Abstract
The airway epithelium secretes proteins that function in innate defense against infection. Bactericidal/permeability-increasing fold-containing family member A1 (BPIFA1) is secreted into airways and has a protective role during bacterial infections, but it is not known whether it also has an antiviral role. To determine a role in host defense against influenza A virus (IAV) infection and to find the underlying defense mechanism, we developed transgenic mouse models that are deficient in BPIFA1 and used these, in combination with in vitro three-dimensional mouse tracheal epithelial cell (mTEC) cultures, to investigate its antiviral properties. We show that BPIFA1 has a significant role in mucosal defense against IAV infection. BPIFA1 secretion was highly modulated after IAV infection. Mice deficient in BPIFA1 lost more weight after infection, supported a higher viral load and virus reached the peripheral lung earlier, indicative of a defect in the control of infection. Further analysis using mTEC cultures showed that BPIFA1-deficient cells bound more virus particles, displayed increased nuclear import of IAV ribonucleoprotein complexes, and supported higher levels of viral replication. Our results identify a critical role of BPIFA1 in the initial phase of infection by inhibiting the binding and entry of IAV into airway epithelial cells.
Similar content being viewed by others
Introduction
The airway epithelium has a fundamental role in the initial defense against pathogens and as such secretes a number of proteins/peptides that function in innate defense.1 Bactericidal/permeability-increasing fold-containing family A1 (BPIFA1; also called SPLUNC1) is a glycoprotein that is highly expressed in the respiratory epithelium and submucosal glands of the upper airways in mice and humans.2, 3, 4, 5, 6, 7, 8, 9 The mouse and human BPIFA1 sequences are homologous.8, 10, 11 Previous studies have demonstrated that BPIFA1 acts as a surfactant,12 can regulate the amiloride-sensitive epithelial sodium channel, ENaC,13 and affects mucociliary clearance in the upper airways.14 It has also been shown to have anti-bacterial roles. For example, Bpifa1 expression is induced after Mycoplasma infection, enhancing interleukin-8 production and bacterial clearance.15 BPIFA1 also has a role in defense against Klebsiella pneumoniae,16 possibly acting through the modulation of macrophage function.17 Less is known about any antiviral role of BPIFA1, although our previous work has shown modulation of BPIFA1 levels after murine γ-herpesvirus 68 infection.18
Influenza A virus (IAV) is an enveloped RNA virus of the Orthomyxovirus genus. Seasonal influenza is a major cause of respiratory infection resulting in substantial morbidity, mortality, and thus economic burden worldwide.19 Pandemic influenza strains emerge sporadically as a result of genetic reassortment and are a substantial global health concern.20 There are vaccines and antiviral drugs to combat influenza. However, because of rapid virus evolution, vaccines need to be re-formulated and re-administered most years,21 and resistance against antiviral drugs is emerging.22 It is therefore important to understand how intrinsic and innate mechanisms modulate influenza virus infection and how these may be used to develop novel therapeutic interventions.
The negative-sense, single-stranded genome of IAV comprises eight segments of viral RNA, which are separately encapsidated into ribonucleoprotein particles (RNPs).23 Infection and entry of IAV into cells involve viral attachment, via the hemagglutinin glycoprotein that is embedded in the virion membrane, to cell surface receptors that contain sialic acid (SA).24 After binding, virus particles enter the cell by receptor-mediated endocytosis. Fusion of the virus membrane with the endosomal membrane results in release of RNPs into the cytoplasm,24, 25 which are then imported into the nucleus, where genome replication and transcription of viral genes take place.26 Understanding the entry process and how the host counters is critical for uncovering inhibitors with therapeutic potential.
In this study, we used transgenic mouse models combined with in vitro three-dimensional culture systems to show that BPIFA1 has a role in the intrinsic defense against IAV infection.
Results
BPIFA1 Expression after IAV Infection
To determine changes in BPIFA1 levels following IAV infection, C57BL/6J mice were infected intranasally with 103 p.f.u. IAV X-31 (or as controls with ultraviolet-inactivated virus or mock-infected) and analyzed at time points post infection (p.i.). IAV X-31 is a mouse-adapted H3N2 strain that generates a sub-lethal infection.27 It thus models most of the IAV-associated disease in humans and also allows for analysis over an extended time course.
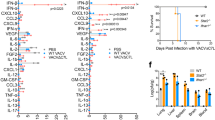
First, levels of BPIFA1 in broncho-alveolar lavage were accurately quantified using (Wes; ProteinSimple, San Jose, CA). The results (Figure 1) showed that BPIFA1 levels decreased markedly after infection (100-fold) to a nadir at day 7 p.i. before returning to a level that was not significantly different to uninfected mice by day 14 p.i. ultraviolet-inactivated virus did not affect the levels of BPIFA1 as measured at day 5 p.i.
BPIFA1 protein levels in broncho-alveolar lavage (BAL) are modulated after influenza A virus (IAV) infection. BAL samples were taken from C57BL/6J mice infected with IAV X-31 at multiple days p.i. The Simple Western (Wes; ProteinSimple) system was used to quantify BPIFA1 protein using a rabbit anti-mBPIFA1 primary antibody. Equal amounts of protein (120 μg as determined by Bicinchoninic acid, ThermoFisher, Paisley, UK) were loaded per lane. The area of the BPIFA1 peak, in relation to the standard curve, was determined using Compass software (ProteinSimple). Bars represent mean±s.e.m. (n=4). Statistically significant differences (one-way analysis of variance with Tukey’s post hoc analysis) between groups are represented by square brackets above. *P<0.05; **P<0.01. ND, not determined.
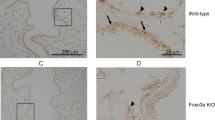
Next, the pattern and quantity of BPIFA1 staining was determined by immunohistology. In mock-infected mice, BPIFA1 was abundant in the epithelium of the trachea and bronchi (Figure 2a,b, arrows), but present only within scattered cells in the bronchioles (Figure 2c, arrows) and absent in the surrounding alveoli (Figure 2c). Following infection with IAV, a substantial decrease in the number of BPIFA1-positive cells was observed and confirmed by the quantitative analysis that showed a 4–8-fold (P<0.05) decrease in the percentage area stained in the trachea and bronchi, respectively, to day 7 p.i.; at this point, there were foci of erosion with reduced cell height, loss of cilia, loss of cellular polarity, and occasional loss of epithelial cells. Recovery of levels of BPIFA1 and evidence of epithelial regeneration with foci of multilayered epithelium and mitoses were seen by day 14 p.i. (Figure 2a,b). The intensity of BPIFA1 staining in the positive areas (i.e., in individual epithelial cells) did not decrease significantly in the trachea, but in the bronchi a similar pattern to that of area stained was noted with an initial decrease followed by a gradual recovery. In contrast, in bronchioles, the intensity of staining in epithelial cells increased fivefold to day 14 p.i., whereas the number of positive cells (area stained) increased threefold by day 14 p.i. (Figure 2c).
BPIFA1 level in airway epithelia is modulated after influenza A virus (IAV) infection. The lungs were dissected from C57BL6/J mice that were either mock- or IAV-infected (intranasally with 103 p.f.u. IAV X-31) at multiple days p.i. as indicated. BPIFA1 was detected by immunohistology using rabbit anti-mBPIFA1, visualized with 3,3′-diaminobenzidine, and counterstained with hematoxylin. The percentage area of epithelium stained (left graph panels) and intensity of staining (right graph panels) in the trachea (a), bronchi (b), and bronchioles (c) were assessed by image analysis. Data are for four mice per group presented as mean±s.e.m. Statistically significant differences (one-way analysis of variance with Tukey’s post hoc analysis) between groups are represented by square brackets above. *P<0.05, **P<0.01, ***P<0.001. Micrographs of representative areas at each time point are shown below the graph panels. Black arrows point to expression of BPIFA1 in respiratory epithelium. Scale Bar=50 μm (a), 10 μm (b), and 20 μm (c).
Mice deficient in BPIFA1 support higher levels of IAV replication
To determine the role of BPIFA1 in defense against IAV, BPIFA1-deficient mice (Bpifa1−/−) were generated, as described in Supplementary Data online and Supplementary Figure S1A. Bpifa1−/− mice expressed no detectable BPIFA1 protein in the lungs (Supplementary Figure S1B), but did not exhibit any morphological changes or altered distribution of cell types in airways and alveoli (Supplementary Figure S1C, as described in Supplementary Data).
C57BL/6J and Bpifa1−/− mice were infected intranasally with 103 p.f.u. IAV X-31. The weight of the mice and lung virus titers were determined over a 28-day period. The results (Figure 3a,b) showed that wt mice were readily infected with IAV. They lost weight (maximum 5%; day 6 p.i.) before recovering. Titers of IAV increased in the lungs, peaking at 106 p.f.u. g−1 tissue on day 5 p.i. Infection in Bpifa1−/− mice followed a similar course. Weight loss was significantly greater (P<0.01) in Bpifa1−/− mice (12% at day 6 p.i.). In addition, IAV titers were significant (P<0.05) and at least 1 log10 higher at all time points. Virus was not detectable in either group at days 7 and 14 p.i.
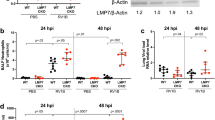
BPIFA1 influences the infection of mice by influenza A virus (IAV). (a,b). C57BL/6J and Bpifa1−/− mice (n=4 per group) were infected intranasally with 103 p.f.u. IAV X-31. (a) Mice were weighed daily and the weights represented as a percentage of the starting weight. Data represent the mean value±s.e.m. Asterisks indicate statistical difference (two-way analysis of variance (ANOVA) with Bonferroni post test; **P<0.01. (b) Lung tissues were taken at multiple days p.i. as indicated and virus titer determined by plaque assay. Data represent the mean value±s.e.m. Asterisks indicate statistical difference (two-way ANOVA with Bonferroni post test; *P<0.05, ***P<0.001). (c) Bpifa1loxP; Scgb1a1-CreERTM mice were treated with either carrier (vegetable oil) or carrier+tamoxifen (tmx) by oral gavage five times as described in the Materials and methods section. Lung tissue (n=4 per group) was analyzed by western blotting for BPIFA1. Equal amounts (100 μg) of protein as determined by Bicinchoninic acid (Pierce) were loaded per lane. The mean (±s.e.m.) levels of BPIFA1 protein were measured by Image J as integrated band intensity. * represents statistical significance (Mann–Whitney U-test; P<0.05) BPIFA1 is clearly knocked down in the Tamoxifen-treated animals. (d) Tamoxifen-inducible, club cell-specific BPIFA1 knockout mice (Bpifa1loxP; Scgb1a1-CreERTM) were dosed on 5 consecutive days by gavage with 0.25 mg g−1 body weight tmx or carrier (vegetable oil). Four days after the last gavage these mice, along with untreated Bpifa1loxP; Scgb1a1-CreERTM and Bpifa1−/− mice (n=4 for all groups) were infected with IAV for 7 days. The lungs were collected, and virus was titer determined by plaque assay. Data represent the mean value±s.e.m. Asterisks indicate statistical difference (one-way ANOVA with Tukey’s post hoc analysis; P<0.05).
To control for potential developmental adaptation in the knockout (KO) mice, a club cell-specific, tamoxifen (tmx)-inducible conditional KO mouse strain (Bpifa1loxP; Scgb1a1-CreERTM) was generated, as described in Supplementary Data. Western blot analysis of lung tissue confirmed a ninefold knockdown in the level of BPIFA1 in Bpifa1loxP; Scgb1a1-CreERTM mice after tmx treatment as compared with mice that were treated with carrier (Figure 3c).
Bpifa1loxP; Scgb1a1-CreERTM mice were treated with either carrier (vegetable oil) or tmx and, along with untreated Bpifa1loxP; Scgb1a1-CreERTM and Bpifa1−/− mice, were infected intranasally with IAV. Virus titers in the lungs were determined at day 7 p.i. The results (Figure 3d) showed that titers of infectious virus were significantly lower in untreated or carrier-treated mice than in tmx-treated Bpifa1loxP; Scgb1a1-CreERTM mice or Bpifa1−/− mice. Thus, reduction of BPIFA1 expression using either total or conditional KO of BPIFA1 enabled IAV to be more pathogenic and replicate to higher titers in the lungs of infected mice.
BPIFA1 limits the initial spread of IAV in vivo
To assess whether BPIFA1 influenced the distribution of IAV infection, C57BL/6J and Bpifa1−/−mice were infected and IAV-infected cells identified by immunohistology. At day 1 p.i., IAV antigen was seen within, and adjacent to, the foci of necrotic respiratory epithelial cells in the nasal cavity, trachea, bronchi, and proximal bronchioles of both wt and Bpifa1−/− mice (Supplementary Figure S2, arrows). However, in Bpifa1−/−mice, epithelial cells in the distal bronchioles were found to be infected, and macrophages in the alveoli around the infected bronchioles were positive for IAV (Figure 4, arrows and red arrowheads, respectively). In contrast, IAV antigen was not observed in distal bronchioles and alveoli of wt mice. At subsequent days p.i., there were no differences in the distribution of IAV antigen between wt and Bpifa1−/− mice.
Influenza A virus (IAV) spreads more rapidly in the absence of BPIFA1. C57BL/6J and Bpifa1−/− mice were infected intranasally with 103 p.f.u. IAV X-31. Lung tissues were collected at 1 day p.i. IAV antigen was detected by immunohistology using goat anti-IAV, visualized with 3,3′-diaminobenzidine, and counterstained with hematoxylin. Micrographs of representative areas from distal bronchioles and alveoli four mice are shown. Large arrows, positive epithelial cells. Red arrowheads, positive macrophages.
Thus, in the absence of BPIFA1, IAV infection reached the distal airways and alveoli earlier during infection.
BPIFA1 restricts IAV infection in normal epithelial cells
Given the differences in viral titer as early as day 1 p.i., we hypothesized that one function of BPIFA1 could be to influence directly the infection of epithelial cells. To investigate this, tracheal cell cultures were established from mice and cultured at the air–liquid interface (ALI) to generate well-differentiated cultures (mouse tracheal epithelial cell, mTEC).28, 29 The validation of these cultures from wt mice is presented in Supplementary Data and Supplementary Figure S3. Differentiated mTEC ALI cultures displayed phenotypic features associated with complex populations of cells including ciliated cells (β-tubulin and FoxJ1), BPIFA1, mucin (MUC5B), and SCGB1A1 (CCSP)-expressing cells that matched with the features of the native rodent airway epithelium. The cultures from Bpifa1−/−mice had similar levels of ciliogenesis (β-tubulin staining) and mucin (MUC5B staining) to those from wt counterparts (Supplementary Figure S4). Quantification of β-tubulin staining showed that ∼17% of cells were ciliated in both types of cultures. In wt cultures, 24% of cells were BPIFA1-positive and were non-ciliated. Thus, mTEC ALI cultures are highly representative of mouse airway epithelium and an ideal model to study the role of BPIFA1 during infection.
mTEC ALI cultures from wt and Bpifa1−/− mice were infected with IAV (0.1 p.f.u. per cell) and infection analyzed by immunofluorescence. In wt mTEC ALI cells, IAV antigen was observed predominantly in epithelial cells that did not express BPIFA1, at both 24 and 48 h p.i. (Figure 5b,c). An increased number of cells containing IAV NP antigen was seen in Bpifa1−/− mTEC ALI cultures as compared with wt cells at both time points (Figure 5e,f). Quantification revealed a more than twofold and nearly twofold higher intracellular IAV NP load in Bpifa1−/− mTEC cultures than in wt mTEC cultures at 24 and 48 h p.i., respectively (Figure 5g; P<0.05). There was also a significant increase in infectious IAV titer in apical wash fluid from Bpifa1−/− cells compared with wt cells (Figure 5h). Thus, BPIFA1 inhibits the infection of normal (mTEC ALI) cells by IAV at a stage up to or before early viral gene expression.
BPIFA1 influences the infection of normal mouse tracheal epithelial cell (mTEC) by influenza A virus (IAV). mTEC air–liquid interface (ALI) cultures were infected with IAV X-31 (0.1 p.f.u. per cell). (a–f) Cells were processed and stained with anti-IAV NP (green) and anti-mBPIFA1 (red), and the nuclei were counterstained with 4′,6-Diamidino-2-phenylindole (blue) and imaged using a confocal microscope. Scale bar=50 μm. Micrographs show representative areas from C57BL/6J mTEC cultures (a–c) and Bpifa1−/− mTEC cultures (d–f) at 2, 24, and 48 h p.i. (g) Mean integrated florescence intensity of IAV nucleoprotein in mTEC ALI shown for three independent biological replicates (±s.e.m.). *P<0.05, one-way analysis of variance (ANOVA) with Tukey’s post hoc analysis. (h) Titer of IAV in apical wash. Data presented as mean±s.e.m.; n=3; *P<0.05, one-way ANOVA with Tukey’s post hoc analysis.
BPIFA1 limits the import of IAV into epithelial cells
Import of virus RNP complexes into the nucleus is one of the initial steps of the IAV replication cycle. To investigate whether BPIFA1 influences the early stages of IAV infection, day 14 mTEC ALI cultures were infected with IAV. Import of RNP complexes into the nucleus was then assessed by immunofluorescence analysis using anti-IAV NP. Blocking protein synthesis with cycloheximide (CHX) was performed to distinguish imported RNPs from de novo NP production. The amount of purified virus used was sufficient to enable the detection of RNP import in 100% of cells in an equivalent assay performed on MDCK cells (data not shown). The results (Figure 6a,b) showed that, in the absence of CHX, IAV NP was readily observed 4 h after infection in the nuclei of cells from wt mice, but, as with the previous experiment, was present in 3.5 × more nuclei in cells from Bpifa1−/− mice. In the presence of CHX, RNP complexes were observed in the nuclei of 1% of wt cells but in a significantly (P<0.05) higher proportion (25 ×) of nuclei in Bpifa1−/− cells (Figure 6a, right panels; Figure 6b,c). Thus, BPIFA1 appreciably decreased the number of cells with visible accumulation of IAV RNPs after infection, indicating that it acts at an early stage of the viral life cycle.
BPIFA1 decreases the nuclear import of influenza A virus (IAV) RNPs into normal mouse tracheal epithelial cell (mTEC). mTEC air–liquid interface cultures from wt C57BL/6J or Bpifa1−/− mice were incubated with purified IAV A/PR/8/34 in the presence or absence of cycloheximide (CHX) as indicated. Cycloheximide blocks protein synthesis and so indicates the import of pre-existing IAV RNP from incoming virus particles. After 4-h incubation at 37 °C, cultures were processed and stained with anti-IAV NP and counterstained with 4′,6-Diamidino-2-phenylindole before visualization using a confocal microscope. Scale bar=50 μm. (a) Representative micrographs from cultures incubated with IAV in the absence or presence of CHX (b) micrographs from cultures incubated with IAV in the presence of CHX at a higher magnification. (c) Nuclei that were positive for IAV NP in the presence of cycloheximide were counted at × 40 magnification in five fields (one center and four peripheral fields) per sample and presented as a percentage of positive cells over total cells counted. The mean percentage positive nuclei are shown for three independent biological replicates (±s.e.m.). * represents statistical significance (one-way analysis of variance with Tukey’s post hoc analysis; P<0.05).
BPIFA1 inhibits IAV binding to respiratory epithelial cells
To determine whether BPIFA1 affected virus binding, mTEC ALI cultures derived from Bpifa1−/− and wt mice were incubated with an Alexa Fluor 488-labeled IAV (IAV-488), and bound virus was assessed using a confocal microscopy. The amount of labeled virus used was sufficient to enable the detection of binding in 100% of cells in an equivalent assay performed on MDCK cells (data not shown). wt cells showed IAV binding predominantly to the surface of Foxj1-positive (ciliated) cells that were negative for BPIFA1 (Figure 7a). The BPIFA1-positive subpopulation of cells (non-ciliated; Supplementary Figure S3A,C) presented as focal clusters in mTEC ALI cultures (outlined in yellow) and were predominantly free from IAV binding (Figure 7a). Analysis of the distribution of α2,3-linked and α2,6-linked SA residues using the lectins MAA and SNA, respectively, showed that there were numerous cells with α2,3-linked residues on the cell surface in both wt and Bpifa1−/− cultures (mean n=3; 25% and 22%, respectively), and that these were on the BPIFA1-negative cells (Supplementary Figure S5A). The few α2,6-positive cells present in the wt and Bpifa1−/− cultures (mean n=3; 3% and 5%, respectively) were mostly BPIFA1-positive (Supplementary Figure S5B–E). Quantification of bound IAV-488 showed that the integrated fluorescence intensity of Bpifa1−/−cells was fourfold greater than that from wt cells (Figure 7b,c). Thus, our data show that IAV bound predominantly to ciliated cells that were BPIFA1-negative, α2,3-linked SA-positive, and that the presence of BPIFA1 decreased the binding of IAV to these cells.
BPIFA1 restricts binding of influenza A virus (IAV) to normal epithelial cells in mouse tracheal epithelial cell (mTEC) air–liquid interface (ALI) cultures. mTEC ALI cultures were incubated for 1 h with IAV Alexa-fluor 488-labeled purified IAV A/PR/8/34 (IAV-488), and then fixed with formalin, stained with anti-BPIFA1 and anti-FoxJ1 antibodies, and imaged using a confocal microscope. (a) Representative images of two different fields of day 14 C57BL/6J mTEC ALI culture showing IAV-488 (green) preferentially binding to FoxJ1-positive (white) ciliated cells but not to BPIFA1 (red)-positive cell populations (circled by yellow dotted line). (b) Micrograph showing IAV-488 binding (green) to the epithelium of wt C57BL/6 and Bpifa1−/− cultures. (c) Integrated florescence intensity of IAV-488 virus binding to the surfaces of mTEC culture. Images were captured from at least five fields per sample by confocal microscope at × 20, × 40, and × 60 magnification. The levels of IAV-488 binding to mTEC cell surfaces were measured using Image J as integrated fluorescence intensity. The mean fluorescent intensity is shown for three independent biological replicates (±s.e.m.). * represents statistical significance (Mann–Whitney U-test; P<0.05).
Discussion
BPIFA1 is constitutively expressed and secreted by the airway epithelium2, 6, 7, 18, 30, 31 including the submucosal glands.9, 28, 30, 31 However, its precise biological functions remain elusive. Our studies have uncovered an important role of BPIFA1 in the host defense against IAV infection. BPIFA1 levels were modulated after infection, and genetic KO of BPIFA1 led to a higher viral titer both in vivo and in mTEC ALI cultures. There was also greater nuclear import of virus RNP complexes and binding of virus to cells in mTEC cultures lacking BPIFA1.
BPIFA1 has previously been shown to have a defensive role in M. pneumoniae and K. pneumoniae infection, enhancing bacterial clearance and inhibiting biofilm formation.15, 16 Here we show, for the first time, that BPIFA1 also has a major role in the defense against IAV. Notably, higher weight loss was observed and higher viral titers (>1 log10) were recovered from the lungs of Bpifa1−/− mice as compared with wt controls. A more rapid spread of IAV to the lung parenchyma was also seen in Bpifa1−/− mice. Specifically, IAV antigen was observed in the bronchiolar epithelium and alveoli of Bpifa1−/− mice but not of wt mice at 24 h p.i., indicating a possible role of BPIFA1 both in intrinsic and innate defense during early infection and in the adaptive immune response to IAV. The higher viral titers in the lungs after conditional knockdown of BPIFA1 in Scgb1a1-expressing club cells indicate that the observations seen in the Bpifa1−/− mice are not due to developmental adaptations to the total loss of BPIFA1 expression.
To investigate intrinsic functions of BPIFA1 in the airway epithelium, we cultured mTEC cells from wt and Bpifa1−/− mice. Although we did not formally rescue the Bpifa1−/− cells using exogenous protein, we controlled for differences between cultures by performing extensive phenotypic characterization. Thus, ∼25% of the cells in wt mTEC had α2,3-linked SA residues on the surface. These cells were BPIFA1-negative. A similar proportion of cells (22%) was α2,3 SA-positive in the Bpifa1−/− cultures. Very few cells on wt and Bpifa1−/− cultures were α2,6 SA-positive (3% and 5%, respectively). Mouse-adapted IAV strains such as those used in this study (X-31 and PR8) will use both α2,3- and α2,6-linked SA residues as a receptor.32, 33 The similar proportions of cells with α2,3- and α2,6-linked SA residues between wt and Bpifa1−/− cultures indicate that differences in infection and binding between wt and Bpifa1−/− cultures are due to the absence or presence of BPIFA1 and are not due to differences in the number of cells expressing receptors. Infection of mTEC cultures with IAV paralleled our observations in vivo in that there was a significantly more rapid spread and higher viral titers in cultures lacking BPIFA1. In line with the distribution of α2,3-linked SA receptors, IAV had a preference for infecting the BPIFA1-negative population. This confirms a role for BPIFA1 in the intrinsic defense against IAV at the airway epithelium during the first few days of infection, and shows that the secreted protein exerts its effects on the epithelial cell surface rather than solely on the cells that produce it.
Further analysis revealed a much greater import of IAV RNPs into the nuclei of cells and significantly increased virus binding in the Bpifa1−/−cultures. The fact that only a percentage of cells were infected in these assays reflects partly the complex nature of mTEC ALI cultures and where only a proportion of the cells express the α2,3- and α2,6 SA receptors. The greater level of inhibition in the RNP import assay compared with the binding assay suggests an important role of BPIFA1 in decreasing post-binding entry into cells via endosomes as well as initial binding of IAV to normal ciliated epithelial cells in airways. BPIFA1 is secreted and is present in the periciliary layer,34 so it is easy to envisage how it might be part of a barrier to IAV infection at the epithelial surface. We did not characterize whether and how BPIFA1 binds to virus particles. However, BPIFA1 could either form a non-specific barrier to IAV engaging with cellular receptors or bind to IAV and prevent receptor interactions. Gel-forming mucins such as MUC5AC inhibit virus infection,35 and BPIFA1 has been shown to associate with the mucus-rich portion of the pericellular lining fluid.34, 36 It has been hypothesized that, in the case of IAV, this is because of competitive inhibition of receptors as MUC5AC from mice contains abundant α2,3-linked SA residues.35 The glycosylation on BPIFA1 contains SA residues37 and could potentially act by binding to IAV particles by specific SA–hemagglutinin interactions. Alternatively, BPIFA1 is a member of the wider tubular lipid-binding (TULIP) superfamily,38 has a hydrophobic cavity, and binds several lipids found in mammalian membranes.39 It therefore has the potential to bind to lipids found in the IAV membrane and interfere with receptor binding and endosomal fusion. Further studies are required to determine precisely how BPIFA1 blocks binding and entry of IAV into normal epithelial cells.
Previous studies have shown that infection by a number of pathogens (e.g., P. aeruginosa, S. pneumoniae, M. pneumoniae, murine γ-herpesvirus 68, and IAV) causes an initial transient increase in secretion of the protein in the first 2–3 h p.i., but this is followed by a decrease over 2–7 days.15, 18, 40 Our results in vivo extend these observations using IAV (Figures 1 and 2). Decrease in BPIFA1 levels after infection is mediated by pathogen-associated molecular patterns and interferon-γ.40 The IAV strains used here target BPIFA1-negative ciliated epithelial cells, and BPIFA1 is produced by non-ciliated epithelial cells. Thus, although IAV induces necrosis of respiratory epithelial cells, the decrease in BPIFA1 expression is unlikely the result from direct killing of BPIFA1-expressing cells. Also, intact respiratory epithelia exhibited decreased BPIFA1 expression at day 7 p.i. (Figure 2). IAV induces signaling by a number of pathogen-associated molecular patterns and so the decrease in BPIFA1 could be due to this mechanism.41 Inactivated IAV had no effect, suggesting that active replication is required to mediate a decrease in BPIFA1 levels. After resolution of the initial infection, BPIFA1 levels returned to pre-infection levels in broncho-alveolar lavage and the tracheal and bronchial epithelium.
It seems unlikely that BPIFA1 has evolved to have a role in the defense against IAV alone and thus it is likely that the protein will influence the pathogenesis of other viruses. It has been shown to have an inhibitory effect on the replication of Epstein–Barr virus (a membrane-bound herpesvirus) in lymphoblastoid cells.42 Downregulation of BPIFA1 expression43 and polymorphisms in the BPIFA1 gene are also associated with susceptibility to Epstein–Barr virus-associated nasopharyngeal carcinoma.44 It will be of interest to determine whether BPIFA1 affects the pathogenesis of other respiratory viruses.
These studies have uncovered a significant and unappreciated role of BPIFA1 in host defense against experimental IAV infections in murine models. It remains to be seen if this protein is able to modulate IAV infection in the human host in a similar manner. The observation of elevated levels of BPIFA1 in nasal aspirates of children with IAV45 suggests that some modulation of the protein does occur during clinical infection and thus is a potential biomarker. We propose that BPIFA1 may be one component of the airway surface lining fluid critical to antiviral host defense. Further studies are needed to determine whether natural variations in BPIFA1 levels due to genetic polymorphisms44 may pre-dispose to more severe IAV-associated disease.
Methods
Virus
Influenza virus strains A/X-31 (X-31, H3N2) and A/PR/8/34 (PR8, H1N1) were propagated in the allantoic cavity of 9-day-old embryonated chicken eggs and titered by plaque assay on MDCK cells.46
Mice
Animal work was reviewed and approved by the local University of Liverpool Animal Welfare Committee and performed under UK Home Office Project Licences 40/2483 and 70/8599. Mice were all specified pathogen-free and maintained under barrier conditions in individually ventilated cages. Transgenic Bpifa1−/−mice were bred at the University of Liverpool under UK Home Office Project Licence 70/8378. Transgenic mice deficient in BPIFA1 (Bpifa1−/−) and conditional KO were generated (Bpifa1loxP; Scgb1a1-CreERTM), as described in S1 text and Supplementary Figure S1. Both transgenic strains were backcrossed 10 generations to C57BL/6J. Wild-type sex- and age-matched C57BL/6J control mice were purchased from Charles River (Margate, UK).
Tmx administration
A 20 mg/ml tmx stock solution was dissolved in Mazola corn oil. Tmx was administered by gavage of 0.25 mg g−p body weight to Bpifa1loxP; Scgb1a1-CreERTM adults daily for 5 days. Mice were then infected with IAV 4 days after the last gavage.
Virus infection of mice
Animals were randomly assigned into multiple cohorts, anesthetized lightly with KETASET i.m. (Zoetis, London, UK), and inoculated intranasally with 103 p.f.u. IAV in 50 μl sterile phosphate-buffered saline, or were mock-infected with a similar volume of allantoic fluid. Mice were killed at variable time points after infection by cervical dislocation. Tissues were removed immediately for downstream processing.
Histology and immunohistology
Histology and immunohistology were performed on paraformaldehyde-fixed, paraffin-embedded tissue using the peroxidase anti-peroxidase method, as previously described.47, 48 Primary antibodies used were rabbit anti-mBPIFA16 and goat anti-IAV (Meridian Life Sciences, B65141G).
Quantitative histopathological assessment
The percentage area and density of 3,3′-diaminobenzidine staining within airway epithelium were quantified using whole-slide images scanned and analyzed using Image-Pro Premier image analysis software version 9.1 (MediaCybernetics, Cambridge, UK). An automated macro and app (named Trachea.ipp and Airways.ipx, respectively) were developed in conjunction with MediaCybernetics. They were designed to identify and outline regions of interest comprising the airway epithelium of the trachea, bronchi, and bronchioles, while excluding other cells. The macro and app analyze thresholds of hue, luminosity, and saturation, which were set relative to the chromogen (DAB) utilized for BPIFA1 localization. These data provided an average density of staining for each region of interest. The percentage area stained was defined as the brown area (positive immunostaining) divided by the total brown area+blue area (hematoxylin; negative staining) × 100.
Western blotting
Samples were analyzed by 12.5% SDS–PAGE gel electrophoresis and blotted using rabbit anti-mBPIFA1.6 Band density was measured using the gel analysis tool of Image J software (NIH, Bethesda, MD).
Quantitative protein analysis
The Wes system was used to identify and quantify BPIFA1 protein in broncho-alveolar lavage and lung samples (ProteinSimple) and rabbit anti-mBPIFA1 as a primary antibody. The area of BPIFA1 peak, in relation to the standard curve, was determined and virtual blot-like images were generated using Compass software (ProteinSimple).
mTEC cultures
mTECs were isolated and differentiated into upper airway-like epithelium in the ALI culture following previously described methodology with slight modification.28, 29 Whole tracheas were excised from 6- to 8-week-old mice. Five or six tracheas were pooled for each evaluation.
Immunofluorescent labeling and confocal imaging
Transwell membranes were fixed with 10% buffered formalin, and then stained using primary and secondary antibodies. Primary antibodies used were as follows: rabbit anti-BPIFA1 (1:200),6 rabbit anti-LPLUNC1 (1:100),6 mouse anti-β tubulin (1:100; Sigma Aldrich, Dorset, UK, catalog no.: T5201), mouse anti-Foxj1 (1:100; eBioscience, Dorset, UK, catalog no.: 14-9965-82, Clone-2A5), goat anti-SCGB1A1 (1:500; Gift from Barry Stripp), rabbit anti-MUC5B (1:100; Santa Cruz, Dorset, UK; H-300, catalog no.: sc-20119), and mouse anti-Influenza A virus NP (anti-IAV NP; 1:200; H16-L10-4R5 (ATCC HB-65, Manassass, VA, USA)). Secondary antibodies are as follows (all from Life Technology, OA; 1:200): Alexa Fluor 568 Goat anti-rabbit antibody (catalog no.: A11011), Alexa Fluor 488 Goat anti-mouse antibody (catalog no.: A11001), Alexa Fluor 488 Rabbit anti-goat antibody (catalog no.: A11078), and Alexa Fluor 633 Goat anti-mouse IgG (catalog no.: A21050). Samples were mounted on glass slides with 4′,6-Diamidino-2-phenylindole Vectashield (Vector Laboratories, Burlingame, CA, catalog no.: H-1200) and visualized with an Olympus Fluoview 1000 Confocal microscope (Center Valley, PA).
Virus labeling and IAV-binding assay
IAV A/PR/8/34 was purified from the allantoic fluid of infected eggs by pelleting through a 30% sucrose/phosphate-buffered saline cushion followed by banding on 15–60% sucrose/phosphate-buffered saline density gradients. Virus was then labeled using a green Alexa-fluorophore labeling kit (Life Technologies, Carlsbad, CA) to yield Alexa Fluor 488-labeled virus (IAV-488). Unreacted dye was removed by pelleting the virus at 125,000 g. For binding assays, IAV-488 was added to the apical surfaces of day 14 mTEC ALI cultures. After incubation for 1 h and washing off unbound virus, membranes were immediately fixed with 10% buffered formalin and co-stained with anti-BPIFA1 and anti-FoxJ1 antibodies. Images were captured from at least five fields per sample using a confocal microscope. Integrated fluorescence intensity was measured using Image J to assess the levels of IAV-488 binding to the cell surface of mTEC cultures. Three independent experiments were conducted with cells from three different batches of mice.
Nuclear import assay of IAV virus ribonucleoprotein (virus RNP) complexes
Apical surfaces of day 14 ALI mTEC cultures were infected with purified IAV A/PR/8/34. Following infection, cells were overlaid with DMEM/F-12 media±CHX at a final concentration of 100 μg ml−1 (no CHX was the control). Cells were incubated in 5% CO2 at 37 °C for 4 h, then washed three times with pre-warmed Hank's Balanced Salt Solution and fixed with 10% buffered formalin as above. Samples were processed and dual immunostained with anti-IAV NP as described above. Cells displaying nuclear localization of viral RNP were counted at × 40 magnification in five fields (one center and four peripheral fields) per sample and presented as a percentage of positive cells over total cells counted. Three independent experiments were conducted with cells from three biologically different batches of mice.
References
Whitsett, J.A. & Alenghat, T. Respiratory epithelial cells orchestrate pulmonary innate immunity. Nat. Immunol. 16, 27–35 (2015).
LeClair, E.E. et al. Genomic organization of the mouse plunc gene and expression in the developing airways and thymus. Biochem. Biophys. Res. Commun. 284, 792–797 (2001).
LeClair, E.E. et al. Cloning and expression of a mouse member of the PLUNC protein family exclusively expressed in tongue epithelium. Genomics 83, 658–666 (2004).
Bingle, C.D. & Craven, C.J. PLUNC: a novel family of candidate host defence proteins expressed in the upper airways and nasopharynx. Hum. Mol. Genet. 11, 937–943 (2002).
Bingle, C.D. & Gorr, S.U. Host defense in oral and airway epithelia: chromosome 20 contributes a new protein family. Int. J. Biochem. Cell Biol. 36, 2144–2152 (2004).
Musa, M. et al. Differential localisation of BPIFA1 (SPLUNC1) and BPIFB1 (LPLUNC1) in the nasal and oral cavities of mice. Cell Tissue Res. 350, 455–464 (2012).
Weston, W.M. et al. Differential display identification of plunc, a novel gene expressed in embryonic palate, nasal epithelium, and adult lung. J. Biol. Chem. 274, 13698–13703 (1999).
Bingle, C.D. & Bingle, L. Characterisation of the human plunc gene, a gene product with an upper airways and nasopharyngeal restricted expression pattern. Biochim. Biophys. Acta 1493, 363–367 (2000).
Di, Y.P., Harper, R., Zhao, Y., Pahlavan, N., Finkbeiner, W. & Wu, R. Molecular cloning and characterization of spurt, a human novel gene that is retinoic acid-inducible and encodes a secretory protein specific in upper respiratory tracts. J. Biol. Chem. 278, 1165–1173 (2003).
Bingle, C.D., LeClair, E.E., Havard, S., Bingle, L., Gillingham, P. & Craven, C.J. Phylogenetic and evolutionary analysis of the PLUNC gene family. Protein Sci. 13, 422–430 (2004).
Bingle, C.D., Bingle, L. & Craven, C.J. Distant cousins: genomic and sequence diversity within the BPI fold-containing (BPIF)/PLUNC protein family. Biochem. Soc. Trans. 39, 961–965 (2011).
Gakhar, L. et al. PLUNC is a novel airway surfactant protein with anti-biofilm activity. PLoS ONE 5, e9098 (2010).
Garcia-Caballero, A. et al. SPLUNC1 regulates airway surface liquid volume by protecting ENaC from proteolytic cleavage. Proc. Natl Acad. Sci. USA 106, 11412–11417 (2009).
McGillivary, G. & Bakaletz, L.O. The multifunctional host defense peptide SPLUNC1 is critical for homeostasis of the mammalian upper airway. PLoS ONE 5, e13224 (2010).
Chu, H.W. et al. Function and regulation of SPLUNC1 protein in Mycoplasma infection and allergic inflammation. J. Immunol. 179, 3995–4002 (2007).
Liu, Y. et al. SPLUNC1/BPIFA1 contributes to pulmonary host defense against Klebsiella pneumoniae respiratory infection. Am. J. Pathol. 182, 1519–1531 (2013).
Thaikoottathil, J.V. et al. SPLUNC1 deficiency enhances airway eosinophilic inflammation in mice. Am. J. Respir. Cell Mol. Biol. 47, 253–260 (2012).
Leeming, G.H. et al. Gammaherpesvirus infection modulates the temporal and spatial expression of SCGB1A1 (CCSP) and BPIFA1 (SPLUNC1) in the respiratory tract. Lab. Invest. 95, 610–624 (2015).
World Health Organization Influenza (seasonal) fact sheet. Available at http://www.who.int/mediacentre/factsheets/fs211/en/ 2014.
Wright, P.F., Neumann, G. & Kawaoka, Y. Orthomyxoviruses. In Fields Virology, Vol 2 (Knipe, D.M. & Howley P.M., eds) 1691–1741 (Philadelphia, Pennsylvania: Lippincott, Williams and Wilkins, 2007).
Krammer, F. & Palese, P. Advances in the development of influenza virus vaccines. Nat. Rev. Drug Discov. 14, 167–182 (2015).
Zaraket, H. et al. Genetic makeup of amantadine-resistant and oseltamivir-resistant human influenza A/H1N1 viruses. J. Clin. Microbiol. 48, 1085–1092 (2010).
Portela, A. & Digard, P. The influenza virus nucleoprotein: a multifunctional RNA-binding protein pivotal to virus replication. J. Gen. Virol. 83 (Pt 4), 723–734 (2002).
Skehel, J.J. & Wiley, D.C. Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin. Annu. Rev. Biochem. 69, 531–569 (2000).
Wharton, S.A., Belshe, R.B., Skehel, J.J. & Hay, A.J. Role of virion M2 protein in influenza virus uncoating: specific reduction in the rate of membrane fusion between virus and liposomes by amantadine. J. Gen. Virol. 75 (Pt 4), 945–948 (1994).
Boulo, S., Akarsu, H., Ruigrok, R.W. & Baudin, F. Nuclear traffic of influenza virus proteins and ribonucleoprotein complexes. Virus Res. 124, 12–21 (2007).
Huang, L., Li, L., Klonowski, K.D., Tompkins, S.M., Tripp, R.A. & Mellor, A.L. Induction and role of indoleamine 2,3 dioxygenase in mouse models of influenza a virus infection. PLoS ONE 8, e66546 (2013).
You, Y., Richer, E.J., Huang, T. & Brody, S.L. Growth and differentiation of mouse tracheal epithelial cells: selection of a proliferative population. Am. J. Physiol. Lung Cell. Mol. Physiol. 283, L1315–L1321 (2002).
You, Y. & Brody, S.L. Culture and differentiation of mouse tracheal epithelial cells. Methods Mol. Biol. 945, 123–143 (2013).
Leclair, E.E. Four BPI (bactericidal/permeability-increasing protein)-like genes expressed in the mouse nasal, oral, airway and digestive epithelia. Biochem. Soc. Trans. 31 (Pt 4), 801–805 (2003).
Bingle, L. et al. SPLUNC1 (PLUNC) is expressed in glandular tissues of the respiratory tract and in lung tumours with a glandular phenotype. J. Pathol. 205, 491–497 (2005).
Bouvier, N.M. & Lowen, A.C. Animal models for influenza virus pathogenesis and transmission. Viruses 2, 1530–1563 (2010).
Meng, B., Marriott, A.C. & Dimmock, N.J. The receptor preference of influenza viruses. Influenza Other Respir. Viruses 4, 147–153 (2010).
Kesimer, M., Ehre, C., Burns, K.A., Davis, C.W., Sheehan, J.K. & Pickles, R.J. Molecular organization of the mucins and glycocalyx underlying mucus transport over mucosal surfaces of the airways. Mucosal Immunol. 6, 379–392 (2013).
Ehre, C. et al. Overexpressing mouse model demonstrates the protective role of Muc5ac in the lungs. Proc. Natl Acad. Sci. USA 109, 16528–16533 (2012).
Radicioni, G. et al. The innate immune properties of airway mucosal surfaces are regulated by dynamic interactions between mucins and interacting proteins: the mucin interactome. Mucosal Immunol. 9, 1442–1454 (2016).
Ghafouri, B., Irander, K., Lindbom, J., Tagesson, C. & Lindahl, M. Comparative proteomics of nasal fluid in seasonal allergic rhinitis. J. Proteome Res. 5, 330–338 (2006).
Alva, V. & Lupas, A.N. The TULIP superfamily of eukaryotic lipid-binding proteins as a mediator of lipid sensing and transport. Biochim. Biophys. Acta 1861 (8 Pt B), 913–923 (2016).
Ning, F. et al. Structural characterization of the pulmonary innate immune protein SPLUNC1 and identification of lipid ligands. FASEB J 28, 5349–5360 (2014).
Britto, C.J., Liu, Q., Curran, D.R., Patham, B., Dela Cruz, C.S. & Cohn, L. Short palate, lung, and nasal epithelial clone-1 is a tightly regulated airway sensor in innate and adaptive immunity. Am. J. Respir. Cell. Mol. Biol. 48, 717–724 (2013).
Chu, H.W. et al. SPLUNC1 regulation in airway epithelial cells: role of Toll-like receptor 2 signaling. Respir. Res. 11, 155 (2010).
Zhou, H.D. et al. Effect of SPLUNC1 protein on the Pseudomonas aeruginosa and Epstein-Barr virus. Mol. Cell. Biochem. 309, 191–197 (2008).
Zhang, B. et al. Identification of tissue-specific genes in nasopharyngeal epithelial tissue and differentially expressed genes in nasopharyngeal carcinoma by suppression subtractive hybridization and cDNA microarray. Genes Chromosomes Cancer 38, 80–90 (2003).
He, Y. et al. Association of PLUNC gene polymorphisms with susceptibility to nasopharyngeal carcinoma in a Chinese population. J. Med. Genet. 42, 172–176 (2005).
Teran, L.M. et al. Immune response to seasonal influenza A virus infection: a proteomic approach. Arch. Med. Res. 43, 464–469 (2012).
Matrosovich, M., Matrosovich, T., Garten, W. & Klenk, H.D. New low-viscosity overlay medium for viral plaque assays. Virol. J. 3, 63 (2006).
Stewart, J.P., Kipar, A., Cox, H., Payne, C., Vasiliou, S. & Quinn, J.P. Induction of tachykinin production in airway epithelia in response to viral infection. PLoS ONE 3, e1673 (2008).
Kipar, A., Kohler, K., Leukert, W. & Reinacher, M. A comparison of lymphatic tissues from cats with spontaneous feline infectious peritonitis (FIP), cats with FIP virus infection but no FIP, and cats with no infection. J. Comp. Pathol. 125, 182–191 (2001).
Acknowledgements
This work was supported by Biotechnology and Biological Sciences Research Council (UK) grants BB/K009664/1 (to JPS, AK, and GHL), BB/K009737/1 (CDB and LB), BBS/E/D/20241864 (PD) and the Georgia Research Alliance to RAT. We wish to thank Barry Stripp for the generous gift of anti-SCGB1A1 and Emma Rawlins for help and advice regarding the use of Scgb1a1-Cre mice and conditional KO mouse technology. We thank the technical staff in the Histology Laboratories, Veterinary Laboratory Services, School of Veterinary Science, University of Liverpool, for excellent technical assistance.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
Author contributions
C.D.B., L.B., A.K., and R.A.T. planned the project, designed experiments, and analyzed the data. J.P.S. planned the project, designed experiments, analyzed the data, interpreted results, and wrote the manuscript. R.V.S. planned and designed the transgenic K.O. strategies. K.M.A. designed and performed the in vitro mTEC experiments, analyzed the data, and interpreted the results. N.A.M. designed and performed the in vivo experiments, analyzed the data, and interpreted the results. A.S. generated the Bpifa1loxP and Bpifa1−/−transgenic mouse lines. S.J. and S.H. helped perform the RNP import and binding assays. P.D. designed and interpreted the SNP import and binding assays. G.H.L. performed and interpreted the histology and immunohistology examinations. All authors reviewed, revised, and approved the manuscript for submission.
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution4.0InternationalLicense.Theimagesor otherthirdpartymaterialinthisarticleareincludedinthearticle's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, userswill need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Akram, K., Moyo, N., Leeming, G. et al. An innate defense peptide BPIFA1/SPLUNC1 restricts influenza A virus infection. Mucosal Immunol 11, 71–81 (2018). https://doi.org/10.1038/mi.2017.45
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2017.45
This article is cited by
-
Genome Scan for Variable Genes Involved in Environmental Adaptations of Nubian Ibex
Journal of Molecular Evolution (2021)
-
Loss of the homeostatic protein BPIFA1, leads to exacerbation of otitis media severity in the Junbo mouse model
Scientific Reports (2018)