Abstract
There is increasing evidence that, in addition to their presence, the propensity of circulating tumour cells to form multi-cellular clusters bears significant information about both cellular resistance to chemotherapy and overall prognosis. We present a novel two-stage, stopped-flow, continuous centrifugal sedimentation strategy to measure the size distributions of events (defined here as cells or clusters thereof) in a blood sample. After off-chip removal of red blood cells, healthy white blood cells are sequestered by negative-immunocapture. The purified events are then resolved along a radially inclined rail featuring a series of gaps with increasing width, each connected to a designated outer collection bin. The isolation of candidate events independent of target-specific epitopes is successfully demonstrated for HL60 (EpCAM positive) and sk-mel28 (EpCAM negative) cells using identical protocols and reagents. The propensity to form clusters was quantified for a number of cell lines, showing a negligible, moderate or elevated tendency towards cluster formation. We show that the occupancy distribution of the collection bins closely correlates with the range of cluster sizes intrinsic to the specific cell line.
Similar content being viewed by others
Introduction
The presence of circulating tumour cells (CTCs) in the blood of a patient, as found in most forms of epithelial cancer1–5, has been identified as a reliable indicator of the prognosis of various cancer types. More recently, the propensity of CTCs to occur in multi-cellular clusters (sometimes referred to as circulating tumour microemboli or CTM) has been observed by a number of groups using a number of technologies. On examination, the occurrence of such clusters can provide important information about patients’ outcomes, as these clusters appear to increase the risk of metastasis6,7. The associated surge in metastatic potential is dramatically increased, by up to 50-fold, when the outcome of mice that have been injected with either single CTCs or CTC clusters is observed6,8. The biological processes that result in CTC clusters having a higher metastatic potential than single CTCs do are still under investigation; nevertheless, studies have provided interesting results suggesting that the microenvironment generated by the multi-cellular nature of the clusters provides certain protection from physiological processes that normally compromise the viability of single CTCs. As the number of single CTCs is far in excess of the number of metastatic lesions that develop in a patient, it can be inferred that the majority of CTCs are either inert (from a metastatic perspective) or do not survive long enough to seed a new lesion. Most CTCs are probably lost through anoikis (a form of apoptosis in which cells lose cell–cell or cell–matrix interactions) or stresses induced by shear forces in the blood. However, CTCs within a cluster have been shown to resist anoikis/apoptosis, possibly due to the maintenance of cell–cell junctions within the cluster and protection from shear forces by virtue of the outer cells7,9,10. Furthermore, molecular investigations of the cells within a cluster have shown elevated levels of pro-survival9 and pro-epithelial-to-mesenchymal transition signals11. These clusters’ genesis is still under investigation, but studies have proposed that rather than emerging from the proliferation of a single CTC or from aggregation in the bloodstream, clusters may represent tissue that breaks away from the site of a primary tumour6,7. CTCs’ lack of proliferation therefore (theoretically) renders these cells resistant to chemotherapies that actively target rapidly dividing cells, thus increasing the diagnostic and prognostic potential of identifying the cluster load in a blood sample.
A number of microfluidic systems for isolating candidate CTC events have been developed, as reviewed by Yu et al. 12. In these systems, various strategies have been employed and combined to isolate CTCs from blood, including epitope-specific capture13–15, epitope-specific magnetic bead binding16–19 and physical isolation based on size20,21 and/or deformability22,23. The majority of microfluidic lab-on-a-chip systems in which clusters have been isolated seem to have been originally designed for the isolation of single CTCs. However, by virtue of fortuitously and efficiently designed tolerance to size, CTC clusters have sometimes been co-isolated along with the single events14,24. The size ranges of clusters retrieved in these systems were elucidated by scanning the capture regions of the chips microscopically and reporting the cell count in each captured cluster on an individual basis. The HB-chip developed by Stott et al.14 was recently advanced to allow more efficient isolation of CTC events, and concordantly, the geometry of this new CTC-iChip was chosen to capture clusters in addition to single-cell events. However, individual optical identification of clusters was still required to quantify the clusters’size25.
To the best of our knowledge, the current work is the first to present a proof-of-concept strategy for directly estimating the size range of CTC clusters in a blood sample without the need to serially examine and enumerate each individual event. Blood samples are initially processed off-chip using standard laboratory protocols to remove red blood cells (RBCs), and negative-mode isolation with specifically functionalised magnetic beads is used to remove non-CTC candidate events. Negative-mode isolation25–27 is chosen to circumvent the issues that arise due to the inherent phenotypic heterogeneity of these events when directly targeting CTC candidates. Using stopped-flow centrifugal particle sedimentation, CTC events (single- and multi-cellular) are binned into discrete collection chambers that are individually connected to a differential size-filtration structure along a rail. As cellular clusters are absent in healthy blood, qualitative analysis of the collection chambers correlates with the frequencies of cluster sizes present in the original sample that was loaded into the cartridge. Simple digital interpretation of the occupancy of the collection bins (i.e., assigning “1” to an occupied bin and “0” to an empty bin) allows quantification of the CTC cluster distribution.
Operational principle
Analysis of a sample is arranged in two stages: initial off-chip blood processing, followed by an on-chip differential size-sorting step. After hypotonic RBC lysis using standard protocols, CTC events are purified through bead-based, immuno-magnetic removal of normal, non-CTC candidate cells. This negative mode was chosen to offset the inherent heterogeneity within CTC populations due to the lack of known epitopes that are common to all CTCs. For example, EpCAM-expressing cells in the blood are commonly used in positive isolation strategies for the identification of CTCs. However, certain CTCs do not express EpCAM, and indeed, in many carcinomas, EpCAM is under expressed or even absent28.
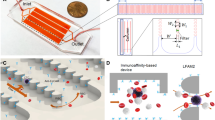
For the negative-mode isolation used here, white blood cells (WBCs) are incubated with a mixture of CD15- and CD45-specific super-paramagnetic beads, which bind to all naturally occurring WBCs. Abnormal events (CTC or CTC-based cluster candidates) lack expression of CD15 and CD45 and therefore remain unbound. Healthy WBCs are then gently eliminated from the solution by placing the tube in proximity to a magnet (Supplementary Figure S1). The small numbers of abnormal cellular events that remain suspended are forwarded into the second phase. The entire, dead-end microfluidic cell-sorting chamber is first fluidic ally primed so that the system operates in stopped-flow sedimentation mode. The isolated abnormal events are introduced via the loading port. Next, the disc is spun at 12 Hz for 25 min. Upon exiting the sample-loading chamber, events are guided along the size-exclusion rail, which is inclined with respect to the direction of the centrifugal field (Figure 1a, green box). This rail is composed of flat-edged pillars featuring interspersed gaps of successively increasing size that gate the entry. The paths to the outer bins are aligned parallel to the radial direction so that events penetrating the rail centrifugally sediment straight to the base. The gate size successively increases from 4.7 μm in the initial bin (which isolates debris and highly deformable cells) to 130 μm in the last of the eight bins, as defined in Figure 1a. The aperture sizes of the rails were inspired by the size distribution of native blood cells in relation to cancer cells. For example, it is anticipated that the fragments of RBCs that are present following hypotonic lysis as well as platelets (2–3 µm) would resolve to the first bin, i.e., the smallest aperture bin. Any remaining WBCs (9–15 µm) would then resolve to the second bin. Thereafter, cancer cells and multi-cellular events will occupy the subsequent bins (bins 3–8). Events that resolve to the base of the collection bins can be observed by microscopy.
Description of the centrifugal size-exclusion rail. (a) A single full chamber is schematically represented, highlighting the operational characteristics and features of interest. The green box shows a magnified section of the critical size-exclusion rail gating entry into two of the underlying bins, with the direction of a particle travelling along the rail shown with green hatched arrows and the centrifugal field shown with blue and yellow solid arrows. The sizes of the apertures gating entry into each of the bins are listed. The purple box shows a magnified section of the particle collection bins, with particles of increasing size represented as blue, yellow and red circles. (b) Operation of the size-exclusion rail. (1) Negative selection of abnormal cells proceeds off-disc in a disposable sample tube. (2) Isolated abnormal cells are loaded in equal aliquots between eight distinct chambers, marked here as A–H. The direction and centre of rotation of the disc are shown with a white arrow. (3) At a spin rate of 12 Hz, cellular events travel from the sample-loading reservoir to the size-exclusion rail. Small cells and debris (blue) enter the first bin, but the diameter of larger events will bar them from bin #1, and these will continue along the rail until a sufficiently wide aperture is encountered that will allow passage of the events into the underlying bin. Single cells are shown in green (i), small clusters are shown in yellow (ii) and larger clusters are shown in red (iii). For clarity, only four chambers are represented.
Arrays of shaped pillars have been previously used in the microfluidics literature to separate materials, using either a filtering strategy based on gap sizes29 or the so-called deterministic lateral displacement approach, in which material ranging from DNA molecules30 to beads31 to cells32 has been successfully separated. However, to the best of our current knowledge, the strategy presented here is the first to combine shaped pillars and a centrifugal sedimentation mode (i.e., in the absence of flow) to specifically generate data to measure the event-size distribution in a cellular sample via a collective approach, i.e., without the requirement of serial testing of individual captured events.
Materials and methods
Standard protocols for disc fabrication, polydimethylsiloxane priming, blood processing and cell culture were used and have been described elsewhere33–37. These protocols are also briefly outlined in the Supplementary Information.
DynabeadTM CD15/CD45-mediated negative selection of normal WBCs
Negative selection-based removal of normal WBCs was carried out on spiked and non-spiked WBC samples that were prepared as described in the “Materials and Methods” section of the ESI. Following the removal of RBCs, each WBC sample was placed into a 15-ml disposable centrifuge tube (Sarstedt, Nümbrecht, Germany), and the volume was adjusted to 14 ml using priming buffer. In total, 16 and 80 µl of CD15- and CD45-specific magnetic Dynabeads® (Life Technologies, Carlsbad, CA, USA), respectively, were added to the tube. A double layer of Parafilm (Pechiney Plastic Packaging Company, Chicago, IL, USA) was placed between the cap and the tube, which was then rotated on a SB2 Rotator (Stuart, Staffordshire, UK) at 20 rpm for 10 min at an angle of 15° from the vertical at room temperature.
After incubation, the bead-bound WBCs were immobilized by placing the tube horizontally onto a flatbed rocker (Stuart, UK) operated at 45 Hz. A strong N45 NdFeB permanent magnet (30 × 30 × 15 mm3, Supermagnete, Gottmadingen, Germany) was slowly lowered from above the tube to gently move the magnetic beads/cells to the tube wall (Supplementary Figure S1). Critically, this step was performed with a slow approach of the magnet to minimize undesirable capture of abnormal cells during the migration of the magnetic events. The magnet was initially placed approximately 10 cm above the tube and was then lowered at such a speed that it made contact with the tube after approximately 1 min. The rocking motion of the flatbed rocker maintained the lateral movement of the non-magnetic cells, thus minimizing inappropriate capture. When in contact with the tube, the magnet was held in place for 30 s to ensure maximum immobilization of the magnetic events; thereafter, the liquid phase of the sample was transferred to a fresh 50-ml tube. Abnormal cells that may have been caught in the bead pack were recovered by resuspension of the beads in 14 ml of priming buffer and two re-isolation steps, as described. The liquid phases of all three isolations were merged, and candidate events were purified by centrifugation at 200× g for 5 min. Any resolved pellet was resuspended in 32 µl of priming buffer.
A 4-µl volume of resuspended pellet was loaded into each of the microfluidic chambers, and discs were spun on a custom-built test stand composed of a Cool Muscle CM2-X-56B10A motor (Muscle Corp., Osaka, Japan) with a custom chuck that was machined to securely attach discs to the servo-motor shaft. A desktop PC (Dell, Round Rock, TX, USA) was used to control the spin speed of the motor. Experimental discs were spun at 12 Hz in the anti-clockwise direction for 40 min.
Microscopic imaging and analysis of bin occupancy
Following completion of a spincycle, the occupancy of the collection bins was visualized by differential interference contrast (DIC) and/or epifluorescence imaging using an inverted epifluorescence microscope (Olympus IX81, Tokyo, Japan). Where experiments using beads are shown, the distribution was measured using ImageJ (NIH, Bethesda, MD, USA) by drawing a multi-point region of interest (ROI) to surround the area occupied by the beads at the base of the bins. The internal area of this ROI was then calculated and compared across the eight bins. Where shown, real-time video of a spinning disc was created from a series of still images. The polystyrene beads used in the images and video were supplied pre-coloured from the manufacturer (Microparticles GmbH, Berlin, Germany).
In cases where cells were stained with NucBlue, the relative fluorescent signal across the bins was calculated as described elsewhere36 and as also briefly outlined in the Supplementary information. For qualitative analysis in which only the binary state of occupancy was required, the presence of events in each bin was visually observed using DIC microscopy.
Experimental results
Size-based binning of beads using the size-exclusion rail
To demonstrate the feasibility of using the size-exclusion rail to correctly assign events of particular sizes to appropriate bins, polystyrene bead samples of increasing size were processed, and the bins to which the beads resolved were analysed. Four bead diameters (5, 20, 35 and 50 µm) were chosen to represent the expected size range of cellular clusters and also to test the aperture widths gating the collection bins of the rail. These beads were introduced into the loading chamber as homogeneous samples and were processed by centrifugation, as described in the “Materials and Methods” section.
Following the resolution of the beads to the capture bins, the distribution of the various bead sizes was investigated. In general, the majority of a homogeneous bead population resolved to the bin whose gate most closely matched the bead size under analysis (Figure 2a). For example, 87% of the 20-µm beads resolved to the third bin, accessible via an aperture of 22 µm, and 79% of the 50-µm beads arrived in the sixth bin after passing through the 50.6-µm gap in the rail (Figure 2b). Although it would be expected that the gate aperture must exceed the bead diameter, certain beads managed to penetrate gates that were set slightly lower than the bead size. This was observed, for example, in the case of the sedimentation of 97.1% of the 5-µm beads in the first bin, which possessed a gate aperture of only 4.7 µm. This finding is most probably attributable to slight manufacturing tolerance of the posts that composed the rail in this region. For example, the reported aperture for the first bin was 4.7 µm based on the original CAD drawing, in which this value was specified. We chose to report this value for consistency. When the aperture was measured using scanning electron microscopy, the actual measurement for the fabricated structured was 4 ± 2 µm, thus allowing a large number of the particles to penetrate the rail at this point. Resolution of the 5-μm beads to the first bin may also have been due to non-uniformity in the shape of the beads. For example, certain beads may not have been perfectly spherical and instead may have had a more ovular cross-section. Nevertheless, despite a degree of sedimentation at locations ±1 bin from the expected destination, the critical size-exclusion rail successfully resolved beads of specific sizes to distinct collection locations. When a heterogeneous sample composed of 5- and 35-µm beads was processed, both bead species were directed to the same bins as in the experiments with monodisperse bead distributions. However, it was observed that the large beads apparently carried several of the 5-µm beads towards the 35-µm bin, possibly due to binding of the polystyrene (Figure 2c). This caveat slightly compromised experiments with more poly-disperse populations, but as the beads were merely used as proof of concept, this is not considered to be a restriction of the device for cell-based applications. A video of the binning of beads is available in the Supplementary information.
Determination of the distribution of polystyrene beads using the size-exclusion rail. (a) Images of the eight capture bins when homogeneous bead populations of defined size were processed. The area outlined with the blue hatched box is shown. Images representing bead diameters of 5 µm (blue), 20 µm (yellow), 35 µm (red) and 50 µm (purple) are shown. (b) Bar chart showing the relative distribution of beads following processing on a size-exclusion rail. The maximum of the scale is 100%. Gate apertures and bin numbers are shown below the chart. Error bars represent one standard deviation. The data were collected from a total of three chambers run on three separate discs. (c) Image from a real-time experiment in which a heterogeneous sample composed of 5-µm and 35-µm beads was processed. The 5-µm beads are shown entering bin #1, whereas the 35-µm beads continue along the rail. All scale bars are 500 µm.
Examination of the clustering potential inherent in specific cultured cells
As discussed, the size-exclusion rail was designed for the quantification of the range and extent of cellular clustering in a blood sample. To this end, we first validated the basic functionality of the rail by spiking WBC samples with three cell lines, representing single cells (HL60, sourced from the Deutsche Sammlung von Mikroorganismen und Zellkulturen [DSMZ], Braunschweig, Germany) and cells of low (colo794) or high (sk-mel28) cluster size range (both kind gifts from Prof. J. O’Leary, Trinity College Dublin, Ireland; Figure 3). Although so far only the overall fraction of all clusters compared with single cells has been reported, the current work uniquely quantifies the precise cluster size distributions for these cell lines.
Detailed cluster size classification of the experimental cultured cell lines. (a) Examples of cellular events scored as cluster sizes ranging from 1 to 8 cells or more cells during the cluster size analysis. The scale bar is 20 µm. (b) Images taken of cell populations for cluster size analysis for HL60 (single cells), colo794 (low clustering) and sk-mel28 (high clustering). Scale bars are 100 µm. (c) Bar charts representing the observed range and distribution of cluster sizes for the three cell lines. Error bars represent one standard deviation. The data were collected for three separate passages of cells.
To examine the capability of the size-exclusion rail to profile the cluster size from a starting sample of whole blood, individual aliquots of blood were spiked with homogeneous populations of NucBlue-stained cell lines (Figure 4). For these experiments, whole blood samples of 1-ml volume were spiked with 1 × 103 events. RBCs were lysed, and CD15/CD45-positive cells were removed by negative magnetic isolation, as described in the “Materials and Methods” section. The isolated abnormal events were then loaded into the chambers of the experimental disc and processed by centrifugation. On the completion of the sample processing, the fluorescence signal from the spiked cells was measured in each collection bin.
Distribution of cellular events according to cluster size fora starting sample of spiked whole blood. (a) Fluorescence data (black lines) showing the relative fluorescent signal detected in each bin (as a percentage of the total signal detected over the eight bins) per cell line. These data are overlaid on the bar charts to display the range and distribution of the expected cellular clustering of the three cell lines, as originally shown in Figure 3. The correlation coefficient between the distributions of fluorescence and cluster size is presented on each graph. Error bars represent a single standard deviation in the fluorescence distribution and were measured over a total of four individual chambers run on separate discs. Error bars are shown only for the fluorescence data, so as not to overlap with those of the bar charts; error bars for the bar charts are shown in Figure 3c. (b) Examples of fluorescent and merged DIC/DAPI images of the capture bins following processing of blood spiked with fluorescently stained cultured cells. For clarity, fluorescent signals are assigned a blue colour in the merged images. Magnified images of the bins of interest are shown in the hatched boxes. Scale bars are 500 µm. In both (a) and (b), data pertaining to HL60, colo794 and sk-mel28 are labelled green, orange and red, respectively.
Supplementary Figure S2 shows that when un-spiked and healthy blood is processed, certain material resolves to bin #1. As platelets express neither CD15 nor CD45, much of this background material is assumed to represent a combination of platelets and debris resulting from the lysis of the RBCs. However, as it is present in all samples tested but always resolves to bin #1, this material does not interfere with the search for larger cellular events in the higher bins. When blood is spiked with single-cell HL60, these cells are also nearly entirely diverted to bin #1 (Figure 4a and 4b, green labels). This finding is interesting, as the gate size for bin #1 is 4.7 µm, whereas an average size of 12.4 ± 1.2 µm has been reported for HL60 cells38. This passage of the larger cells may be linked to the fact that living cells, and HL60 cells in particular39,40, are known to deform under certain circumstances. Therefore, it is likely that the centrifugal force suffices to squeeze the HL60 cells through the 4.7-µm aperture. Supplementary Figure S3 displays an image of an HL60 dimer after passage through the gate of bin #2. It can be observed that the width of the aperture is narrower than either the width or the length of the HL60 dimer, suggesting that the dimer deformed to penetrate the rail. Although interesting in terms of deformability, a potential oncogenic marker in and of itself41, the detection of multi-cellular clusters, as the primary objective of our work, was not compromised by the resolution of the individual HL60 cells to bin #1.
Samples were spiked with either the low-range-clustering colo794 cells or the high-range-clustering sk-mel28 cells; both cell lines were directed to the higher level bins in addition to bin #1 (Figure 4a and 4b, orange and red labels). Analysis of the distribution of the fluorescence signal in these bins (Figure 4a, black lines) revealed that the colo794 cells largely occupied bins #1–2, with a degree of low occupancy of bin #3. Notably, the sk-mel28 cells exhibited a fluorescence distribution profile that spanned from bins #1–8, with the highest peaks for the first and last bins, thus reflecting the cells’ wider distribution of cluster sizes. When the observed cluster size data for each cell line are overlaid on the fluorescence distribution data across the capture bins of the size-exclusion chamber (Figure 4a), a close correlation between the expected cluster size and bin occupancy is found. HL60 shows a near perfect correlation (ρ = 0.9998), as was expected given the largely single-cell nature of the cell line. Furthermore, colo794 (ρ = 0.99) and sk-mel28 (ρ = 0.95) are strongly correlated, thus demonstrating that the size-exclusion rail can be used to assess the tendency towards clustering within a sample based on the occupancy distribution along the capture bins.
Analysis method for showing the cellular clustering potential in a sample
Although the data in Figure 4 are based on the measurement and processing of fluorescence in the collection bins, they are shown primarily as an example of the performance of the size-exclusion rail. One of the design aims of binning events to discrete bins was to facilitate simple interpretation of the data, without the need for fluorescence or similar specialized microscopy. To this end, inspection of each bin using a simple bright field microscope (i.e., even without fluorescent light sources or imagers) allows rapid occupancy scoring of each. As detectable clustering events are absent in (un-spiked) healthy blood (Supplementary Figure S2), it is likely that any occupation of bins to which clusters would resolve is of interest for an experimental sample. As the candidate events from a sample are distributed across all disc-based chambers (A–H) and given that each chamber contains eight collection bins, when the occupancy profile of the sample is displayed, a two-dimensional 8 × 8 array will result. To facilitate graphical interpretation, locations in the array representing void and occupied bins are coloured green and red, respectively (Figure 5a). Viewing such an array provides an immediate and easily interpreted indication of the range of cluster sizes harboured in the sample. For example, in the case that the array is overwhelmingly green, with certain areas of red on the right, the array suggests that there were few events larger than 2–4 cells (i.e., colo794 in Figure 5a). Conversely, a nearly entirely red array suggests extensive clustering (i.e., sk-mel28 in Figure 5a).
Presentation of clustering data using digital “void”/“occupied” visual inspection of the collection bins. (a) The data are arranged in a two-dimensional 8 × 8 array representing chamber identification (A–H) and bin identification (1–8). For each sample, a green box indicates a chamber that was identified as void, and a red box indicates a chamber that was occupied. (b) Data from (a) were further processed to generate a one-dimensional profile of the occupancy probabilities of the eight discrete bin addresses across all of the chambers (A–H). A colour scheme is applied to the data to facilitate rapid visual communication, which allows comparison of the clustering ranges inherent to each sample. The colour gradient goes from white (case: all bins are void) to red (case: all bins are occupied).
These data can be further processed from the two-dimensional array to an one-dimensional profile, if desired. By assigning a numerical value of 0 to void bins and a value of 1 to occupied bins, the representative value for each bin address (1–8) can be calculated by reporting the mean value of each specific bin address across all of the chambers used. This concept is shown as the “Event Distribution” in Figure 5b and is sufficient to visualize the relative load of multi-cellular clusters present in the original blood sample. Rather than displaying the specific distribution of cluster sizes, such a single-figure compression of data mirrors the overall tendency of cells to form clusters of various sizes in the examined sample and further allows easy comparison of blood samples (Figure 5b).
Summary and outlook
Recent literature has provided strong evidence that the cluster size distribution in an oncological sample represents a marker of high diagnostic value. In this work, we developed a novel centrifugal size-exclusion rail scheme for the examination of the cluster load in a blood sample, and we demonstrated that samples harbouring single-cell candidate events (or no candidate events at all) will exclusively resolve to a given collection bin. When spiking cancer cells from cell lines with a known propensity to cluster, their respective tendencies to form clusters are reflected, with high correlation, by the occupancy distribution across the bins.
We acknowledge that the concentration of target cellular events that we used in the spiked samples is in excess of that expected in a real liquid biopsy from a patient; however, our aim was to present a proof-of-concept strategy for specifically utilizing a size-exclusion rail to detect and characterize the cluster load in a blood sample. Continuing work on the size-exclusion rail is focused on integrating recovery technology developed by our group to allow arraying and removal of cells/clusters of interest for specific downstream molecular analysis (these technologies are detailed in Ref 42 and Ref 43) and also on using the rail to analyse patient samples, with a particular focus on increasing the limit of detection for clinical deployment. Footnote 1
Notes
Supplementary information for this article can be found on the Microsystems & Nanoengineering website (http://www.nature.com/micronano).
References
Hou J, Krebs M, Ward T et al. Circulating tumor cells, enumeration and beyond. Cancers 2010; 2: 1236–1250.
Cohen SJ, Punt CJ, Iannotti N et al. Relationship of circulating tumor cells to tumor response, progression-free survival, and overall survival in patients with metastatic colorectal cancer. Journal of Clinical Oncology 2008; 26: 3213–3221.
de Bono JS, Scher HI, Montgomery RB et al. Circulating tumor cells predict survival benefit from treatment in metastatic castration-resistant prostate cancer. Clinical Cancer Research 2008; 14: 6302–6309.
Mateo J, Gerlinger M, Rodrigues D et al. The promise of circulating tumor cell analysis in cancer management. Genome Biology 2014; 15: 448.
Alix-Panabières C, Pantel K. Circulating tumor cells: Liquid biopsy of cancer. Clinical Chemistry 2013; 59: 110–118.
Aceto N, Bardia A, Miyamoto DT et al. Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell 2014; 158: 1110–1122.
Hou JM, Krebs MG, Lancashire L et al. Clinical significance and molecular characteristics of circulating tumor cells and circulating tumor microemboli in patients with small-cell lung cancer. Journal of Clinical Oncology 2012; 30: 525–532.
Liotta LA, Kleinerman J, Saldel GM. The significance of hematogenous tumor cell clumps in the metastatic process. Cancer Research 1976; 36: 889–894.
Duda DG, Duyverman AM, Kohno M et al. Malignant cells facilitate lung metastasis by bringing their own soil. Proc Natl Acad Sci U S A 2010; 107: 21677–21682.
Robson EJ, Khaled WT, Abell K et al. Epithelial-to-mesenchymal transition confers resistance to apoptosis in three murine mammary epithelial cell lines. Differentiation 2006; 74: 254–264.
Yu M, Bardia A, Wittner BS et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 2013; 339: 580–584.
Yu M, Stott S, Toner M et al. Circulating tumor cells: Approaches to isolation and characterization. Journal of Cell Biology 2011; 192: 373–382.
Nagrath S, Sequist LV, Maheswaran S et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 2007; 450: 1235–1239.
Stott SL, Hsu CH, Tsukrov DI et al. Isolation of circulating tumor cells using a microvortex-generating herringbone-chip. Proceedings of the National Academy of Sciences of the United States of America 2010; 107: 18392–18397.
Dharmasiri U, Njoroge SK, Witek MA et al. High-throughput selection, enumeration, electrokinetic manipulation, and molecular profiling of low-abundance circulating tumor cells using a microfluidic system. Analytical Chemistry 2011; 83: 2301–2309.
Kang JH, Krause S, Tobin H et al. A combined micromagnetic-microfluidic device for rapid capture and culture of rare circulating tumor cells. Lab on a Chip 2012; 12: 2175–2181.
Kirby D, Glynn M, Kijanka G et al. Rapid and cost-efficient enumeration of rare cancer cells from whole blood by low-loss centrifuge-magnetophoretic purification under stopped-flow conditions. Cytometry Part A 2014; 87: 74–80.
Lien K, Chuang Y, Hung L et al. Rapid isolation and detection of cancer cells by utilizing integrated microfluidic systems. Lab on a Chip 2010; 10: 2875–2886.
Thege FI, Lannin TB, Saha TN et al. Microfluidic immunocapture of circulating pancreatic cells using parallel EpCAM and MUC1 capture: Characterization, optimization and downstream analysis. Lab on a Chip 2014; 14: 1775–1784.
Zheng S, Lin HK, Lu B et al. 3D microfilter device for viable circulating tumor cell (CTC) enrichment from blood. Biomedical Microdevices 2011; 13: 203–213.
Hur SC, Mach AJ, Di Carlo D. High-throughput size-based rare cell enrichment using microscale vortices. Biomicrofluidics 2011; 5: 022206-1–022206-10.
Tan SJ, Yobas L, Lee GYH et al. Microdevice for the isolation and enumeration of cancer cells from blood. Biomedical Microdevices 2009; 11: 883–892.
Hur SC, Henderson-MacLennan NK, McCabe ER et al. Deformability-based cell classification and enrichment using inertial microfluidics. Lab on a Chip 2011; 11: 912–920.
Hou J, Krebs M, Ward T et al. Circulating tumor cells as a window on metastasis biology in lung cancer. American Journal of Pathology 2011; 178: 989–996.
Ozkumur E, Shah AM, Ciciliano JC et al. Inertial focusing for tumor antigen-dependent and -independent sorting of rare circulating tumor cells. Science Translational Medicine 2013; 5: 179ra47.
Balasubramanian P, Yang L, Lang JC et al. Confocal images of circulating tumor cells obtained using a methodology and technology that removes normal cells. Molecular Pharmaceutics 2009; 6: 1402–1408.
Tkaczuk KHR, Goloubeva O, Tait NS et al. The significance of circulating epithelial cells in breast cancer patients by a novel negative selection method. Breast Cancer Research and Treatment 2008; 111: 355–364.
Kim M, Choi NY, Lee EK et al. Identification of novel markers that outperform EpCAM in quantifying circulating tumor cells. Cellular Oncology 2014; 37: 235–243.
Sochol RD, Corbett D, Hesse S et al. Dual-mode hydrodynamic railing and arraying of microparticles for multi-stage signal detection in continuous flow biochemical microprocessors. Lab on a Chip 2014; 14: 1405–1409.
Huang LR, Cox EC, Austin RH et al. Tilted Brownian ratchet for DNA analysis. Analytical Chemistry 2003; 75: 6963–6967.
Huang LR, Cox EC, Austin RH et al. Continuous particle separation through deterministic lateral displacement. Science 2004; 304: 987–990.
Green JV, Radisic M, Murthy SK. Deterministic lateral displacement as a means to enrich large cells for tissue engineering. Analytical Chemistry 2009; 81: 9178–9182.
Kirby D, Siegrist J, Kijanka G et al. Centrifugo-magnetophoretic particle separation. Microfluidics and Nanofluidics 2012; 13: 899–908.
Steigert J, Haeberle S, Brenner T et al. Rapid prototyping of microfluidic chips in COC. Journal of Micromechanics and Microengineering 2007; 17: 333–341.
Glynn MT, Kinahan D, Ducree J. Rapid, low-cost and instrument-free CD4 cell counting for HIV diagnostics in resource-poor settings. Lab on a Chip 2014; 14: 2844–2851.
Glynn M, Kirby D, Chung D et al. Centrifugo-magnetophoretic purification of CD4+ cells from whole blood toward future HIV/AIDS point-of-care applications. J Lab Autom 2013; 19: 285–296.
Hosokawa K, Sato K, Ichikawa N et al. Power-free poly (dimethylsiloxane) microfluidic devices for gold nanoparticle-based DNA analysis. Lab on a Chip 2004; 4: 181–185.
Rosenbluth MJ, Lam WA, Fletcher DA. Force microscopy of nonadherent cells: A comparison of leukemia cell deformability. Biophysical Journal 2006; 90: 2994–3003.
Tsai MA, Waugh RE, Keng PC. Cell cycle-dependence of HL-60 cell deformability. Biophysical Journal 1996; 70: 2023–2029.
Hallows KR, Frank RS. Changes in mechanical properties with DMSO-induced differentiation of HL-60 cells. Biorheology 1992; 29: 295–309.
Swaminathan V, Mythreye K, O’Brien ET et al. Mechanical stiffness grades metastatic potential in patient tumor cells and in cancer cell lines. Cancer Research 2011; 71: 5075–5080.
Burger R, Reith P, Kijanka G et al. Array-based capture, distribution, counting and multiplexed assaying of beads on a centrifugal microfluidic platform. Lab on a Chip 2012; 12: 1289–1295.
Burger R, Kurzbuch D, Gorkin R et al. An integrated centrifugo-opto-microfluidic platform for arraying, analysis, identification and manipulation of individual cells. Lab on a Chip 2015; 15: 378–381.
Acknowledgements
This work was supported in part by Enterprise Ireland (Grant No. CF 2011 1317) and Science Foundation Ireland (Grant No. 10/CE/B1821). The authors would like to thank Prof. J. O’Leary for the kind gift of the colo794 and sk-mel28 cell lines.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 Unported License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0
About this article
Cite this article
Glynn, M., Nwankire, C., Lemass, K. et al. Cluster size distribution of cancer cells in blood using stopped-flow centrifugation along scale-matched gaps of a radially inclined rail. Microsyst Nanoeng 1, 15018 (2015). https://doi.org/10.1038/micronano.2015.18
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/micronano.2015.18
This article is cited by
-
Particle/cell separation on microfluidic platforms based on centrifugation effect: a review
Microfluidics and Nanofluidics (2017)