Abstract
The first generation of genome-wide association studies (GWA studies) for psychiatric disorders has led to new insights regarding the genetic architecture of these disorders. We now start to realize that a larger number of genes, each with a small contribution, are likely to explain the heritability of psychiatric diseases. The contribution of a large number of genes to complex traits can be analyzed with genome-wide profiling. In a discovery sample, a genetic risk profile for depression was defined based on a GWA study of 1738 adult cases and 1802 controls. The genetic risk scores were tested in two population-based samples of elderly participants. The genetic risk profiles were evaluated for depression and anxiety in the Rotterdam Study cohort and the Erasmus Rucphen Family (ERF) study. The genetic risk scores were significantly associated with different measures of depression and explained up to ∼0.7% of the variance in depression in Rotterdam Study and up to ∼1% in ERF study. The genetic score for depression was also significantly associated with anxiety explaining up to 2.1% in Rotterdam study. These findings suggest the presence of many genetic loci of small effect that influence both depression and anxiety. Remarkably, the predictive value of these profiles was as large in the sample of elderly participants as in the middle-aged samples.
Similar content being viewed by others
Introduction
Genetic factors have an important role in the susceptibility to depression. A meta-analysis of twin studies on major depressive disorder (MDD) estimated the heritability at 37%.1 However, the success of studies aiming to find genes underlying the vulnerability for depression has been limited. An overview of promising results of linkage studies on MDD and neuroticism, a related personality trait, shows some overlap in regions of interest, but, so far, no single locus has been identified.2 Candidate gene studies, mostly focusing on genes involved in neurotransmitter circuits or in reactions to stress, have also not been able to unambiguously identify a genetic variants explaining differences in the vulnerability for depression.3, 4
An important issue in research on the etiological factors of MDD has been the frequent comorbidity with anxiety disorders. In the National Comorbidity Survey Replication, 59% of the subjects with a lifetime diagnosis of MDD also fulfilled the criteria for a lifetime anxiety disorder diagnosis.5 A review of twin and family studies indicated that this comorbidity might be explained by shared, mostly genetic factors.6 Still, an overview of promising results of linkage studies of anxiety only showed one overlapping region of interest with MDD and neuroticism.2
The recent success of genome-wide association (GWA) studies has fueled expectations on finding genes for MDD. One of the first GWA studies of depression showed evidence for the role of the presynaptic protein piccolo (PCLO) gene.7, 8 Recently, this result was replicated with a P-value of 2 × 10−9 in a meta-analysis of the results in three population-based samples, but not when five clinical samples were also included.7, 9 However, the first GWA studies of MDD as well as those of other psychiatric phenotypes have also shown that genome-wide significant findings are rare and explain a small part of heritability.7, 10, 11 This might be due to a lack of power. The Genetic Association Information Network (GAIN)-MDD GWA study, for example, including ∼1700 cases and 1800 controls, had 80% statistical power to detect relative risks of 1.59, 1.40 and 1.35 with a P-value of 1 × 10−7, for minor allele frequencies of 0.10, 0.25 and 0.40. This is well comparable to other first generation GWA studies. However, the first results of GWA studies suggest that the strongest odds ratios may be <1.2.12 Another explanation for the scarce genome-wide significant findings could be that there is not a distinct number of genes for MDD with moderate-to-small risks but rather a large number of variations spread over the genome, each with small effects. Such a polygenic model predicts that the more markers are used, the better the disease is predicted and it implies that everybody carries risk variants but patients carry more than non-diseased people. We examined if a polygenetic component influences MDD implying that a large number of genetic variants are involved in explaining its heritability.
The evidence for a polygenic origin has recently been examined for schizophrenia and the hypothesis of a polygenic component was directly tested using GWA data.13 In this approach, the joint effect of multiple single-nucleotide polymorphisms (SNPs) are tested rather than the effects of individual SNPs. These individual SNPs are not required to reach a genome-wide significance level by themselves. This approach aims to test whether the genetic disease liability reflects, at least in part, the additive effect of a large number of variants spread across the genome whose joint action may be captured in a genome-wide genetic risk score. To obtain this risk score, a discovery set is used to select SNPs based on specific P-value thresholds (for example, 0.0001, 0.001, 0.01, and so on) from a genome-wide scan for the disease of interest. In the target samples, genetic risk scores are calculated for each individual for each set of SNPs. The selected SNPs will contain false positives but if they are enriched with true-associated variants with low effect size then the genetic risk score might still be significantly associated with the disease in an independent sample. The problem is to distinguish truly associated SNPs from the false positives, which occur massively around liberal P-value thresholds. In the schizophrenia study, the genetic risk scores based on the multiple SNPs in the discovery sample were associated with schizophrenia in three independent samples. The variance explained by the risk scores increased as more SNPs were included, that is, risk scores based on SNPs that had a P-value below 0.5 in the discovery sample explained more variance in the replication sample than the risk scores based on SNPs selected at P-value below 0.1.13 Moreover, the genetic risk scores for schizophrenia were also significantly associated with bipolar disorder assessed in two samples suggesting that the genes influencing schizophrenia and bipolar disorder partly overlap.
This study applied the risk score approach to investigate whether a polygenic component can be detected for depression and whether this polygenic component also influences anxiety. As the study samples differ in age, differences in the effect of the polygenic component may indicate that the genetic factors influencing depression and/or anxiety differ across the lifespan. Twin studies have already shown that the relative influence of genetic factors for depression decreases with age,14, 15, 16 but that the genes influencing depression remain the same across the years.17 This can be investigated directly in this study.
The discovery set consisted of the GAIN–MDD sample, including 1738 cases and 1802 controls7, 18 with over 400 000 SNPs genotyped. The target sets consisted of two independent Dutch samples. The first sample was based on the Rotterdam Study cohort and consisted of 178 depressive disorder patients diagnosed according to Diagnostic and Statistical Manual of Mental Disorders, 4th edition (DSM-IV) and 915 controls at low liability of depression as well as 222 cases for anxiety and 290 controls at low liability for anxiety. The second target sample was the Erasmus Rucphen Family (ERF) study in which symptoms of anxiety and depression were measured in 1886 participants. The subjects in the GAIN–MDD sample and the ERF sample were around 45 years of age. The Rotterdam sample was an elderly sample with a mean age of around 70. Height and intraocular pressure (IOP), phenotypes unrelated to psychiatric disorders, were additionally analyzed to examine if the association with the genetic risk scores was specific to depression and anxiety.
Materials and methods
Discovery sample
The discovery sample consisted of subjects from two large-scale longitudinal studies: the Netherlands Study of Depression and Anxiety19 and the Netherlands Twin Registry.20 The chances of success of genetic risk score analyses depend primarily on the size of the discovery or training set. If the sample size is too small, the risk profiles will be based on random noise and are not expected to explain variance in the target set. To increase the chances of success, the power of the discovery set should therefore be maximized.21 The size of the GAIN–MDD study made it more suitable to be used as the discovery set than the Rotterdam and ERF studies, which thus supplied the target samples.
The Netherlands Study of Depression and Anxiety and the Netherlands Twin Registry studies were approved by the medical ethics committee of all participating institutes. Collection of the phenotype data and quality control of the genotype data as well as the statistical methods are described in detail elsewhere.7, 18 In brief, inclusion criteria for MDD cases were a lifetime diagnosis of DSM-IV MDD22 assessed with the Composite International Diagnostic Interview,23 age of 18–65 years and self-reported western European ancestry. Persons who were not fluent in Dutch and those with a primary diagnosis of schizophrenia or schizoaffective disorders, obsessive-compulsive disorder, bipolar disorder or severe substance use dependence were excluded. Inclusion criteria for control subjects were availability of biological samples and survey data with assessments of depression, anxiety and neuroticism, no report of MDD at any measurement occasion and low genetic liability for MDD based on the survey data. In addition, controls and their parents were required to have been born in the Netherlands or Western Europe. Only one control per family was selected. The cases and controls were carefully matched on age and sex.
The genotypic data used in the discovery sample were part of one of the six initial GAIN studies sponsored by the Foundation for the National Institutes of Health (NIH).20 Individual genotyping was conducted by Perlegen Sciences (Mountain View, CA, USA) using a set of four proprietary, high-density oligonucleotide arrays. The SNPs on these arrays were selected to tag common variation in the HapMap European and Asian panels. Of the 3820 samples sent to Perlegen, genotypes were delivered for 3761 samples (98.5%) of which 3540 subjects passed quality controls and were available in the final analysis data set including 1738 MDD cases and 1802 controls. The SNP quality control process and the precautions against population stratification are detailed in Sullivan et al.7 A total of 427 037 SNPs on chromosome 1 to chromosome 22 met all selection criteria and were included in the final association analyses, which were performed in PLINK.24
Target samples
Rotterdam study
The Rotterdam Study is a prospective cohort study in the district Ommoord of Rotterdam.25 In 1990, all inhabitants aged 55 years and over were invited and 7983 persons agreed to participate. The medical ethics committee of the Erasmus Medical Center, Rotterdam, approved the study. Written informed consent was obtained from all participants.
Ascertainment of depressive symptoms and incident depressive disorders was described previously.26 Depression and anxiety symptoms were assessed with the Center for Epidemiological Studies Depression Scale (CES-D) and Hospital Anxiety and Depression Scale (HADS). The CES-D scale consists of 20 items with scores ranging from 0 to 60. A score of 16 or higher on the CES-D is considered indicative of a depressive disorder. The HADS-Depression (HADS-D) and HADS-Anxiety (HADS-A) scales each consist of seven items with scores ranging from 0 to 21 with higher scores indicating more symptoms of depression. These questionnaires are valid and reliable self report measures of symptoms of depression.27, 28 The HADS was assessed during the second visit in a randomly selected subgroup of individuals (N=2231). Depression was measured with the CES-D three times during the follow-up.
Among 7983 subjects who agreed to participate, 5974 were successfully genotyped, 524 died before depression screening and 747 did not participate in depression screening. In the remaining sample, 587 persons scoring higher than 16 on the CES-D in the third or fourth visit were invited for a semi-structured interview with the Present State Examination29 by a clinician. In addition, general practitioner records and specialist letters were surveyed actively for the occurrence of depression. Furthermore, physicians conducted repeated interviews to assess self-reported depression in the interval period. This effort identified 178 persons with current DSM-IV defined depressive disorder (145 MDD, 15 dysthymia and 18 depression-not otherwise specified cases) and eligible genotype data. The control group consisted of 915 persons, who scored in the lowest quartile (CES-D=‘0’) on CES-D in the third visit (N=3879) and who did not report any depressive symptoms during the follow up.
Anxiety disorders were assessed during the fourth visit in the total sample by trained lay interviewers who conducted a slightly adapted version of the Munich Composite International Diagnostic Interview to obtain DSM-IV diagnoses of generalized anxiety disorder, panic disorder, agoraphobia, social phobia and specific phobia. Obsessive compulsive disorder and post-traumatic stress disorder were not included. The current sample is selected from the 2779 persons who had valid Munich Composite International Diagnostic Interview assessment and genotype data. Out of 2779, 222 persons were anxiety disorder cases. The control group consists of 290 persons who did not have any anxiety disorder and scored in the lowest quartile (HADS-A=0) on the HADS-A measured in 1322 persons of the interviewed and sample with eligible genotype data during the second visit.
Genome-wide SNP data were available from the Illumina HumanHap550K (Illumina, Inc., San Diego, CA, USA) array for all cases and controls. Data were excluded based on call rate <97.5%, sex mismatch, excess autosomal heterozygosity and outliers identified by the clustering analysis. MACH 1.0 software (v1.0.16)30, 31 was used to impute to ∼2.54 million SNPs based on the HapMap CEU phased haplotypes (release 22). SNPs included in imputation met thresholds of minor allele frequency (MAF) ⩾1%, Hardy-Weinberg equilibrium (HWE) P-value ⩾10−6 and SNP call rate ⩾98.0%. GWA analysis of MDD was performed with Mach2Dat (logistic regression on allele dosage) using the GRIMP interface.30, 32 Age and sex were included as covariates in the analysis.
ERF study
The ERF study is part of the Genetic Research in Isolated Population program. The study population essentially consists of one extended family of descendents from 20 related couples who lived in the isolate between 1850 and 1900 and had at least six children baptized in the community church. The detailed information regarding ERF isolate can be found elsewhere.33, 34, 35 The medical ethical committee of the Erasmus Medical Center, Rotterdam approved the study and informed consent was obtained from all participants.
Symptoms of depression and anxiety were assessed using the HADS and the CES-D27, 28 in 2385 participants who also underwent an extensive medical examination.
Data on height and IOP were collected during the medical examination. The height of participants was determined using a stadiometer and bilateral IOP measurements were performed using Goldmann applanation tonometry.36
Among 2385 persons with phenotypes, high-density genotype data were available for 1886 subjects. The genotype data were available for this population on four different genotyping platforms which were Illumina 6K, Illumina 318K, Illumina 370K and Affymetrix 250K (Affymetrix, Inc., Santa Clara, CA, USA), which were merged and ∼2.54 × 106 SNPs were imputed using MACH 1.0 software (v1.0.16),30, 31 using build 36 HapMap (release 22) CEU population as reference. Within each genotyping batch, only SNPs showing a call rate >98%, MAF>1% and HWE P-value >10−6 were used for imputations.
As the ERF study included related individuals, GWA analyses were performed using a mixed model by ‘mmscore’ option in GenABEL software,37 which combines the Family Based Score Test for Association (FASTA) method of Abecasis et al.,38 and kinship matrix estimated from genotyped SNPs.39
Risk score profiling
The score profiling method tests the association of a genetic score variable that reflects a combined effect of a number of selected SNPs with a trait. For a more detailed description of the method, we refer to Purcell et al.13 In brief, SNPs were selected using the results from the GAIN-MDD GWA studies7 (the ‘discovery sample’). These sets of SNPs were used to calculate the genetic scores in the target samples. SNPs were selected on the basis of their nominal P-value (Pdiscovery) for association with MDD in the discovery sample. Genetic risk scores were calculated for Pdiscovery thresholds ranging from 0.00001 to 1.0.
Only those SNPs were included that were directly genotyped in the discovery set (N=427 049 SNPs). To avoid ambiguity A/T −G/C SNPs were excluded. As an A to T or G to C change will result in the same nucleotides on the opposite strands, this change might be missed during the genotype analysis. SNPs for which the quality of imputation had an R2<0.95 in target samples were also excluded. After all quality checks and exclusions, a total of 181 582 SNPs that were available in both ERF and Rotterdam study samples were selected for calculations of genetic risk scores.
For each individual in the two independent target samples, the genetic score was calculated by multiplying the number of risk alleles per SNP (0, 1 or 2) with the log odds ratio, summed over all SNPs in the considered set of SNPs.40 We calculated individual scores for each set of SNPs using the PLINK (v1.06) software.24
Logistic regression models were used to test the association of the individual genetic risk scores for depressive and anxiety disorders. Linear regression models were used to test the association between genetic risk scores and the total CES-D, HADS-D and HADS-A scores as well as for height and IOP. Sex and age were used as covariates. As an alternative control for false positivity, 10% of the non-associated cluster of SNPs with Pdiscovery>0.9 (N=1569 SNPs) in the discovery set was selected and used for computing the risk profile in both target samples.
For the Rotterdam Study regression analysis were performed in SPSS 15.0 for Windows (SPSS, Chicago, IL, USA). As the ERF sample includes relatives, data are not independent, which can lead to biased estimates of standard errors and test statistics if this dependency between measures is not taken into account.41 Association analysis of genetic risk score and the traits in ERF population were performed in Sequential Oligogenic Linkage Analysis Routines 4.1.5 software package (Southwest Foundation for Biomedical Research, San Antonio, TX, USA)42 using the ‘polygenic’ option to adjust for pedigree kinship. Among 1886 people both genotyped and phenotyped, 1697 were clustered into pedigrees (using the pedigree splitting algorithm PedCut43) and included in the family-based analysis. The remaining persons were not included in the analysis because they were also (distantly) related. The difference of the explained variance in the null and alternative model was considered as the variance explained by the genetic score. A genetic risk score with a P-value <0.05 in the model was considered as significantly associated with the trait.
Results
Table 1 shows the descriptive data for the case–control studies of GAIN–MDD and the Rotterdam Study, and Table 2 for the ERF study. As in the Rotterdam study subjects were ascertained on the basis of age 55 or more, the mean age was 74 years. This was higher than in the GAIN–MDD and ERF study in which the mean ages were around 45 years. Level of education was higher in the GAIN–MDD sample than in the other two samples. In the target samples, subjects diagnosed with a depressive disorder or an anxiety disorder were more often women and were older. In the discovery sample, cases and controls were matched based on age and sex.
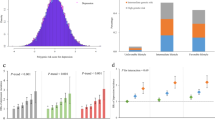
In the ERF study, CES-D, HADS-D and HADS-A scores were highly correlated (r∼0.7 pair wise for all three). Figure 1a shows the variance explained by the genetic risk scores in the logistic regression analyses performed in the Rotterdam Study using depressive disorder as dependent variable. The genetic score based on the first cluster of six SNPs (Pdiscovery <0.00001) significantly explained 0.66% of the variance in depressive disorder in the Rotterdam Study (P=0.03). This association is explained in large part by a cluster of three SNPs (rs2715148, rs2522833 and rs2522840) in the PCLO gene as after removing the PCLO SNPs in linkage disequilibrium (LD), the risk score was not significantly associated with depression in the target sample anymore. More importantly, the scores based on SNPs with Pdiscovery <0.1 to Pdiscovery<0.4 were associated with depressive disorder in the Rotterdam Study explaining up to 0.65% of the variance, with a P<0.05. As shown in Figure 1b, the Rotterdam Study anxiety disorder case–control sample analysis yielded the highest percentage of variance explained with the genetic risk scores from GAIN–MDD study. The risk scores based on SNPs with Pdiscovery<0.1 to Pdiscovery<1.0 significantly explained up to 2.1% of the variance (P=0.0025). For Pdiscovery values of 0.1, 0.2 and 0.3, the percentage of variance increased from 1 to 2% when a higher number of SNPs were included in risk scoring.
Percentage of variance explained by genetic risk scores in Rotterdam Study. Percentage of variance represented as difference in Nagelkerke R2 after adjustment for age and sex. (a) Analyses based on comparison of DD persons (n=178) to persons scoring in the lowest quartile of CES-D (CES-D=‘0’) scale and who did not report any depressive complaints during the follow-up (n=915). (b) Analyses based on comparison of persons with anxiety disorder (n=222) to persons scoring in the lowest quartile of HADS-A (HADS-A=‘0’) scale and who did not report any depression or anxiety symptoms during the follow-up (n=290). *P value <0.05, **P value <0.001.
Figures 2a–c show the linear regression results for the analysis using the continuous scores on the CES-D, HADS-D and HADS-A in the ERF study. For CES-D, the scores based on SNPs with Pdiscovery values <0.01 and 0.1 explained ∼0.5% of the variance (P=0.007 and P=0.008). For the HADS-D, the score based on SNPs with Pdiscovery<0.01 significantly explained 0.3% of the variance (P=0.03). The MDD-based genetic score was also significantly associated with anxiety measured with the HADS-A explaining up to 0.5% of the variance (P=0.01).
Percentage of variance explained in depression and anxiety symptoms in ERF study by the genetic risk scores. Percentage of variance represented as difference in R2 after adjusting for age sex and family relations. (a–c), analyses of continuous scales. *P value <0.05, **P value <0.01.
To examine whether these results were due to chance, we tested whether the MDD-based genetic risk score predict also variation in height and IOP measured in ERF. Heritability of IOP was 35% and 86% for height36, 44 and none of the traits was correlated to depression or anxiety in the ERF study. The genetic risk score for MDD failed to predict IOP and height (Figures 3a and b) suggesting that this relation is specific to depression and anxiety. Moreover, a genetic score of SNPs with Pdiscovery<0.9 in the discovery set did not show significant association with any of the phenotypes in the target samples (data not shown).
Predicting height (a) and IOP (b) in ERF study. Linear regression analysis of height and intraocular pressure. Percentage of variance represented as difference in R2 after adjusting for age sex and family relations.
Discussion
The aim of this study was to investigate the genetic architecture of depression and the potential overlap in genetic risk factors with anxiety. Owing to the availability of an elderly cohort, it was also possible to examine whether the genetic factors influencing anxiety and depression change across the life span. Using genetic risk scores derived from the association results of the GAIN–MDD study in two independent target samples, we evaluated the evidence for a genome-wide signature for several measures of depression and anxiety used as outcome variables in the target samples. For depression, either diagnosed according to the DSM-IV or measured with the CES-D or HADS-D, we could explain up to ∼1% of the variance with the genetic risk scoring approach. Moreover, the genetic risk scores for depression were also associated with anxiety explaining up to 2.1% of the variance when approximately half of the genome-wide SNP data were included in the score. The explained variance was highest in the elderly sample indicating that the genetic factors influencing anxiety and depression hardly change with age. No significant results were found for the control variables height and IOP, implying that the association of the genetic risk score with depression and anxiety does not reflect chance alone. Overall, these findings suggest the presence of many loci, each with a small effect influencing depression as well as anxiety.
We checked whether our results were only because of SNPs in high LD segregating together. We performed a strict LD pruning (200 SNPs sliding window with r2snp−snp threshold of 0.25). Considering the CES-D scale in ERF study, percentage of variance explained by the risk scores based on SNPs with Pdiscovery<0.01 dropped slightly after LD pruning from 0.52 to 0.49 but remained significant (P=0.01) whereas a less strict pruning with an r2snp−snp threshold of 0.50 improved the percentage of explained variance to 0.62 (P=0.0003). Results with HADS-A and HADS-D scales were similar which shows that LD pruning itself does not add a major difference to the method (data not shown). Excluding the SNPs with minor allele frequency <0.05 did not change the explained variance. This finding suggest the common disease common variant hypothesis is explaining MDD heritability, on the other hand, the power to detect the effect of rare variants in the discovery and target sets, was low and such rare variants may be detected by other approaches such as linkage or deep sequencing.
Our results are in agreement with the results from the International Schizophrenia Consortium13 that pointed out a polygenic component influencing schizophrenia as well as bipolar disorder. There was a somewhat higher amount of explained variance for schizophrenia (3.2% compared with 1%). This may be due to power issues such as differences in sample size (∼3300 cases for International Schizophrenia Consortium vs ∼1800 cases for GAIN–MDD), MDD being a common disease with clear non-genetic influence because of life events, and lower heritability compared with schizophrenia (∼40 vs ∼80%).
The percentage of explained variance in anxiety (2.1%) supports the idea of shared genetic background between these disorders. This has already been suggested by twin studies45 and is confirmed by our results. The trend of an increase in R2 for anxiety with different Pdiscovery thresholds is different from the trend that we observe in depression, pointing out that the effect sizes are different, but the direction of effect is the same. We would like to stress that difference in explained variance between the target samples can evenly well be explained by chance. Moreover, it is important to note that 70% of the GAIN–MDD cases had a comorbid lifetime anxiety diagnosis. However, this high comorbidity is exactly what is expected if two disorders are influenced by similar genes and diagnoses are not mutually exclusive. Future research, preferably with a more balanced proportion of pure depressed and comorbid cases, can shed more light on the overlap in genetic factors influencing anxiety and depression.
A limitation of this study was that there were some differences between the discovery set and the target samples. Different instruments were used to measure depression and anxiety. In the GAIN–MDD study, the Composite International Diagnostic Interview was used to diagnose MDD and anxiety disorders, while in the Rotterdam study, the Present State Examination (PSE) was used. However, both instruments aim to make diagnoses according to the DSM-IV criteria and have adequate agreement for overall syndromes.46 In the ERF population, symptoms of depression and anxiety were measured using the CES-D and HADS. Several validation studies on various types of patients using different diagnostic tools have shown that HADS performs well in assessing the symptom severity and case status of anxiety disorders and depression in both somatic, psychiatric and primary care patients and in the general population.47 The HADS-D subscale has shown high sensitivity (∼0.9) and specificity (∼0.7) for MDD as diagnosed by DSM-IV in various studies.48 The CES-D scale was found to be satisfactory in a semi-clinical sample of the elderly and in general population (sensitivity=0.9 and specificity=0.6) for life-time MDD and also performed excellent for 1 month of prevalence of MDD as diagnosed by DSM-IV (sensitivity=1.0 and specificity=0.9) among elderly Dutch.49, 50, 51 Considering the HADS-A subscale, the sensitivity and specificity for DSM-IV generalized anxiety disorder was reported to be 0.9 and 0.8, respectively.52 In addition, the discovery set in this study included lifetime MDD cases, whereas the Rotterdam study recorded depressive disorders during a 9-year follow-up rather than lifetime MDD. Similarly, CES-D and HADS measure depressive and anxious symptoms in the last week. This means that subjects in the control groups in the target samples may be non-current but life-time MDD or anxiety cases. To summarize, although the measurements of anxiety and depression used in the three study samples are definitely related to each other, the fact that they are not entirely similar implies some heterogeneity, biasing the results toward the null hypothesis.
Another point involves the difference in gender ratios between discovery and target samples. In the discovery sample, the cases and controls were carefully matched on age and sex. Meta-analysis of twin studies suggests that genetic factors that influence depression are mostly shared between men and women.1, 53, 54 Sex was also used as a covariate when predicting depression or anxiety in the target samples. Thus, it seems unlikely that the gender ratio may have a major effect in the replication of the findings.
There was also heterogeneity in education level as a measure of socioeconomical status. In spite of these differences, we still found a significant effect of the genetic risk score suggesting that the effects of the risk scores are actually even stronger. In both the International Schizophrenia Consortium study and the current studies, the low variance explained compared with the heritability of the disorders will also reflect that the analyses did not include the X chromosome, that gene–gene or gene–environment interactions are not considered and that the current generation of genotyping platforms do not fully tag genomic variance.55
This study is the second study showing direct evidence for a polygenic component influencing the susceptibility for a psychiatric disorder as well as overlap in genetic risk factors with another psychiatric condition. In addition, this study suggests that the genetic factors influencing anxiety and depression hardly change with age. The results imply that causal SNPs or the SNPs in LD with such variants do exist, but have lower effect sizes than the first generation of GWA studies on psychiatric disorders was powered to detect. This provides optimism that variants associated at genome-wide levels of significance will be detectable as sample sizes increase in the next generation of GWA studies and their meta/mega-analyses. Moreover, it confirms that genome-wide profiling is a useful approach to analyze the genetic architecture of disorders, that is, similarities and differences in genetic factors influencing several disorders or influencing the same disorder across the lifespan.
Accession codes
References
Sullivan PF, Neale MC, Kendler KS . Genetic epidemiology of major depression: review and meta-analysis. Am J Psychiatry 2000; 157: 1552–1562.
Middeldorp C, Boomsma D . Genetics and Psychopathology. In: Berntson GG CJ (ed). Handbook of Neuroscience for Behavioral Sciences. Wiley: New York, 2009.
Levinson DF . The genetics of depression: a review. Biol Psychiatry 2006; 60: 84–92.
Lopez-Leon S, Janssens AC, Gonzalez-Zuloeta Ladd AM, Del-Favero J, Claes SJ, Oostra BA et al. Meta-analyses of genetic studies on major depressive disorder. Mol Psychiatry 2008; 13: 772–785.
Kessler RC, Berglund P, Demler O, Jin R, Koretz D, Merikangas KR et al. The epidemiology of major depressive disorder: results from the National Comorbidity Survey Replication (NCS-R). JAMA 2003; 289: 3095–3105.
Middeldorp CM, Cath DC, Van Dyck R, Boomsma DI . The co-morbidity of anxiety and depression in the perspective of genetic epidemiology. A review of twin and family studies. Psychol Med 2005; 35: 611–624.
Sullivan PF, de Geus EJ, Willemsen G, James MR, Smit JH, Zandbelt T et al. Genome-wide association for major depressive disorder: a possible role for the presynaptic protein piccolo. Mol Psychiatry 2009; 14: 359–375.
Bochdanovits Z, Verhage M, Smit AB, de Geus EJ, Posthuma D, Boomsma DI et al. Joint reanalysis of 29 correlated SNPs supports the role of PCLO/Piccolo as a causal risk factor for major depressive disorder. Mol Psychiatry 2009; 14: 650–652.
Hek K, Mulder CL, Luijendijk HJ, van Duijn CM, Hofman A, Uitterlinden AG et al. The PCLO gene and depressive disorders: replication in a population-based study. Hum Mol Genet 2010; 19: 731–734.
Psychiatric GCCC, Cichon S, Craddock N, Daly M, Faraone SV, Gejman PV et al. Genomewide association studies: history, rationale, and prospects for psychiatric disorders. Am J Psychiatry 2009; 166: 540–556.
Muglia P, Tozzi F, Galwey NW, Francks C, Upmanyu R, Kong XQ et al. Genome-wide association study of recurrent major depressive disorder in two European case-control cohorts. Mol Psychiatry 2010; 15: 589–601.
Craddock N, O’Donovan MC, Owen MJ . Genome-wide association studies in psychiatry: lessons from early studies of non-psychiatric and psychiatric phenotypes. Mol Psychiatry 2008; 13: 649–653.
International Schizophrenia Consortium Purcell SM Wray NR Stone JL Visscher PM, O’Donovan MC et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 2009; 460: 748–752.
Baldwin RC, Tomenson B . Depression in later life. A comparison of symptoms and risk factors in early and late onset cases. Br J Psychiatry 1995; 167: 649–652.
Jansson M, Gatz M, Berg S, Johansson B, Malmberg B, McClearn GE et al. Gender differences in heritability of depressive symptoms in the elderly. Psychol Med 2004; 34: 471–479.
Johnson W, McGue M, Gaist D, Vaupel JW, Christensen K . Frequency and heritability of depression symptomatology in the second half of life: evidence from Danish twins over 45. Psychol Med 2002; 32: 1175–1185.
Gillespie NA, Kirk KM, Evans DM, Heath AC, Hickie IB, Martin NG . Do the genetic or environmental determinants of anxiety and depression change with age? A longitudinal study of Australian twins. Twin Res 2004; 7: 39–53.
Boomsma DI, Willemsen G, Sullivan PF, Heutink P, Meijer P, Sondervan D et al. Genome-wide association of major depression: description of samples for the GAIN Major Depressive Disorder Study: NTR and NESDA biobank projects. Eur J Hum Genet 2008; 16: 335–342.
Penninx BW, Beekman AT, Smit JH, Zitman FG, Nolen WA, Spinhoven P et al. The Netherlands Study of Depression and Anxiety (NESDA): rationale, objectives and methods. Int J Methods Psychiatr Res 2008; 17: 121–140.
Boomsma DI, de Geus EJ, Vink JM, Stubbe JH, Distel MA, Hottenga JJ et al. Netherlands Twin Register: from twins to twin families. Twin Res Hum Genet 2006; 9: 849–857.
Wray NR, Goddard ME, Visscher PM . Prediction of individual genetic risk to disease from genome-wide association studies. Genome Res 2007; 17: 1520–1528.
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders 4th edn. American Psychiatric Association, Washington, DC, 1994.
World Health Organization. Composite International Diagnostic Interview (version 2.1). WHO: Geneva, 1992.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007; 81: 559–575.
Hofman A, Breteler MM, van Duijn CM, Janssen HL, Krestin GP, Kuipers EJ et al. The Rotterdam Study: 2010 objectives and design update. Eur J Epidemiol 2009.
Luijendijk HJ, van den Berg JF, Dekker MJ, van Tuijl HR, Otte W, Smit F et al. Incidence and recurrence of late-life depression. Arch Gen Psychiatry 2008; 65: 1394–1401.
Radloff . The CES-D scale: a self report depression scale for research in theb general population. Appl Pshycol Measurement 1977; 3: 385–401.
Zigmond AS, Snaith RP . The hospital anxiety and depression scale. Acta Psychiatr Scand 1983; 67: 361–370.
Wing J, Cooper JE, Sartorius N . 1974.
Li Y, Abecasis GR . Mach 1.0: rapid haplotype reconstruction and missing genotype inference. Am J Hum Genet S79–2290, 2006.
Nothnagel M, Ellinghaus D, Schreiber S, Krawczak M, Franke A . A comprehensive evaluation of SNP genotype imputation. Hum Genet 2009; 125: 163–171.
Estrada K, Abuseiris A, Grosveld FG, Uitterlinden AG, Knoch TA, Rivadeneira F . GRIMP: a web- and grid-based tool for high-speed analysis of large-scale genome-wide association using imputed data. Bioinformatics 2009; 25: 2750–2752.
Lopez-Leon S, Chi Choy W, Aulchenko YS, Claes SJ, Oostra BA, Mackenbach JP et al. Genetic factors influence the clustering of depression among individuals with lower socioeconomic status. PLoS ONE 2009; 4: e5069.
Santos R, Zillikens M, Rivadeneira F, Pols H, Oostra B, van Duijn C et al. Heritability of fasting glucose levels in a young genetically isolated population. Diabetologia 2006; 49: 667–672.
Aulchenko YS, Heutink P, Mackay I, Bertoli-Avella AM, Pullen J, Vaessen N et al. Linkage disequilibrium in young genetically isolated Dutch population. Eur J Hum Genet 2004; 12: 527–534.
van Koolwijk LME, Despriet DDG, van Duijn CM, Pardo Cortes LM, Vingerling JR, Aulchenko YS et al. Genetic contributions to glaucoma: heritability of intraocular pressure, retinal nerve fiber layer thickness, and optic disc morphology. Invest Ophthalmol Vis Sci 2007; 48: 3669–3676.
Aulchenko YS, Ripke S, Isaacs A, van Duijn CM . GenABEL: an R library for genome-wide association analysis. Bioinformatics 2007; 23: 1294–1296.
Abecasis GR, Cherny SS, Cookson WO, Cardon LR . Merlin--rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet 2002; 30: 97–101.
Amin N, van Duijn CM, Aulchenko YS . A genomic background based method for association analysis in related individuals. PLoS One 2007; 2: e1274.
Evans DM, Visscher PM, Wray NR . Harnessing the information contained within genome-wide association studies to improve individual prediction of complex disease risk. Hum Mol Genet 2009; 18: 3525–3531.
Rebollo I, de Moor MH, Dolan CV, Boomsma DI . Phenotypic factor analysis of family data: correction of the bias due to dependency. Twin Res Hum Genet 2006; 9: 367–376.
Almasy L, Blangero J . Multipoint quantitative-trait linkage analysis in general pedigrees. Am J Hum Genet 1998; 62: 1198–1211.
Liu F, Kirichenko A, Axenovich TI, van Duijn CM, Aulchenko YS . An approach for cutting large and complex pedigrees for linkage analysis. Eur J Hum Genet 2008; 16: 854–860.
Axenovich TI, Zorkoltseva IV, Belonogova NM, Struchalin MV, Kirichenko AV, Kayser M et al. Linkage analysis of adult height in a large pedigree from a Dutch genetically isolated population. Hum Genet 2009; 126: 457–471.
Kendler KS, Prescott CA, Myers J, Neale MC . The structure of genetic and environmental risk factors for common psychiatric and substance use disorders in men and women. Arch Gen Psychiatry 2003; 60: 929–937.
Andrews G, Peters L . The psychometric properties of the composite international diagnostic interview. Soc Psychiatry Psychiatr Epidemiol 1998; 33: 80–88.
Bjelland I, Dahl AA, Haug TT, Neckelmann D . The validity of the hospital anxiety and depression scale. An updated literature review. J Psychosom Res 2002; 52: 69–77.
Silverstone PH . Concise assessment for depression (CAD): a brief screening approach to depression in the medically ill. J Psychosom Res 1996; 41: 161–170.
Beekman AT, Deeg DJ, Van Limbeek J, Braam AW, De Vries MZ, Van Tilburg W . Criterion validity of the Center for Epidemiologic Studies Depression scale (CES-D): results from a community-based sample of older subjects in The Netherlands. Psychol Med 1997; 27: 231–235.
Donker T, van Straten A, Marks I, Cuijpers P . Brief self-rated screening for depression on the Internet. J Affect Disord 2010; 122: 253–259.
Haringsma R, Engels GI, Beekman AT, Spinhoven P . The criterion validity of the Center for Epidemiological Studies Depression Scale (CES-D) in a sample of self-referred elders with depressive symptomatology. Int J Geriatr Psychiatry 2004; 19: 558–563.
Olsson I, Mykletun A, Dahl AA . The Hospital Anxiety and Depression Rating Scale: a cross-sectional study of psychometrics and case finding abilities in general practice. BMC Psychiatry 2005; 5: 46.
Kendler KS, Gatz M, Gardner CO, Pedersen NL . A Swedish national twin study of lifetime major depression. Am J Psychiatry 2006; 163: 109–114.
Middeldorp CM, Birley AJ, Cath DC, Gillespie NA, Willemsen G, Statham DJ et al. Familial clustering of major depression and anxiety disorders in Australian and Dutch twins and siblings. Twin Res Hum Genet 2005; 8: 609–615.
Ng SB, Turner EH, Robertson PD, Flygare SD, Bigham AW, Lee C et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature 2009; 461: 272–276.
Acknowledgements
GAIN-MDD: We acknowledge support from NWO: genetic basis of anxiety and depression (904-61-090); resolving cause and effect in the association between exercise and well-being (904-61-193); twin-family database for behavior genomic studies (480-04-004); twin research focusing on behavior (400-05-717), Center for Medical Systems Biology (NWO Genomics); Spinozapremie (SPI 56-464-14192); Neuroscience Campus Amsterdam (NCA-VU); genome-wide analyses of European twin and population cohorts (EU/QLRT-2001-01254); genome scan for neuroticism (NIMH R01 MH059160); Geestkracht program of ZonMW (10-000-1002); matching funds from universities and mental health care institutes involved in NESDA. Genotyping was funded by the Genetic Association Information Network (GAIN) of the Foundation for the US National Institutes of Health, and analysis was supported by grants from GAIN and the NIMH (MH081802). CM Middeldorp is financially supported by NWO (VENI grant 916-76-125). Genotype data were obtained from dbGaP (http://www.ncbi.nlm.nih.gov/dbgap, accession number phs000020.v1.p1). Statistical analyses were carried out on the Genetic Cluster Computer (http://www.geneticcluster.org) which is financially supported by the NWO (480-05-003). Dr Sullivan was also supported by R01s MH074027 and MH077139. Dr Wray is supported by the Australian National Health and Medical Research Council (grant 496688). Dr Middeldorp was supported by NWO (VENI grant 916-76-125). Rotterdam Study: Genome-wide genotyping of the Rotterdam Study was supported by NWO (175.010.2005.011). ERF study: The ERF study was supported by grants from the Netherlands Organisation for Scientific Research, Erasmus MC, The Netherlands Brain Foundation (HsN) and the Centre for Medical Systems Biology (CMSB). The genotyping was founded by EUROSPAN (European Special Populations Network) Consotium. We are grateful to all study participants and their relatives, general practitioners and neurologists for their contributions and to P Veraart for her help in genealogy, J Vergeer for the supervision of the laboratory work and P Snijders for his help in data collection.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Demirkan, A., Penninx, B., Hek, K. et al. Genetic risk profiles for depression and anxiety in adult and elderly cohorts. Mol Psychiatry 16, 773–783 (2011). https://doi.org/10.1038/mp.2010.65
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2010.65
Keywords
This article is cited by
-
A promising microbial α-amylase production, and purification from Bacillus cereus and its assessment as antibiofilm agent against Pseudomonas aeruginosa pathogen
Microbial Cell Factories (2023)
-
Association of time spent in outdoor light and genetic risk with the incidence of depression
Translational Psychiatry (2023)
-
Protocol for a systematic review and network meta-analysis of randomised controlled trials examining the effectiveness of early parenting interventions in preventing internalising problems in children and adolescents
Systematic Reviews (2020)
-
Machine learning for effectively avoiding overfitting is a crucial strategy for the genetic prediction of polygenic psychiatric phenotypes
Translational Psychiatry (2020)
-
Genetic susceptibility, inflammation and specific types of depressive symptoms: evidence from the English Longitudinal Study of Ageing
Translational Psychiatry (2020)