2 gene is expressed in the cerebral cortex and thus is a potential candidate for affecting AAO in AD. These findings need to be confirmed in additional well-powered samples.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Slooter AJ, Cruts M, Kalmijn S, Hofman A, Breteler MM, Van Broeckhoven C et al. Risk estimates of dementia by apolipoprotein E genotypes from a population-based incidence study: the Rotterdam Study. Arch Neurol 1998; 55: 964–968.
Seshadri S, Drachman DA, Lippa CF . Apolipoprotein E epsilon 4 allele and the lifetime risk of Alzheimer's disease. What physicians know, and what they should know. Arch Neurol 1995; 52: 1074–1079.
Lambert JC, Heath S, Even G, Campion D, Sleegers K, Hiltunen M et al. Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer's disease. Nat Genet 2009; 41: 1094–1099.
Harold D, Abraham R, Hollingworth P, Sims R, Gerrish A, Hamshere ML et al. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer's disease. Nat Genet 2009; 41: 1088–1093.
Seshadri S, Fitzpatrick AL, Ikram MA, DeStefano AL, Gudnason V, Boada M et al. Genome-wide analysis of genetic loci associated with Alzheimer disease. JAMA 2010; 303: 1832–1840.
Hollingworth P, Harold D, Sims R, Gerrish A, Lambert JC, Carrasquillo MM et al. Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer's disease. Nat Genet 2011; 43: 429–435.
Naj AC, Jun G, Beecham GW, Wang LS, Vardarajan BN, Buros J et al. Common variants at MS4A4/MS4A6E, CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer's disease. Nat Genet 2011; 43: 436–441.
Gatz M, Reynolds CA, Fratiglioni L, Johansson B, Mortimer JA, Berg S et al. Role of genes and environments for explaining Alzheimer disease. Arch Gen Psychiatry 2006; 63: 168–174.
Daw EW, Payami H, Nemens EJ, Nochlin D, Bird TD, Schellenberg GD et al. The number of trait loci in late-onset Alzheimer disease. Am J Hum Genet 2000; 66: 196–204.
Li YJ, Scott WK, Hedges DJ, Zhang F, Gaskell PC, Nance MA et al. Age at onset in two common neurodegenerative diseases is genetically controlled. Am J Hum Genet 2002; 70: 985–993.
Kamboh MI, Minster RL, Kenney M, Ozturk A, Desai OO, Kammerer CM et al. Alpha-1-antichymotrypsin (ACT or SERPINA3) polymorphom may affect aga-at-onset and disease duration of Alzheimer's disease. Neurobiol Aging 2006; 27: 1435–1439.
Lambert JC, Sleegers K, González-Pérez A, Ingelsson M, Beecham GW, Hiltunen M et al. The CALHM1 P86L polymorphism is a genetic modifier of age at onset in Alzheimer's disease: a meta-analysis study. J Alzheimers Dis 2010; 22: 247–255.
Bertram L, Lange C, Mullin K, Parkinson M, Hsiao M, Hogan MF et al. Genome-wide association analysis reveals putative Alzheimer's disease susceptibility loci in addition to APOE. Am J Hum Genet 2008; 83: 623–632.
Burns LC, Minster RL, Demirci FY, Barmada MM, Ganguli M, Lopez OL et al. Replication study of genome-wide associated SNPs with late-onset Alzheimer's disease. Am J Med Genet B Neuropsychiatr Genet 2011; 156: 507–512.
Carrasquillo MM, Zou F, Pankratz VS, Wilcox SL, Ma L, Walker LP et al. Genetic variation in PCDH11X is associated with susceptibility to late-onset Alzheimer's disease. Nat Genet 2009; 41: 192–198.
Saykin AJ, Shen L, Foroud TM, Potkin SG, Swaminathan S, Kim S et al. Alzheimer's Disease Neuroimaging Initiative biomarkers as quantitative phenotypes: genetics core aims, progress, and plans. Alzheimers Dement 2010; 6: 265–273.
Aisen PS, Petersen RC, Donohue MC, Gamst A, Raman R, Thomas RG et al. Clinical Core of the Alzheimer's Disease Neuroimaging Initiative: progress and plans. Alzheimers Dement 2010; 6: 239–246.
Petersen RC, Aisen PS, Beckett LA, Donohue MC, Gamst AC, Harvey DJ et al. Alzheimer's Disease Neuroimaging Initiative (ADNI): clinical characterization. Neurology 2010; 74: 201–209.
Kamboh MI, Aston CE, Hamman RF . The relationship of APOE polymorphism and cholesterol levels in normoglycemic and diabetic subjects in a biethnic population from the San Luis Valley, Colorado. Atherosclerosis 1995; 112: 145–159.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D et al. PLINK: a toolset for whole-genome association and population-based linkage analysis. Am J Hum Genet 2007; 81: 559–575.
Brockwell SE, Gordon IR . A comparison of statistical methods for meta-analysis. Stat Med 2001; 20: 825–840.
Higgins JP, Thompson SG, Deeks JJ, Altman DG . Measuring inconsistency in meta-analyses. BMJ 2003; 327: 557–560.
Höng JC, Ivanov NV, Hodor P, Xia M, Wei N, Blevins R et al. Identification of new human cadherin genes using a combination of protein motif search and gene finding methods. J Mol Biol 2004; 337: 307–317.
McClay JL, Adkins DE, Aberg K, Stroup S, Perkins DO, Vladimirov VI et al. Genome-wide pharmacogenomic analysis of response to treatment with antipsychotics. Mol Psychiatry 2011; 16: 76–85.
Coultas L, Terzano S, Thomas T, Voss A, Reid K, Stanley EG et al. Hrk/DP5 contributes to the apoptosis of select neuronal populations but is dispensable for haematopoietic cell apoptosis. J Cell Sci 2007; 120: 2044–2052.
Clark ME, Kelner GS, Turbeville LA, Boyer A, Arden KC, Maki RA . ADAMTS9, a novel member of the ADAM-TS/ metallospondin gene family. Genomics 2000; 67: 343–350.
Zeggini E, Scott LJ, Saxena R, Voight BF, Marchini JL, Hu T et al. Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat Genet 2008; 40: 638–645.
Sakai K, Tiebel O, Ljungberg MC, Sullivan M, Lee HJ, Terashima T et al. A neuronal VLDLR variant lacking the third complement-type repeat exhibits high capacity binding of apoE containing lipoproteins. Brain Res 2009; 1276: 11–21.
Helbecque N, Berr C, Cottel D, Fromentin-David I, Sazdovitch V, Ricolfi F et al. VLDL receptor polymorphism, cognitive impairment, and dementia. Neurology 2001; 56: 1183–1188.
Lachman HM, Fann CS, Bartzis M, Evgrafov OV, Rosenthal RN, Nunes EV et al. Genome-wide suggestive linkage of opioid dependence to chromosome 14q. Hum Mol Genet 2007; 16: 1327–1334.
Heard-Costa NL, Zillikens MC, Monda KL, Johansson A, Harris TB, Fu M et al. NRXN3 is a novel locus for waist circumference: a genome-wide association study from the CHARGE Consortium. PLoS Genet 2009; 6: e1000539.
Shen L, Kim S, Risacher SL, Nho K, Swaminathan S, West JD et al. Whole genome association study of brain-wide imaging phenotypes for identifying quantitative trait loci in MCI and AD: a study of the ADNI cohort. Neuroimage 2010; 53: 1051–1063.
Acknowledgements
This study was supported by the National Institute on Aging grants AG030653 (MIK), AG005133 (ADRC), AG027224 (RAS) and AG18023 (SGY). We thank Drs Peter Gregersen and Annette Lee of the Feinstein Institute of Medical Research where the genotyping of the discovery sample was performed. We acknowledge Drs Neill R Graff-Radford, Dennis W Dickson and Ronald C Petersen for their key contribution in collecting the Mayo replication sample. One of the replication data sets was funded by the Alzheimer's Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904; RC2 AG036535). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: Abbott, AstraZeneca AB, Bayer Schering Pharma AG, Bristol-Myers Squibb, Eisai Global Clinical Development, Elan Corporation, Genentech, GE Healthcare, GlaxoSmithKline, Innogenetics, Johnson and Johnson, Eli Lilly and Co, Medpace, Inc, Merck and Co, Inc, Novartis AG, Pfizer Inc, F Hoffman-La Roche, Schering-Plough, Synarc, Inc, as well as non-profit partners the Alzheimer's Association and Alzheimer's Drug Discovery Foundation, with participation from the US Food and Drug Administration. Private sector contributions to ADNI are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer's Disease Cooperative Study at the University of California, San Diego. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of California, Los Angeles. This research was also supported by NIH grants P30 AG010129, K01 AG030514 and the Dana Foundation. Some sample collection was supported by the NIA National Cell Repository for Alzheimer's Disease (NCRAD; U24 AG021886) and additional support for data analysis was provided by R01 AG19771 and P30 AG10133. We thank the following people for their contributions to the ADNI genotyping project: Tatiana Foroud, Kelley Faber, Li Shen, Steven G Potkin, Matthew J Huentelman, David W Craig, Sungeun Kim, Kwangsik Nho and Bryan M DeChairo.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
Appendix
Appendix
Data used in preparation of this article were obtained from the Alzheimer's Disease Neuroimaging Initiative (ADNI) database (http://www.adni.loni.ucla.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data, but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at http://adni.loni.ucla.edu/wp-content/uploads/how_to_apply/ADNI_Authorship_List.pdf
Rights and permissions
About this article
Cite this article
Kamboh, M., Barmada, M., Demirci, F. et al. Genome-wide association analysis of age-at-onset in Alzheimer's disease. Mol Psychiatry 17, 1340–1346 (2012). https://doi.org/10.1038/mp.2011.135
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2011.135
Keywords
This article is cited by
-
The Role of Changes in the Expression of Inflammation-Associated Genes in Cerebral Small Vessel Disease with Cognitive Impairments
Neuroscience and Behavioral Physiology (2024)
-
Six genetically linked mutations in the CD36 gene significantly delay the onset of Alzheimer's disease
Scientific Reports (2022)
-
Genomics and Functional Genomics of Alzheimer's Disease
Neurotherapeutics (2022)
-
A machine learning approach to brain epigenetic analysis reveals kinases associated with Alzheimer’s disease
Nature Communications (2021)
-
Risk prediction of late-onset Alzheimer’s disease implies an oligogenic architecture
Nature Communications (2020)