Abstract
Metastases, or migration of cancers, are common and severe cancer complications. Although the 5-year survival rates of primary tumors have greatly improved, those of metastasis remain below 30%, highlighting the importance of investigating specific mechanisms and therapeutic approaches for metastasis. Microfluidic devices have emerged as a powerful platform for drug target identification and drug response screening and allow incorporation of complex interactions in the metastatic microenvironment as well as manipulation of individual factors. In this work, we review microfluidic devices that have been developed to study cancer cell migration and extravasation in response to mechanical (section ‘Microfluidic investigation of mechanical factors in cancer cell migration’), biochemical (section ‘Microfluidic investigation of biochemical signals in cancer cell invasion’), and cellular (section ‘Microfluidic metastasis-on-a-chip models for investigation of cancer extravasation’) signals. We highlight the device characteristics, discuss the discoveries enabled by these devices, and offer perspectives on future directions for microfluidic investigations of cancer metastasis, with the ultimate aim of identifying the essential factors for a ‘metastasis-on-a-chip’ platform to pursue more efficacious treatment approaches for cancer metastasis.
Similar content being viewed by others
Introduction
Metastasis is the migration of cancers, which occurs when cancer cells break off from the primary tumor, travel in the lymph or blood vessels, and eventually extravasate into a tissue and establish a secondary tumor1. Although only 0.001–0.02% of cancer cells that enter circulation can form a metastatic tumor2, successful metastasis significantly increases the morbidity and mortality of patients. Metastases are difficult to treat because they signal a systemic disease that can affect multiple organs. Although the 5-year survival rates for localized cancers are relatively high, those for metastases are <30% and have not improved significantly over the last 10 years3, making metastasis the major cause of death from cancer, as reported by the World Health Organization4.
Significant efforts have been spent on investigating cancer metastases, as summarized in several recent review articles5–9. Many of these studies focused on the causes of metastatic specificity. For example, breast cancer mainly metastasizes to bone, lung, brain, and liver tissues, whereas prostate cancer primarily metastasizes to bone10. This tendency suggests that cancer cells receive and respond to signals from the secondary site, leading to preferential migration. These signals might take the form of chemokines released from the secondary tumor site, mechanical properties of the environment that guide the migration of cancer cells, or interactions with other types of cells. Metastatic cancer cells were shown to adjust their migration mechanisms in response to these signals and more robustly than non-metastatic and normal cells11. It was also found that certain cancer drugs can lead to increased metastasis12. These factors highlight the need to investigate treatments designed specifically for metastasis.
Currently, treatments for cancer and metastasis are often tested in vivo using mouse models in which cancer cells are injected orthotopically or into circulation. Cell-labeling techniques have also recently enabled the observation of cell–cell interactions13,14. These in vivo models have the advantage of strong physiological relevance compared with in vitro models. However, use of human cancer cells in mice requires immunocompromised mice, which renders the power of this approach limited because immune cells play an important role in metastasis15,16. This and other differences between mice models and humans might be the reason that many drugs tested in mice have failed to work in humans17,18. Even for the drugs that do work in patients, their effects cannot be extrapolated across all patients due to tumor heterogeneity19. These observations prompt the need for individualized medicine. A few in vitro systems have been developed to predict individualized drug response for primary tumors20, and these systems often involve isolation and culture of primary tumor cells with chemotherapy drugs. The growth of these tumor cells can be monitored as well as their gene expression and other measures. More sophisticated systems have used tumor spheroids21 and incorporated extracellular matrix (ECM) and other cell types22. However, the response of cancer cells to drugs might differ because they encounter different environments during metastasis. Therefore, systems specifically for metastasis are direly needed8.
The first conventional in vitro system used to study metastasis was the 2D wound-healing assay, which involves creation of a ‘wound’ gap in a cell monolayer and observation of cell migration to close the ‘wound’ gap. Although simple to use, this approach fails to replicate the 3D environment and the signal gradients that are present in vivo22,23. To address these limitations, the Boyden chamber assay was developed in which cells are seeded on well inserts with semi-permeable membranes, and their migration across the membrane toward a chemoattractant is quantified. The membrane can also be coated with ECM or seeded with other types of cells. However, this technique permits only end-point measurement but not real-time imaging, which could be important for observing changes in cell morphology in response to chemoattractants or drugs. Furthermore, this method does not allow for single-cell analysis and cannot reveal intratumor heterogeneity24. Microfluidics has emerged as a powerful platform for study of cancer metastases25–28 and drug screening29,30. Due to their micro-scaled structures, microfluidic devices require low numbers of cells and offer the potential for high-throughput screening. Microfluidic devices also allow for additional types of cells to co-exist in 3D while maintaining each cell population in their appropriate environment, which enables the intricate physiological environment during metastasis to be replicated.
The metastatic cascade is extremely complex. Growth of the primary tumor involves the recruitment of cells, such as cancer-associated fibroblasts and endothelial cells, for angiogenesis. This scenario creates a gradient of chemokines and ECM stiffness to guide cancer cell migration toward the circulation, where cancer cells must subsequently degrade the ECM and intravasate across endothelial cells to enter the circulation. Cancer cells that have survived circulation might reach a secondary organ, where they might adhere or be trapped physically in small vessels. In this case, the cells might extravasate out of the vessels, degrade the ECM, and grow to establish a secondary tumor. Because metastasis is a multi-step process and every step requires unique gene expression and interaction with other cells, each step can be targeted for treatment. For example, studies have investigated the potential for targeting αvβ3 and αvβ5 integrin to prevent tumor cell adhesion to the ECM31, C-X-C ligand-12 (CXCL-12; also known as SDF-1α) to reduce recruitment of myeloid bone marrow-derived cells that can support angiogenesis and tumor growth32, and receptor activator of nuclear factor kappa-B ligand to block the differentiation of osteoclasts that can facilitate bone metastasis33. Therefore, several of these steps have been investigated using microfluidic devices, as summarized in recent reviews25–28. Nevertheless, many patients already showed signs of disseminated tumor cells at the time of primary tumor diagnosis34, indicating that drugs targeting the early stages in metastasis might not be ideal. In addition, cancer cell activity prior to arrival at the vasculature of the secondary site is thought to primarily follow the blood flow35, and it is more difficult to treat metastasis once a secondary tumor is established3.
Therefore, this review is focused on cancer cell extravasation and migration at the secondary site under (a) mechanical stimulation (section ‘Microfluidic investigation of mechanical factors in cancer cell migration’), (b) biochemical factors (section ‘Microfluidic investigation of biochemical signals in cancer cell invasion’), and (c) cells of the secondary site (section ‘Microfluidic metastasis-on-a-chip models for investigation of cancer extravasation’), as summarized in Figure 1. An overview of the capabilities of current microfluidic devices could aid in identifying important factors in metastasis that might be investigated as drug targets or should be included in a ‘metastasis-on-a-chip’ device for drug screening.
Several factors that guide cancer cell invasion to secondary tissue have been investigated using microfluidics devices: Mechanical (confinement, ECM stiffness, and fluid flow) (section ‘Microfluidic investigation of mechanical factors in cancer cell migration’), biochemical factors (section ‘Microfluidic investigation of biochemical signals in cancer cell invasion’), and interaction with cells at the secondary tumor site (section ‘Microfluidic metastasis-on-a-chip models for investigation of cancer extravasation’).
Microfluidic investigation of mechanical factors in cancer cell migration
When cancer cells metastasize, they must migrate through small spaces in the ECM at the primary and secondary tumor sites (as small as 1 μm in rat collagen matrices36) as well as the capillaries or small lymphatic vessels (5–10 μm). These sites all have different confinement and stiffness characteristics, which are two mechanical cues that are known to affect cell migration37–39. The stiffness gradient created by cancer-associated fibroblasts (CAFs) had been shown to guide cancer cell migration40. Furthermore, chemotherapies that effectively increase leukemia apoptosis were suggested to increase cancer cell invasion in capillary-sized constrictions41. Therefore, microfluidic devices were developed to investigate cancer cell response to mechanical cues and drug impact on these responses (Figure 2)42–47.
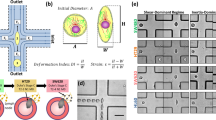
Modeling of mechanical signals received by metastasizing cancer cells. (a) Confinement channels fabricated in 96-well plates. Adapted from Ref. 42 with permission. (b) Cell migration into sub-nuclear confinement channels of different lengths. Adapted from Ref. 43 with permission. (c) Aggregates of cancer cells showing rearrangement and migration into confinement channels (time i to iv). Adapted from Ref. 44 with permission. (d) Device for cancer cell isolation after confined migration for single-cell analysis. Adapted from Ref. 45 with permission. (e) Device for analysis of cell stiffness and size during migration in confined channels. Adapted from Ref. 46 with permission. (f) Device for interstitial flow. Adapted from Ref. 47 with permission.
Irimia et al.42 developed an early microfluidic device for investigating the effect of confinement on cancer cell migration (Figure 2a). To simulate different levels of confinement that cancer cells experience in vivo in a high-throughput manner, polydimethylsiloxane (PDMS) devices consisting of four arrays of 50 channels (3–12 μm×6–100 μm) were plasma-bonded on coverslips at the bottom of 96-well plates. In these confined channels, cancer cells migrated continuously in one direction in the absence of chemical gradients while 3T3 fibroblasts frequently changed directions and paused, suggesting that confinement alone is sufficient to drive persistent migration and that cancer cells are more responsive to mechanical cues. The migration speed of MDA-MB-231 breast cancer cells was shown to be fastest in 3 μm×25 μm channels at 50 μm h−1. Application of drugs that alter microtubule dynamics (Nocodazole and Taxol) reduced this migration speed. A similar finding was observed with a polyethylene (glycol) diacrylate device constructed using digital micromirror device-based projection printing48 that has the advantage of being able to print exact replicas of capillaries. However, due to the complexity of the printed capillaries, the researchers made a simplified honeycomb model (25–120 μm) instead and observed that HeLa cervical cancer cells migrated faster in narrower channels, whereas the migration speed of non-cancerous 10T1/2 cells did not change with the channel width.
To more closely recapitulate the gradual change in confinement levels that cancer cells experience as they move into smaller vessels, a microfluidic device was developed to observe the effect of tapering angles on cancer cell migration49. This device consists of an array of PDMS microchannels with varying tapering angles (1–40°) between the large (15 μm×10 μm) and the small (4 μm×10 μm) channels. It was observed that >80% of metastatic MDA-MB-231 breast cancer cells underwent permeation (invasion into the small channel) rather than repolarization (turning back to the large channel at the tapering region) at all tapering angles. MDA-MB-231 cells were also found to be more likely to permeate at high-gradient tapering regions (>7°) than the non-cancerous MCF-10A cells.
Because the smallest space that cancer cells experience during metastasis can reach sub-nuclear levels50 and nuclear deformation might lead to downstream mechanotransduction pathways51, several devices sufficiently small to confine the cancer cell nucleus were constructed to observe nuclear deformation43,52–55. These devices demonstrated the protrusion of cancer cell cytoplasm into the sub-nuclear channels prior to nuclear deformation43,52. This approach might aid in reducing the cell size and facilitating nutrient finding. It was also observed that migration from the larger channel (15 μm×10 μm) through the sub-nuclear channel (15 μm×3.3 μm, 10 or 60 μm long) is reduced if the sub-nuclear channel is longer than a typical cell (Figure 2b)43. Furthermore, the application of 5ʹ-deoxy-5ʹ-methylthioadenosine (a methylase inhibitor that causes chromatin de-condensation)52 and Taxol (that alters microtubule dynamics)43 reduced the percentage and speed of cancer cell migration into the sub-nuclear channels, whereas the application of phospholipid sphingosylphosphorylcholine (that reduces cell stiffness)53 increased these measures. Cancer cells were shown to migrate through constrictions as narrow as 2 μm formed by PDMS pillars54, but nuclear translocation was greatly impeded when the sub-nuclear channel (2–20 μm×5 μm) was narrower than 5 μm (Ref. 55). These studies advanced the understanding of the morphological changes of cancer cells when they invade spaces with sub-nuclear confinements.
The difference in cancer cell behavior during sub-nuclear migration under drug treatments was further investigated56,57 by observing MDA-MB-231 breast cancer cell migration in channels of various sizes (3–50 μm×10 μm). The cortical F-actin and microtubules of migrating cells were shown to be uniformly distributed in 50 μm channels, yet redistributed to the cell poles with no focal adhesion in 3 μm channels. The researchers also found that the application of Y-27632 (a Rho-associated kinase inhibitor that can eliminate stress fibers and focal adhesions) in 50 μm channels induced long protrusions that could not be retracted, leading to a significant reduction in cell displacement. In contrast, the migration speed was increased in 3 μm channels. The researchers also demonstrated that ML-7 (an inhibitor of myosin light-chain kinase), CT04 (an inhibitor of Rho A, B, and C), and β1-integrin neutralizing antibody reduced migration only in 50 μm and not in 3 μm channels. Actin polymerization inhibitors cytochalasin D and latrunculin-A had no effect on migration in 3 μm channels. Blebbistatin (an inhibitor of non-muscle myosin II a, important for cell contractility) even enhanced migration speed in 3 μm channels. Finally, the application of Taxol (an inhibitor of microtubule depolymerization) and colchicine (a promoter of microtubule disassembly) significantly reduced migration in 3 μm channels.
To further investigate the response of cancer cells to different levels of confinement, Mak et al. developed a microfluidic device in which migrating cancer cells encountered a branch point leading to two channels, one 10 μm wide and the other 3.3 μm wide58. The two channels were configured in a circular (two branches splitting at the same angle) or semi-circular (smaller channel collinear with the original path leading to the branch point) pattern. Cancer cells were observed to either enter directly into one path or first extend into both channels. Approximately 90% of the cells entered the larger channel for circular branch points, whereas 68% did so for semi-circular branch points. Moreover, only 35% of cells treated with blebbistatin (inhibitor of non-muscle myosin II a) entered the larger channel at semi-circular branch points, and Taxol significantly reduced the probability of cancer cells making a path decision and migrating further.
Because clusters of cancer cells had been isolated from venous circulation59,60 and were shown to be more invasive than single cancer cells61, Au et al.44 instigated extravasation of cancer cell clusters (2–20 cells) through narrow constrictions (Figure 2c). A microfluidic device was constructed with 16 microchannels (30 μm×30 μm) tapering into constrictions of 5–10 μm. The outlet was lowered to create a hydrostatic pressure and flow. Primary cancer cell clusters and MDA-MB-231 breast cancer cells were cultured in ultralow attachment well plates. Using the microfluidic device, 90% of clusters (up to 20 cells) were found to migrate through the narrowest constriction (5 μm). As the clusters approached the constriction, they unfolded into a chain-like organization and reformed into clusters as they exited. For small clusters of <5 cells, the migration velocity of the cluster through the constriction was observed to be equal to the velocity of an individual cell of the same diameter as the largest cell in the cluster. For larger clusters, the migration velocity was more similar to the sum of resistances of the cluster’s individual cells.
Due to increased recognition of the importance of intra-tumor heterogeneity, a microfluidic device was developed to isolate and capture invasive and non-invasive subsets of tumor cells for analysis (Figure 2d)45. The device consisted of two symmetrical serpentine channels (40 μm tall) connected at each turn through a central migration channel (20 μm tall) and a ‘capture gap’ (20 μm tall and 10 μm wide). Because the serpentine channel was longer and had a higher resistance, cells loaded into one serpentine channel went down the central migration channel and were trapped in the capture gap. A chemoattractant was applied to the other serpentine channel to stimulate cancer cell migration across the central channel. Finally, cells were detached using trypsin from both sides of the central channel to isolate the migratory and non-migratory cell populations. It was shown that the migrated cells had a more elongated and mesenchymal morphology. Daughter cells of the migrated subset also remained more chemotactic in character and showed higher expression of RhoC GTPase and p38γ (known mediators of migration) after four days of culture. The microfluidic device was further modified to incorporate a series of 100 μm long ‘choke points’ (6–30 μm) in each central migration channel to mimic cancer cell migration in lymphatic vessels. It was shown that cancer cells extended pseudopodia beyond the choke points, and that cells with p38γ knockdown were not able to migrate through the narrowest choke points.
Other than mechanical confinement of cancer cells, surface stiffness was also suggested to affect the behavior of cancer cells62,63, and exogenous tissue stiffening can facilitate tumorigenesis64. Therefore, a device was designed to observe the 3D migration of H1299 lung cancer cells in gels of different composition and stiffness65. The PDMS device consisted of a central hydrogel channel laterally connected to two media channels separated by 3 posts. Cancer cells were suspended in hydrogels of different compositions (2 mg mL−1 of type I collagen, 0–4 mg mL−1 of Matrigel) and loaded into the hydrogel channel. In addition, 20% fetal bovine serum (FBS) was loaded in the media channel to stimulate migration. The hydrogel with the highest concentration of Matrigel was verified as the stiffest, with the thickest fibers and fewer but larger pores. Cancer cells in the hydrogel with the most Matrigel were the most motile and had a lobopodial as opposed to mesenchymal phenotype. Anti-β1 and anti-β3 neutralizing antibodies triggered amoeboid migration and reduced migration speeds in hydrogels with 0–2 mg mL−1 Matrigel but increased those in hydrogel with 4 mg mL−1 Matrigel.
As stated, the above device altered both the stiffness and the size of the pores that confine migrating cancer cells. Therefore, Pathak et al. constructed a device to isolate the effect of stiffness and confinement66. The researchers fabricated microfluidic channels of different widths (10–40 μm) and stiffness (0.4–120 kPa) through polymerization and gelation of polyacrylamide hydrogels. Enhanced migration of U373-MG glioma cells in narrower channels was observed for a fixed stiffness. The dependence of migration speed on confinement was found to be the strongest on the stiffest ECM. The migration speed increased with stiffness (up to 120 kPa) in the 10 μm channels but reached a maximum at a stiffness value of 10 kPa in wider channels. Interestingly, the application of blebbistatin (an inhibitor of non-muscle myosin II a) saturated the dependence of migration speed on stiffness in all channels.
In addition to the biomaterial surface stiffness that cancer cells encounter, cell stiffness and deformability are major factors that regulate the cell migration speed in confined channels67. Therefore, several microfluidic devices were developed with the ability to measure the mechanical properties of cancer cells68–70, and one device was able to do so while measuring the speed of cancer cells passing through constrictions (Figure 2e)46. The device contained a comparator region at the end of two parallel confined channels (15 μm). With no cells in either confined channel, the liquid interface between fluids from the two channels was balanced at the center of the comparator region. When a cell passed through a channel, this interface moved toward the channel with the cell due to the reduced flow rate. This fluid interface displacement was dependent on the cell size and stiffness. This device showed that considerable differences in fluid interface displacement existed between benign cells and A712 glioblastoma cells even when the cells were of the same sizes, indicating that the difference is likely due to cell stiffness. Surprisingly, L0329 and L0367 normal astrocytes were found to be less stiff and migrated faster than cancerous glioblastoma A172 and astrocytoma 1321N1 cells. This observation is in striking contrast to the previous observations that breast cancer cells are softer than normal breast tissue cells70,71.
Another factor that plays an important role in cell migration is fluidic flow72,73. Cancer cells in vivo experience blood flow or interstitial flow in all stages of metastasis. Physiological levels of hemodynamic shear stress (15–30 dynes per cm2) have been shown to trigger apoptosis in non-metastatic cancer cells but enhance transwell migration of metastatic cells74. A microfluidic device was developed to model cancer cell movement in interstitial flow (Figure 2f)47. The device consisted of three parallel cell channels separated by microfabricated PDMS ridges and a flow channel perpendicular to and running through the cell channels. Cells suspended in collagen were subsequently loaded to the cell channels, and flow was introduced at 2 μm s−1, which is within the range of interstitial flow in healthy tissue75. Because there were only ridges and no walls, the obstruction of flow was greatly reduced. Using the device, it was found that flow promoted an amoeboid over a mesenchymal phenotype, exhibiting reduced cell dimensions and evenly distributed actin filaments. Because fibronectin is an adhesion molecule essential in migration with a mesenchymal phenotype, the researchers hypothesized that the flow washed away fibronectin, leading to an amoeboid phenotype. The observation that exogenous fibronectin (added to the cell-collagen solution at 100 μg mL−1) promoted a mesenchymal phenotype confirmed this hypothesis. Interestingly, although flow increased migration speed, the probability that cancer cells migrated persistently in one direction was reduced. This observation contrasts with the previous observation that interstitial flow guided cancer cell migration76 and suggests that further studies on interstitial fluid flow and cancer cell migration are needed. For example, gene expression studies can be useful in identifying the difference in cells that responded differently to interstitial flow and are expected to aid in a better understanding of how different cancer cells migrate in tissues and how migration can be deterred to prevent metastasis.
Microfluidic investigation of biochemical signals in cancer cell invasion
Metastasis to secondary tumor sites involves a plethora of chemokines. After cancer cells are arrested in the capillaries or slowed down in the sinusoid blood vessels, gradients of biochemical factors secreted by cells in the microenvironment might signal the tumor cells to extravasate out of the vessel. For example, epidermal growth factor77, CXCL-1278, hepatocyte growth factor79, and vascular endothelial growth factor were shown to stimulate cancer cell migration. Early models used to study biochemical factors used cell migration through bulk gels in response to factors including EGF80,81, CXCL-12 (Ref. 81), hypoxia82,83, and serum concentration84. These models allowed the study of cancer cell migration in a 3D environment, which is known to occur via mechanisms different from those in a 2D environment85,86. With these devices, morphological changes and matrix degradation were observed during cancer cell migration toward a nutrient gradient. The efficacy of drugs in inhibition of cancer cell migration and invasion were also compared. These devices have significant advantages over the wound-healing assay due to the incorporation of gradients and 3D migration environments, but they still lack intrinsic co-culture and cross-talk between cells.
The metastatic niche is highly dynamic, with interactions between tumor and resident cells. In addition to responding to chemokines, tumor cells actively produce factors to modify ECM, disrupt endothelial integrity, and recruit cells to form a favorable environment. For example, tumor cells upregulate VEGF to recruit endothelial cells and promote angiogenesis, transforming growth factor β (TGF-β) and macrophage colony-stimulating factor to recruit tumor-associated macrophages87, and platelet-derived growth factor and TGF-β to recruit CAFs88. These three cell types have been indicated to promote metastasis to a variety of secondary tumor sites. Therefore, more advanced microfluidic devices began to involve co-cultures with macrophages89,90, CAFs91–93, and endothelial cells94–101. One device used macrophages and cancer cells suspended in 2.5 mg mL−1 collagen (Figure 3a)90. Experiments demonstrated that the presence of macrophages enhanced cancer cell migration speed and persistence as well as their production of matrix metalloproteinases (MMPs, an essential factor for matrix degradation). Co-blocking of tumor necrosis factor α (TNF-α) and TGF-β reduced the production of MMPs only when macrophages were present, suggesting that the enhanced invasion was due to macrophage production of the two factors.
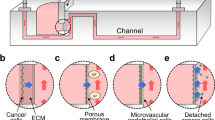
Modeling of cancer cell invasion. (a) Effect of macrophage on invasion of cancer cells. Adapted from Ref. 90 with permission. (b) Invasions of co-culture of cancer cells and cancer-associated fibroblasts. Adapted from Ref. 93 with permission. (c) First microfluidic extravasation device with endothelial cells. Adapted from Ref. 94 with permission. (d) Self-assembled microvasculature. Adapted from Ref. 97 with permission. (e) Cancer cell extravasation under flow. Adapted from Ref. 100 with permission. (f) Collection of cells extravasated under flow for post-analysis. Adapted from Ref. 101 with permission.
A similar device was constructed to study the interaction between salivary gland adenoid cystic carcinoma (ACC) cells and CAFs93 (Figure 3b). This device consisted of a stimulation channel (20% FBS) and a cell channel (co-culture of ACC cells and primary CAFs isolated from patients with ACC) connected by Matrigel-filled side channels. Studies found that co-cultures with CAFs were much more invasive than cultures with only ACC cells or co-cultures with regular fibroblasts. Furthermore, it was demonstrated that CAFs invaded in the front and ACC cells followed, whereas ACCs invaded first when co-cultured with normal fibroblasts. However, when the C-X-C receptor-4 (CXCR-4, receptor for CXCL-12) on ACC cells was blocked with AMD3100, ACC cells did not follow the track of CAFs and sometimes invaded in the front. These devices permitted the observation of intricate interactions between cancer cells and metastasis-supporting cells90,93. Although these approaches are excellent platforms for studying cancer cell migration in ECMs, it could be more insightful to incorporate endothelial cells to model blood vessels because endothelial cells are the first barriers encountered by cancer cells in blood vessels during extravasation.
Figure 3c shows an early device developed to study the transmigration of cancer cells across a layer of endothelial cells94, with a region on the device filled with basement membrane extract (BME) as a model ECM. To develop an endothelial wall, the device was filled with human umbilical vascular endothelial cells (HUVECs) and tipped on its side to attach the cells on the BME. Metastatic adenoid cystic carcinoma (ACC-M) cells aggregates were subsequently seeded on the endothelial cell layer. The application of CXCL-12 was able to induce trans-endothelial migration in the cancer cells. By antagonizing its receptor CXCR-4 with AMD3100, migration was blocked, which highlights the importance of CXCR-4 in metastases. A similar device was used to observe MDA-MB-231 extravasation toward an CXCL-12 gradient95. However, these devices used a non-luminal vasculature, which could affect signaling of endothelial cells102.
To improve the geometry of vascular models, a device was built96 based on the design of a co-culture device103. The device consisted of a central channel coated in Matrigel and seeded on all sides with human microvascular endothelial cells. The transmigration of metastatic breast cancer cells MDA-MB-231 seeded on the endothelial cells was investigated. The device allowed visualization of the damage that cancer cells caused to the endothelial wall during transmigration. It was also observed that the majority of the transmigration occurred shortly after cell adhesion (within 24 h). However, different from a blood vessel, the microfluidic device had a rectangular cross-section, and this shape difference could possibly affect the interaction between cancerous and endothelial cells104.
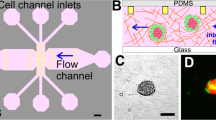
To develop rounded vasculature in a rectangular channel, the ability of endothelial cells to self-assemble microvasculature was leveraged (Figure 3d)97. The device consisted of two fibrinogen cell channels separated by three media channels. To induce microvasculature self-organization, HUVECs were seeded in fibrinogen in one cell channel, and normal lung fibroblasts were seeded in the other. This co-culture allowed for the development of more physiologically relevant microvasculature compared with that of endothelial monocultures105,106. MDA-MB-231 cells were loaded into the central media channel, which induced a pressure drop that caused the cancer cells to enter the microvasculature. In the experiments, the application of proinflammatory cytokine TNF-α greatly increased the permeability of the microvasculature and cancer cell extravasation in a dose-dependent manner. Real-time imaging showed that cancer cells first extended thin filopodial protrusions through the endothelial cell layer, which subsequently grew and branched out. The cell body was reduced in size as it penetrated the endothelial cell layer, and as it entered the new ECM, it developed a ‘spread’ morphology. After extravasation, no breakage in the E-cadherin junctions was observed. However, a small gap (up to 8 μm) was observed to form during transmigration. Furthermore, it was noted that cancer cells trapped in the vessel had a higher extravasation efficiency compared with those that were adhered yet free to move.
The microfluidic device97 was used to further investigate the importance of integrin β1 in breast cancer cell extravasation98. small hairpin RNA (shRNAs) targeting integrin β1 caused a significant reduction in MDA-MB-231 cell protrusion and extravasation at 6 h, whereas those targeting integrin β3 had no effect. Moreover, it was shown that co-blocking of laminin-binding integrin α3 and α6 also reduced extravasation, although not to the level of β1KD cells. A similar approach of self-assembled microvasculature was applied to demonstrate that Taxol not only greatly reduced the migration and growth of MDA-MB-231 breast cancer cells but also disrupted the microvasculature around the cancer cells99.
To model the full extravasation process of circulating tumor cells (CTC) with flow, a microfluidic device was developed (Figure 3e)100 consisting of a perfusion channel with cancer cell suspension and a gel channel loaded with CXCL-12. A monolayer of HUVECs was seeded on Matrigel on the walls of the cancer cell channel. The chemokine channel used Matrigel as a model ECM. Flow (50 μL h−1, 6.7×10−3 Pa) was applied to the cancer cell channel to develop a confluent monolayer of endothelial cells and to pump in the suspension of CTCs. The flow rate was subsequently lowered (1 μL h−1, 1.3×10−4 Pa) to establish a stable CXCL-12 gradient in the channel. The study showed that CXCL-12 increased the numbers of extravasated MDA-MB-231 cells but not MCF7 cells. It should be noted that although this system was the first to address the need for fluid flow on endothelial cells for a valid model, the shear stresses applied were well below physiological levels107.
In addition to observing the effect of different factors on extravasation, due to the increasing interest in intra-tumor heterogeneity, a microfluidic device was developed to isolate invasive subsets for analysis (Figure 3f)101. The device consisted of a porous membrane with 20 μm thickness separating a top channel layer and bottom collection chambers. Pore sizes between 24 and 28 μm were found to allow endothelial cells to cover the pores and to allow cancer cells to maintain their morphology after migration. Because each collection chamber was connected to an independent inlet and outlet controlled by pneumatic microvalves, cells migrating across areas where the endothelial cell layer was intact were collected. In the device, the membrane was coated with 4 μg cm−2 poly-D-lysine and 50 μg mL−1 fibronectin and seeded with HUVECs and cancer cells. CXCL-12 (100 ng mL−1) was subsequently added to the bottom collection chambers, and flow with a shear stress of ~10 dynes per cm2 was applied to the flow channel. After 15 h, 5.2% of MDA-MB-231 invasive breast cancer cells and only 0.4% of MCF-10A normal breast epithelial cells had migrated. Furthermore, when a lower flow rate (shear stress of 2.5 dynes per cm2) was applied, 9.1% of MDA-MB-231 cells migrated, suggesting that the higher shear stress might help to maintain an intact layer of endothelial cells. The migrated cells were subsequently collected and incubated for further analysis. Similar to the isolated subsets of cancer cells migrating through confinement45, the MDA-MB-231 cells capable of trans-endothelial migration were also more spread and revealed a mesenchymal morphology.
Microfluidic metastasis-on-a-chip models for investigation of cancer extravasation
As discussed previously, different types of cancer preferentially metastasize to different tissues. A number of organ-on-a-chip models have been developed for organs that are common secondary sites of metastases (such as lung, liver, and bone), as summarized in several review papers108–110. These organ-on-a-chip models could be subsequently combined with vasculature extravasation models discussed in section ‘Microfluidic investigation of biochemical signals in cancer cell invasion’111 to create metastasis-on-a-chip models that mimic cancer cell extravasation through an endothelial barrier toward the secondary metastasis site. These models are advantageous for investigating the interactions between cancer cells and cells at the secondary site and the mechanisms regulating organ specificity of metastases.
Breast cancer is known to metastasize preferentially to bone. An early microfluidic device investigating this specificity consisted of a blood vessel channel and a bone channel (Figure 4a)112. The bone channel contained osteo-differentiated (OD) human bone marrow-derived mesenchymal stem cells (hBM-MSCs) seeded within a collagen gel. Osteogenic medium was supplied for 3 days to induce bone formation, after which the blood vessel channel was coated with Matrigel and seeded with HUVECs. After another 3 days, the vessel was seeded with MDA-MB-231 metastatic breast cancer cells. With this model, a substantial increase in MDA-MB-231 extravasation rate and migration distance was observed compared with a simple collagen gel without hBM-MSCs. To study the potential mechanisms related to the increased extravasation, the effect of CXCL-5 (a ligand expressed by osteo-cells) was investigated together with its receptor CXCR-2 (on MDA-MB-231 cells). Specifically, the application of CXCL-5 to a simple collagen gel increased extravasation, whereas the application of CXCR-2 blocking antibody decreased extravasation. The development of cancer cell aggregates of at least 4 cancer cells within the bone ECM after extravasation was also observed, showing the ability of cancer cells to proliferate within this environment.
A microfluidic microvasculature model was also reported for studying breast cancer metastases into bone tissue113. In this model, all of the cells were incorporated in one ECM channel, which was surrounded on both sides by media channels. To model bone, the ECM channel was loaded with a fibrin gel containing primary hBM-MSCs, OD hBM-MSCs, and HUVECs. Over a period of 4 days, the HUVECs self-assembled a microvascular network, after which a bone-seeking subclone of MDA-MB-231 (BOKL) was loaded into the network. It was shown that extravasation rates significantly increased when co-cultured with OD cells compared with both control myoblasts (C2C12) and without the addition of cells. RAW264.7 macrophages also increased extravasation rates compared with the myoblast control, but the rates were lower than with the OD cells. To verify the specificity of the metastatic cancer cells to the bone environment, the extravasation rate of non-metastatic mammary epithelial cells (MCF-10A) was investigated, and it was found to be significantly lower. Furthermore, to understand the protective effect that skeletal muscle has against metastases, the A3 adenosine receptor expressed by BOKL was supplied while the matrix was seeded with C2C12. The extravasation rate was significantly increased, indirectly demonstrating the role that adenosine has in anti-metastases. As well, the addition of adenosine to the ECM when supplied with OD cells resulted in a significant decrease in the extravasation rate. Finally, this study addressed the issue that many previous vasculature models failed to study, which is the role of physiological fluid shear stress supplied to the endothelial cells. The researchers preconditioned the endothelial cells with flow before the addition of cancer cells and observed a significantly decreased extravasation rate compared with the statically maintained endothelial cells, which was attributed to the change in microvessel permeability.
A device was also designed to mimic the blood–brain barrier (BBB) and brain metastasis (Figure 4b)114. This device consisted of an array of 16 units, with four identical BBB regions in each unit. The units were formed with one vascular channel, four gel channels (each representing one BBB region), and one gas channel with a gas valve at the end of the vascular channel to control the flow. To form the BBB, collagen gel was first added to the gel channel, and primary rat cerebral astrocytes and brain microvascular endothelial cells (BMECs) were added in respective order to the vascular channel and allowed to adhere to the collagen surface. Media was applied to the vascular channel at 1 μL min−1 (0.1 dyne per cm2 shear stress), mimicking the capillary flow in the brain. It was observed that with the addition of flow and/or astrocytes, BMECs increased their expression of tight junction (ZO-1 and Claudin-5) and adhesion (VE-Cadherin) proteins. Furthermore, the expression of transporter proteins (P-gp and Glut-1) was increased with the addition of astrocytes alone but not flow. Both tracer and transendothelial electrical resistance experiments showed that the presence of flow and/or astrocytes reduced BBB permeability. When the cancer cells were added to the vascular channel, cancer cells that are known to metastasize to the brain (A549 lung cancer cells, MDA-MB-231 breast cancer cells, and M624 melanoma cells) were able to disrupt the integrity of the BBB and migrate past the BBB, but BEL-7402 liver cancer cells did not. U87 glioma cells from brain tumor were also shown to be unable to cross the BBB when seeded in collagen gels.
Ma et al. explored cancer cell extravasation toward different ‘organs’ with a microfluidic device (Figure 5a)115 consisting of a porous polycarbonate membrane (20 μm thick, 8 μm pores) sandwiched between two PDMS layers (300 μm thick) with an array of through-holes in variable patterns. The carefully aligned PDMS layers essentially create a series of microwells with a porous membrane in the middle. Two thick PDMS rings were bonded to the outside for support. The device was placed in oil to avoid media evaporation, and a polymethylmethacrylate (PMMA) ring was placed on top to prevent the device from floating. Droplets of Matrigel (1.1 mg mL−1) were deposited into the microwells with a tapered capillary and a syringe pump. For simple migration studies, the inducer droplet (20% FBS) was deposited on the underside of the membrane, and a cell droplet was deposited on the upside of the membrane, forming a Boyden-chamber-like system. The number of migrated cells was quantified by imaging the total number of cells in the system and subsequently imaging the cells after scraping off the top droplet. It was found that the metastatic MDA-MB-231 breast cancer cells significantly increased migration toward the inducer droplet in the presence of 20% FBS;, whereas the non-metastatic MCF-7 breast cancer cells did not exhibit significant migration with or without 20% FBS. This system was expanded to conduct studies on cancer cell migration toward cell cultures from different organs. This ‘multi-tissue’ set-up consisted of one large cancer cell droplet on the top side, and five inducer droplets, each overlapping with the cancer cell droplet but not touching each other, on the underside of the membrane. The five inducer droplets each contained a different type of cell to realize a 3D co-culture of cells. This system successfully demonstrated the migration of MDA-MB-231 cells toward different inducer droplets, although the preference in migration pattern was not significant.
Modeling of metastasis using multiple ‘organs-on-a-chip’ to test for metastatic specificity. (a) Array of microdroplets with different cell cultures on the underside for study of cancer cell migration toward different ‘droplets’. Adapted from Ref. 115 with permission. (b) Modeling of metastasis from lung to brain, bone, and liver downstream organs (assembly i-vi). Adapted from Ref. 116 with permission.
Figure 5b shows a multi-organ system that mimics lung cancer cell extravasation to common sites for lung metastasis, namely, brain, bone, and liver116. The device consisted of three PDMS layers separated by two PDMS porous membranes coated with ECM, creating three channels. The top bronchial channel contained 16HBE human bronchial epithelial cells seeded on top of the membrane and was exposed to air, whereas the middle microvascular channel was coated with HUVEC endothelial cells and stromal cells (WI38 human lung fibroblasts and stimulated THP-1 macrophages) and filled with media. A cyclic vacuum was applied to the hollow side-chambers to stretch the membrane (10% cyclic strain at 0.2 Hz) to mimic physiological breathing. HA-1800 astrocytes, Fob1.19 osteoblasts, and L-02 hepatocytes were cultured separately in three chambers at the bottom layer that were linked to the microvascular channel via side channels to represent brain, bone, and liver, respectively. A549 human lung cancer cells were co-cultured with 16HBE epithelial cells, and media flowed through the microvascular channel to mimic blood flow (24 mm h−1). The A549 cells co-cultured with stromal cells expressed lower E-cadherin and higher N-cadherin, Snail1, and Snail2 and were more invasive toward the microvascular channel than monocultured A549 cells, similar to the findings reported by Yu et al.92 Furthermore, after A549 metastasis to the three ‘distant organ’ chambers, astrocytes overexpressed CXCR4, osteoblasts overexpressed RANKL, and hepatocytes overexpressed AFP, consistent with previous studies on lung cancer metastasis117–119. The microfluidic device was validated to be capable of mimicking lung cancer metastasis and offers significant potential for studying the mechanisms, drug effects, and site specificity of lung cancer metastasis.
Summary and outlook
This paper reviewed a multitude of microfluidic devices for modeling of cancer cell extravasation during metastasis. By leveraging the advantages of microfluidics, these devices modeled extravasation, manipulated specific factors, and studied single-cell responses. The devices were developed to investigate cancer cell migration under the impact of (a) mechanical factors (section ‘Microfluidic investigation of mechanical factors in cancer cell migration’), (b) biochemical factors (section ‘Microfluidic investigation of biochemical signals in cancer cell invasion’), and (c) microenvironments of the secondary site in metastasis-on-a-chip devices (section ‘Microfluidic metastasis-on-a-chip models for investigation of cancer extravasation’). A summary of these devices is presented in Table 1 that outlines the type of cancer cells studied, the substrate with which cancer cells are in contact, the extravasation matrix, whether flow is involved, and the chemokine sources (applied or cell-derived).
The devices discussed in section ‘Microfluidic investigation of mechanical factors in cancer cell migration’ demonstrated strong mechanical regulation of cancer cell migration. Confinement of the cancer cells increased their migration speed and persistence42,48,66. The migration speed also increased with substrate stiffness in confined channels66. It was observed that as individual cancer cells approached confined regions, they first extended protrusions into the confined channels prior to nucleus deformation, and migration occurred43,52,54,55. These pre-migration protrusions were also observed in cancer cell extravasation across self-assembled endothelial networks97. In addition, the clustered cancer cells transformed into a chain-like organization as they entered the confined channels44. Furthermore, a number of studies have shown that invasive cancer cells were more responsive to changes in the mechanical environment than less invasive and non-cancerous cells42,48,49,52. Chemotherapy drugs (for example, Taxol) were often tested in microfluidic studies and were shown to reduce cancer cell migration speed in confinement42,43,57,58.
Section ‘Microfluidic investigation of biochemical signals in cancer cell invasion’ discussed how biochemical or cellular factors affect cancer cell invasion. These devices allowed the 3D migration of cancer cells to be studied in response to a biochemical gradient or in co-culture with tumor-associated macrophages90 and CAFs93. To better simulate cancer cell extravasation, microfluidic devices began to include endothelial cells in the form of a layer94,95,100, a lumen96, and self-assembled networks97–99. Various factors involved in cancer cell extravasation were investigated, such as CXCL-12 94,95,100, tumor integrin β198, and TNF-α97. Microfluidic devices have also been developed to isolate subpopulations that were able to migrate through confinement45 and endothelial cells101 to further investigate intra-tumor heterogeneity.
The microfluidic devices described in section ‘Microfluidic metastasis-on-a-chip models for investigation of cancer extravasation’ are the most complex and involve cell cultures of the targeted organs. For example, breast cancer extravasation toward bone multicellular cultures was modeled and demonstrated increased extravasation compared with gels without bone cells112,113. Cancer cells that are known to metastasize to the brain in vivo have been shown to disrupt and migrate past the BBB, whereas other types of cancer cells were not able to perform these functions114. Multi-organ devices have also been developed with multiple cancer cell migration pathways leading to cell cultures of different downstream organs115,116, and cancer cells in these devices were able to migrate to different cell cultures and affect protein expressions in the downstream cell cultures116.
These advances demonstrated the potential of microfluidic devices for modeling intricate interactions and resulted in valuable findings. However, Table 1 also makes apparent a few areas where further improvement is needed. First, few of the current extravasation models incorporated the flow that cancer cells experience in vivo. This aspect is particularly important because fluid shear stress is known to affect the behavior of endothelial cells120–122 and cancer cells74,123. Secondly, few of the gel models considered the stiffness of the ECM into which the cell migrates, even though the stiffness of ECM is well known to play a major role in cancer cell migration62,66,124. Notably few existing devices considered both mechanical and biochemical or cellular factors. Furthermore, many of the studies used MDA-MB-231 breast cancer cells (a highly metastatic cell line), probably because breast cancer has a high likelihood of metastasis, as observed clinically. However, metastasis could result from any type of cancer, and studies should move toward other primary tumor types. The use of primary cells isolated from patients should also be increased. Therefore, substantial room still exists in this field for further innovations to improve the physiological relevance of microfluidic devices.
Because the individual ‘organ-on-a-chip’ devices combine multiple factors and improve on physiological relevance, it can be envisioned that they can be integrated synergistically to create ‘body-on-a-chip’ or ‘metastasis-on-a-chip’ devices. These devices can be used to identify metastatic specificity and drug targets and to perform drug screening for personalized medicine. Personalized treatment gained recognition after significant heterogeneity in the tumor and drug responses of patients were observed. Ideally, primary tumor cells isolated from individual patients should be obtained and added to these ‘metastasis-on-a-chip’ devices to test their response to cancer drugs in terms of migration and apoptosis in a physiological environment. Microfluidic devices developed for isolating circulating tumor cells125 could be used to obtain tumor cells and add them to the ‘metastasis-on-a-chip’ devices. Off-target toxicity could also be identified with a more comprehensive ‘body-on-a-chip’ that includes, for example, liver and gut models for drug metabolism. Cancer cell dormancy and the interaction with other cell populations during reactivation should also be included. These devices could also benefit greatly by further improving the isolation and quantification methods. Subpopulations of tumor cells that did not respond to drugs could be investigated for intra-tumor heterogeneity. Downstream assays for gene and protein expression profiles might be used to identify the differences in cancer cells that are responsive or resistant to the drugs applied. This approach might lead to the use of promising immunotherapy126 to modulate immune cells to target the resistant subpopulations. Identifying how different cells in a tumor microenvironment affect cancer cell activities might also lead to drug targets that can reduce host support for metastasis.
To fully utilize the advantages of the in vitro microfluidic devices over current in vivo systems, these devices should incorporate immune cells, primary human cells, and the capability of mass production. Because mice models could only include either immune cells or primary human tumor cells, microfluidics offer advantages for studying complex and important interactions between metastasis and the immune system15. In addition to human tumor cells, microfluidics could include individualized human cells as the techniques to produce induced pluripotent stem (iPS) cells become standard127. Even without iPS cells, additional human cell lines should be incorporated to account for the potential differences between human and mice models. The addition of these human cells (and specifically, immune cells) could greatly improve the ability of future microfluidic devices to model the intricate metastasis environment in vivo. In the meantime, it is also important to preserve the relative simplicity of microfluidic devices and their ability to perform high-throughput studies. Further work is required to identify the minimal essential factors and cells that must be included in the final ‘metastasis-on-a-chip’ platform. Parameters such as the variety of biomaterials needed, the media used to maintain multiple cell types, the ratio of cell numbers from each cell type to better mimic the in vivo environment and avoid over-representation, and the drug concentration also need to be optimized. Only devices designed with simplicity and scale-up capability128 can embrace the full potential of microfluidics.
In summary, current microfluidic devices have demonstrated great potential to fine tune specific physical, biochemical, or cellular signals to investigate metastasis. Integrating the different factors and ‘organ-on-a-chip’ devices into a full ‘metastasis-on-a-chip’ could create a platform for drug target identification and individualized medicine screening. Adopting advances from related fields, such as induced pluripotent stem cells and biomaterials, could further improve the physiological relevance of these models. A ‘metastasis-on-a-chip’ with the correct balance between capturing the complexity of the metastatic cascade and ability to be mass produced could revolutionize cancer treatment and could allow drug investigation and screening in a high-throughput and predictive fashion to identify effective, personalized therapeutic approaches for managing cancer metastasis.
References
Fidler IJ . The pathogenesis of cancer metastasis: the ‘seed and soil’ hypothesis revisited. Nature Reviews Cancer 2003; 3: 453–458.
Weilbaecher KN, Guise TA, McCauley LK . Cancer to bone: a fatal attraction. Nature Reviews Cancer 2011; 11: 411–425.
Steeg PS . Targeting metastasis. Nature Reviews Cancer 2016; 16: 201–218.
Cancer. World Health Orgnanization 2015. http://www.who.int/mediacentre/factsheets/fs297/en/ (accessed 24 October 2016).
Cuddapah VA, Robel S, Watkins S et al. A neurocentric perspective on glioma invasion. Nature Reviews Neuroscience 2014; 15: 455–465.
Tabassum DP, Polyak K . Tumorigenesis: It takes a village. Nature Reviews Cancer 2015; 15: 473–483.
Lim B, Hortobagyi GN . Current challenges of metastatic breast cancer. Cancer Metastasis Reviews 2016; 35: 495–514.
Klemm F, Joyce JA . Microenvironmental regulation of therapeutic response in cancer. Trends in Cell Biology 2015; 25: 198–213.
Plaks V, Kong N, Werb Z . The cancer stem cell niche: How essential is the niche in regulating stemness of tumor cells? Cell Stem Cell 2015; 16: 225–238.
Gupta GP, Massagué J . Cancer metastasis: Building a framework. Cell 2006; 127: 679–695.
Sahai E . Mechanisms of cancer cell invasion. Current Opinion in Genetics & Development 2005; 15: 87–96.
Sleeman J, Steeg PS . Cancer metastasis as a therapeutic target. European Journal of Cancer (Oxford, England: 1990) 2010; 46: 1177–1180.
Jenkins DE, Oei Y, Hornig YS et al. Bioluminescent imaging (BLI) to improve and refine traditional murine models of tumor growth and metastasis. Clinical & Experimental Metastasis 2003; 20: 733–744.
Sahai E . Illuminating the metastatic process. Nature Reviews Cancer 2007; 7: 737–749.
Kitamura T, Qian B-Z, Pollard JW . Immune cell promotion of metastasis. Nature Reviews Immunology 2015; 15: 73–86.
Noy R, Pollard JW . Tumor-associated macrophages: From mechanisms to therapy. Immunity 2014; 41: 49–61.
Caponigro G, Sellers WR . Advances in the preclinical testing of cancer therapeutic hypotheses. Nature Reviews Drug Discovery 2011; 10: 179–187.
Day C-P, Merlino G, Van Dyke T . Preclinical mouse cancer models: A maze of opportunities and challenges. Cell 2015; 163: 39–53.
Kelloff GJ, Sigman CC . Cancer biomarkers: selecting the right drug for the right patient. Nature Reviews Drug Discovery 2012; 11: 201–214.
Volm M, Efferth T . Prediction of cancer drug resistance and implications for personalized medicine. Frontiers in Oncology 2015; 5: 282.
LaBarbera DV, Reid BG, Yoo BH . The multicellular tumor spheroid model for high-throughput cancer drug discovery. .Expert Opinion on Drug Discovery 2012; 7: 819–830.
Xu X, Farach-Carson MC, Jia X . Three-dimensional in vitro tumor models for Cancer Research and drug evaluation. Biotechnology Advances 2014; 32: 1256–1268.
Decaestecker C, Debeir O, Van Ham P et al. Can anti-migratory drugs be screened in vitro? A review of 2D and 3D assays for the quantitative analysis of cell migration. Medicinal Research Reviews 2007; 27: 149–176.
Hanahan D, Weinberg RA . Hallmarks of cancer: The next generation. Cell 2011; 144: 646–674.
Peela N, Truong D, Saini H et al. Advanced biomaterials and microengineering technologies to recapitulate the stepwise process of cancer metastasis. Biomaterials 2017; 133: 176–207.
Samatov TR, Shkurnikov MU, Tonevitskaya SA et al. Modelling the metastatic cascade by in vitro microfluidic platforms. Progress in Histochemistry and Cytochemistry 2015; 49: 21–29.
Portillo-Lara R, Annabi N . Microengineered cancer-on-a-chip platforms to study the metastatic microenvironment. Lab on a Chip 2016; 16: 4063–4081.
Huang YL, Segall JE, Wu M . Microfluidic modeling of the biophysical microenvironment in tumor cell invasion. Lab on a Chip 2017; 17: 3221–3233.
Esch EW, Bahinski A, Huh D . Organs-on-chips at the frontiers of drug discovery. Nature Reviews Drug Discovery 2015; 14: 248–260.
Du G, Fang Q, den Toonder JMJ . Microfluidics for cell-based high throughput screening platforms—A review. Analytica Chimica Acta 2016; 903: 36–50.
Kurozumi K, Ichikawa T, Onishi M et al. Cilengitide treatment for malignant glioma: current status and future direction. Neurologia Medicochirurgica 2012; 52: 539–547.
Dubrovska A, Cojoc M, Peitzsch et al. Emerging targets in cancer management: role of the CXCL12/CXCR4 axis. Oncotargets and Therapy 2013; 6: 1347.
Fizazi K, Carducci M, Smith M et al. Denosumab versus zoledronic acid for treatment of bone metastases in men with castration-resistant prostate cancer: a randomised, double-blind study. Lancet (London, England) 2011; 377: 813–822.
Sun Y, Ma L . The emerging molecular machinery and therapeutic targets of metastasis. Trends in Pharmacological Sciences 2015; 36: 349–359.
Chambers AF, Groom AC, MacDonald IC . Dissemination and growth of cancer cells in metastatic sites. Nature Reviews Cancer 2002; 2: 563–572.
Wolf K, Te Lindert M, Krause M et al. Physical limits of cell migration: Control by ECM space and nuclear deformation and tuning by proteolysis and traction force. The Journal of Cell Biology 2013; 201: 1069–1084.
Peyton SR, Putnam AJ . Extracellular matrix rigidity governs smooth muscle cell motility in a biphasic fashion. Journal of Cellular Physiology 2005; 204: 198–209.
Hawkins RJ, Piel M, Faure-Andre G et al. Pushing off the walls: A mechanism of cell motility in confinement. Physical Review Letters 2009; 102: 58103.
Paul CD, Hung W-C, Wirtz D et al. Engineered models of confined cell migration. Annual Review of Biomedical Engineering 2016; 18: 159–180.
Clark AG, Vignjevic DM . Modes of cancer cell invasion and the role of the microenvironment. Current Opinion in Cell Biology 2015; 36: 13–22.
Prathivadhi-Bhayankaram SV, Ning J, Mimlitz M et al. Chemotherapy impedes in vitro microcirculation and promotes migration of leukemic cells with impact on metastasis. Biochemical and Biophysical Research Communications 2016; 479: 841–846.
Irimia D, Toner M . Spontaneous migration of cancer cells under conditions of mechanical confinement. Integrative Biology: Quantitative Biosciences from Nano to Macro 2009; 1: 506–512.
Mak M, Reinhart-King CA, Erickson D . Elucidating mechanical transition effects of invading cancer cells with a subnucleus-scaled microfluidic serial dimensional modulation device. Lab on a Chip 2013; 13: 340–348.
Au SH, Storey BD, Moore JC et al. Clusters of circulating tumor cells traverse capillary-sized vessels. Proceedings of the National Academy of Sciences of the United States of America 2016; 113: 4947–4952.
Chen Y-C, Allen SG, Ingram PN et al. Single-cell migration chip for chemotaxis-based microfluidic selection of heterogeneous cell populations. Scientific Reports 2015; 5: 9980.
Khan ZS, Vanapalli SA . Probing the mechanical properties of brain cancer cells using a microfluidic cell squeezer device. Biomicrofluidics 2013; 7: 11806.
Huang YL, Tung C, Zheng A et al. Interstitial flows promote amoeboid over mesenchymal motility of breast cancer cells revealed by a three dimensional microfluidic model. Integrative Biology: Quantitative Biosciences from Nano to Macro 2015; 7: 1402–1411.
Huang TQ, Qu X, Liu J et al. 3D printing of biomimetic microstructures for cancer cell migration. Biomedical Microdevices 2014; 16: 127–132.
Mak M, Reinhart-King CA, Erickson D . Microfabricated physical spatial gradients for investigating cell migration and invasion dynamics. PLoS ONE 2011; 6: e20825.
Mecham RP, Heuser J . Three-dimensional organization of extracellular matrix in elastic cartilage as viewed by quick freeze, deep etch electron microscopy. Connective Tissue Research 1990; 24: 83–93.
Dahl KN, Ribeiro AJS, Lammerding J . Nuclear shape, mechanics, and mechanotransduction. Circulation Research 2008; 102: 1307–1318.
Fu Y, Chin LK, Bourouina T et al. Nuclear deformation during breast cancer cell transmigration. Lab on a Chip 2012; 12: 3774–3778.
Rolli CG, Seufferlein T, Kemkemer R et al. Impact of tumor cell cytoskeleton organization on invasiveness and migration: a microchannel-based approach. PLoS ONE 2010; 5: e8726.
Davidson PM, Sliz J, Isermann P et al. Design of a microfluidic device to quantify dynamic intra-nuclear deformation during cell migration through confining environments. Integrative Biology: Quantitative Biosciences from Nano to Macro 2015; 7: 1534–1546.
Malboubi M, Jayo A, Parsons M et al. An open access microfluidic device for the study of the physical limits of cancer cell deformation during migration in confined environments. Microelectronic Engineering 2015; 144: 42–45.
Tong Z, Balzer EM, Dallas MR et al. Chemotaxis of cell populations through confined spaces at single-cell resolution. PLoS ONE 2012; 7: e29211.
Balzer EM, Tong Z, Paul CD et al. Physical confinement alters tumor cell adhesion and migration phenotypes. FASEB Journal: Official Publication of the Federation of American Societies for Experimental Biology 2012; 26: 4045–4056.
Mak M, Erickson D . Mechanical decision trees for investigating and modulating single-cell cancer invasion dynamics. Lab on a Chip 2014; 14: 964–971.
Yu M, Bardia A, Wittner BS et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science (New York, NY) 2013; 339: 580–584.
Molnar B, Ladanyi A, Tanko L et al. Circulating tumor cell clusters in the peripheral blood of colorectal cancer patients. Cancer ResearchCancer ResearchClinical Cancer Research: an Official Journal of the American Association for Cancer Research 2001; 7: 4080–4085.
Aceto N, Bardia A, Miyamoto DT et al. Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell 2014; 158: 1110–1122.
Tilghman RW, Cowan CR, Mih JD et al. Matrix rigidity regulates cancer cell growth and cellular phenotype. PLoS ONE 2010; 5: e12905.
Schrader J, Gordon-Walker TT, Aucott RL et al. Matrix stiffness modulates proliferation, chemotherapeutic response, and dormancy in hepatocellular carcinoma cells. Hepatology (Baltimore, Md) 2011; 53: 1192–1205.
Lam WA, Cao L, Umesh V et al. Extracellular matrix rigidity modulates neuroblastoma cell differentiation and N-myc expression. Molecular Cancer 2010; 9: 35.
Anguiano M, Castilla C, Maška M et al. Characterization of three-dimensional cancer cell migration in mixed collagen-Matrigel scaffolds using microfluidics and image analysis. PLoS ONE 2017; 12: e0171417.
Pathak A, Kumar S . Independent regulation of tumor cell migration by matrix stiffness and confinement. Proceedings of the National Academy of Sciences of the United States of America 2012; 109: 10334–10339.
Guck J, Schinkinger S, Lincoln B et al. Optical deformability as an inherent cell marker for testing malignant transformation and metastatic competence. Biophysical Journal 2005; 88: 3689–3698.
Lange JR, Steinwachs J, Kolb T et al. Microconstriction arrays for high-throughput quantitative measurements of cell mechanical properties. Biophysical Journal 2015; 109: 26–34.
Lautscham LA, Kämmerer C, Lange JR et al. Migration in confined 3D environments is determined by a combination of adhesiveness, nuclear volume, contractility, and cell stiffness. Biophysical Journal 2015; 109: 900–913.
Hou HW, Li QS, Lee GYH et al. Deformability study of breast cancer cells using microfluidics. Biomedical Microdevices 2009; 11: 557–564.
Li QS, Lee GYH, Ong CN et al. AFM indentation study of breast cancer cells. Biochemical and Biophysical Research Communications 2008; 374: 609–613.
Polacheck WJ, Charest JL, Kamm RD . Interstitial flow influences direction of tumor cell migration through competing mechanisms. Proceedings of the National Academy of Sciences of the United States of America 2011; 108: 11115–11120.
Haessler U, Teo JCM, Foretay D et al. Migration dynamics of breast cancer cells in a tunable 3D interstitial flow chamber. Integrative Biology: Quantitative Biosciences from Nano to Macro 2012; 4: 401–409.
Ma S, Fu A, Chiew GGY et al. Hemodynamic shear stress stimulates migration and extravasation of tumor cells by elevating cellular oxidative level. Cancer letters 2017; 388: 239–248.
Chary SR, Jain RK . Direct measurement of interstitial convection and diffusion of albumin in normal and neoplastic tissues by fluorescence photobleaching. Proceedings of the National Academy of Sciences of the United States of America 1989; 86: 5385–5389.
Polacheck WJ, German AE, Mammoto A et al. Mechanotransduction of fluid stresses governs 3D cell migration. Proceedings of the National Academy of Sciences of the United States of America 2014; 111: 2447–2452.
Sun R, Gao P, Chen L et al. Protein kinase C zeta is required for epidermal growth factor-induced chemotaxis of human breast cancer cells. Cancer Research 2005; 65: 1433–1441.
Müller A, Homey B, Soto H et al. Involvement of chemokine receptors in breast cancer metastasis. Nature 2001; 410: 50–56.
Gherardi E, Birchmeier W, Birchmeier C et al. Targeting MET in cancer: rationale and progress. Nature Reviews Cancer 2012; 12: 89–103.
Abhyankar VV, Toepke MW, Cortesio CL et al. A platform for assessing chemotactic migration within a spatiotemporally defined 3D microenvironment. Lab on a Chip 2008; 8: 1507–1515.
Kim BJ, Hannanta-anan P, Chau M et al. Cooperative roles of SDF-1α and EGF gradients on tumor cell migration revealed by a robust 3D microfluidic model. PLoS ONE 2013; 8: e68422.
Acosta MA, Jiang X, Huang P-K et al. A microfluidic device to study cancer metastasis under chronic and intermittent hypoxia. Biomicrofluidics 2014; 8: 54117.
Kalchman J, Fujioka S, Chung S et al. A three-dimensional microfluidic tumor cell migration assay to screen the effect of anti-migratory drugs and interstitial flow. Microfluidics and Nanofluidics 2013; 14: 969–981.
Chaw KC, Manimaran M, Tay FEH et al. Matrigel coated polydimethylsiloxane based microfluidic devices for studying metastatic and non-metastatic cancer cell invasion and migration. Biomedical Microdevices 2007; 9: 597–602.
Even-Ram S, Yamada KM . Cell migration in 3D matrix. Current Opinion in Cell Biology 2005; 17: 524–532.
Baumann K . Switching to 3D. Nature Reviews Molecular Cell Biology 2012; 13: 338.
Gong D, Shi W, Yi S et al. TGFβ signaling plays a critical role in promoting alternative macrophage activation. BMC Immunology 2012; 13: 31.
Karagiannis GS, Poutahidis T, Erdman SE et al. Cancer-associated fibroblasts drive the progression of metastasis through both paracrine and mechanical pressure on cancer tissue. Molecular Cancer Research 2012; 10: 1403–1418.
Huang CP, Lu J, Seon H et al. Engineering microscale cellular niches for three-dimensional multicellular co-cultures. Lab on a Chip 2009; 9: 1740.
Li R, Hebert JD, Lee TA et al. Macrophage-secreted TNFα and TGFβ1 influence migration speed and persistence of cancer cells in 3D tissue culture via independent pathways. Cancer Research 2017; 77: 279–290.
Hsu T-H, Xiao J-L, Tsao Y-W et al. Analysis of the paracrine loop between cancer cells and fibroblasts using a microfluidic chip. Lab on a Chip 2011; 11: 1808–1814.
Yu T, Guo Z, Fan H et al. Cancer-associated fibroblasts promote non-small cell lung cancer cell invasion by upregulation of glucose-regulated protein 78 (GRP78) expression in an integrated bionic microfluidic device. Oncotarget 2016; 7: 25593–25603.
Li J, Jia Z, Kong J et al. Carcinoma-associated fibroblasts lead the invasion of salivary gland adenoid cystic carcinoma cells by creating an invasive track. PLoS ONE 2016; 11: e0150247.
Zhang Q, Liu T, Qin J . A microfluidic-based device for study of transendothelial invasion of tumor aggregates in realtime. Lab on a Chip 2012; 12: 2837–2842.
Roberts SA, Waziri AE, Agrawal N . Development of a single-cell migration and extravasation platform through selective surface modification. Analytical Chemistry 2016; 88: 2770–2776.
Jeon JS, Zervantonakis IK, Chung S et al. In vitro model of tumor cell extravasation. PLoS ONE 2013; 8: e56910.
Chen MB, Whisler JA, Jeon JS et al. Mechanisms of tumor cell extravasation in an in vitro microvascular network platform. Integrative Biology 2013; 5: 1262.
Chen MB, Lamar JM, Li R et al. Elucidation of the roles of tumor integrin β1 in the extravasation stage of the metastasis cascade. Cancer Research 2016; 76: 2513–2524.
Sobrino A, Phan DTT, Datta R et al. 3D microtumors in vitro supported by perfused vascular networks. Scientific Reports 2016; 6: 31589.
Riahi R, Yang YL, Kim H et al. A microfluidic model for organ-specific extravasation of circulating tumor cells. Biomicrofluidics 2014; 8: 24103.
Cui X, Guo W, Sun Y et al. A microfluidic device for isolation and characterization of transendothelial migrating cancer cells. Biomicrofluidics 2017; 11: 14105.
Bischel LL, Sung KE, Jiménez-Torres JA et al. The importance of being a lumen. FASEB Journal: Official Publication of the Federation of American Societies for Experimental Biology 2014; 28: 4583–4590.
Chung S, Sudo R, Mack PJ et al. Cell migration into scaffolds under co-culture conditions in a microfluidic platform. Lab on a Chip 2009; 9: 269–275.
Fiddes LK, Raz N, Srigunapalan S et al. A circular cross-section PDMS microfluidics system for replication of cardiovascular flow conditions. Biomaterials 2010; 31: 3459–3464.
Hsu Y-H, Moya ML, Hughes CCW et al. A microfluidic platform for generating large-scale nearly identical human microphysiological vascularized tissue arrays. Lab on a Chip 2013; 13: 2990–2998.
Kim S, Lee H, Chung M et al. Engineering of functional, perfusable 3D microvascular networks on a chip. Lab on a Chip 2013; 13: 1489–1500.
Ballermann BJ, Dardik A, Eng E et al. Shear stress and the endothelium. Kidney international Supplement 1998; 67: S100–S108.
Skardal A, Shupe T, Atala A . Organoid-on-a-chip and body-on-a-chip systems for drug screening and disease modeling. Drug Discovery Today 2016; 21: 1399–1411.
Kashaninejad N, Nikmaneshi M, Moghadas H et al. Organ-tumor-on-a-chip for chemosensitivity assay: a critical review. Micromachines 2016; 7: 130.
Lee SH, Ha SK, Choi I et al. Microtechnology-based organ systems and whole-body models for drug screening. Biotechnology Journal 2016; 11: 746–756.
Wang X, Phan DTT, Sobrino A et al. Engineering anastomosis between living capillary networks and endothelial cell-lined microfluidic channels. Lab on a Chip 2016; 16: 282–290.
Bersini S, Jeon JS, Dubini G et al. A microfluidic 3D in vitro model for specificity of breast cancer metastasis to bone. Biomaterials 2014; 35: 2454–2461.
Jeon JS, Bersini S, Gilardi M et al. Human 3D vascularized organotypic microfluidic assays to study breast cancer cell extravasation. Proceedings of the National Academy of Sciences of the United States of America 2015; 112: 214–219.
Xu H, Li Z, Yu Y et al. A dynamic in vivo-like organotypic blood-brain barrier model to probe metastatic brain tumors. Scientific Reports 2016; 6: 36670.
Ma Y, Pan J-Z, Zhao S-P et al. Microdroplet chain array for cell migration assays. Lab on a Chip 2016; 16: 4658–4665.
Xu Z, Li E, Guo Z et al. Design and construction of a multi-organ microfluidic chip mimicking the in vivo microenvironment of lung cancer metastasis. ACS Applied Materials & Interfaces 2016; 8: 25840–25847.
Wang L, Wang Z, Liu X et al. High-level C-X-C chemokine receptor type 4 expression correlates with brain-specific metastasis following complete resection of non-small cell lung cancer. Oncology Letters 2014; 7: 1871–1876.
Feeley BT, Liu NQ, Conduah AH et al. Mixed metastatic lung cancer lesions in bone are inhibited by noggin overexpression and Rank:Fc administration. Journal of Bone and Mineral Research: the Official Journal of the American Society for Bone and Mineral Research 2006; 21: 1571–1580.
Zhang H, Yang N, Sun B et al. CD133 positive cells isolated from A549 cell line exhibited high liver metastatic potential. Neoplasma 2014; 61: 153–160.
Mo M, Eskin SG, Schilling WP . Flow-induced changes in Ca2+ signaling of vascular endothelial cells: effect of shear stress and ATP. The American Journal of Physiology 1991; 260: H1698–H1707.
Chiu J-J, Chien S . Effects of disturbed flow on vascular endothelium: pathophysiological basis and clinical perspectives. Physiological Reviews 2011; 91: 327–387.
Davies PF . Flow-mediated endothelial mechanotransduction. Physiological Reviews 1995; 75: 519–560.
Regmi S, Fu A, Luo KQ . High shear stresses under exercise condition destroy circulating tumor cells in a microfluidic system. Scientific Reports 2017; 7: 39975.
Zaman MH, Trapani LM, Sieminski AL et al. Migration of tumor cells in 3D matrices is governed by matrix stiffness along with cell-matrix adhesion and proteolysis. Proceedings of the National Academy of Sciences of the United States of America 2006; 103: 10889–10894.
Patil P, Madhuprasad M, Kumeria T et al. Isolation of circulating tumour cells by physical means in a microfluidic device: a review. RSC Adv 2015; 5: 89745–89762.
Khalil DN, Smith EL, Brentjens RJ et al. The future of cancer treatment: immunomodulation, CARs and combination immunotherapy. Nature Reviews Clinical Oncology 2016; 13: 273–290.
Yoshida Y, Yamanaka S . Recent stem cell advances: induced pluripotent stem cells for disease modeling and stem cell-based regeneration. Circulation 2010; 122: 80–87.
Vladisavljević GT, Khalid N, Neves MA et al. Industrial lab-on-a-chip: design, applications and scale-up for drug discovery and delivery. Advanced Drug Delivery Reviews 2013; 65: 1626–1663.
Acknowledgements
We acknowledge financial support by the Natural Sciences and Engineering Research Council of Canada via Discovery Grants to LDY and YS and by the Canada Research Chairs Program.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ma, YH., Middleton, K., You, L. et al. A review of microfluidic approaches for investigating cancer extravasation during metastasis. Microsyst Nanoeng 4, 17104 (2018). https://doi.org/10.1038/micronano.2017.104
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/micronano.2017.104
This article is cited by
-
Micro-engineering and nano-engineering approaches to investigate tumour ecosystems
Nature Reviews Cancer (2023)
-
Microfluidics engineering towards personalized oncology—a review
In vitro models (2023)
-
Recent advances in cancer-on-a-chip tissue models to dissect the tumour microenvironment
Med-X (2023)
-
Recent advances for cancer detection and treatment by microfluidic technology, review and update
Biological Procedures Online (2022)
-
Towards Green 3D-Microfabrication of Bio-MEMS Devices Using ADEX Dry Film Photoresists
International Journal of Precision Engineering and Manufacturing-Green Technology (2022)