Abstract
Tip-enhanced Raman spectroscopy (TERS) is a powerful surface analysis technique that can provide subnanometer-resolved images of nanostructures with site-specific chemical fingerprints. However, due to the limitation of weak Raman signals and the resultant difficulty in achieving TERS imaging with good signal-to-noise ratios (SNRs), the conventional single-peak analysis is unsuitable for distinguishing complex molecular architectures at the subnanometer scale. Here we demonstrate that the combination of subnanometer-resolved TERS imaging and advanced multivariate analysis can provide an unbiased panoramic view of the chemical identity and spatial distribution of different molecules on surfaces, yielding high-quality chemical images despite limited SNRs in individual pixel-level spectra. This methodology allows us to exploit the full power of TERS imaging and unambiguously distinguish between adjacent molecules with a resolution of ~0.4 nm, as well as to resolve submolecular features and the differences in molecular adsorption configurations. Our results provide a promising methodology that promotes TERS imaging as a routine analytical technique for the analysis of complex nanostructures on surfaces.
Similar content being viewed by others
Introduction
One of the ultimate goals of analytical chemistry is to achieve the identification of different species and their chemical properties at the single-molecule scale. To this end, tip-enhanced Raman spectroscopy (TERS) offers an attractive approach to obtaining a panoramic chemical image of a sample’s surface because it combines the advantages of ultrahigh spatial resolution in scanning probe microscopy and sensitive chemical specificity with respect to vibrational fingerprints in Raman spectroscopy1, 2, 3, 4, 5, 6, 7, 8, 9. Significant progress has been made in TERS with respect to its application to nanoscale chemical identification, from carbon nanotubes10, 11, 12, 13, 14, 15 to organic16, 17, 18, 19, 20 and bio-related molecules21, 22, 23, 24, 25, 26, 27, 28. Recently, its spatial resolution has even been pushed down to the subnanometer scale15, 29, 30, 31.
However, despite this progress, it is still highly challenging to obtain a relatively high pixel density TERS image for an area of closely packed species with subnanometer resolution32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, even if the acquired image only has a very limited signal-to-noise ratio (SNR). This result is due to the acquisition time for each pixel being limited by the stability of the system (for example, the stability of molecules and the tip status, as well as the thermal drift), resulting in relatively weak Raman signals at each pixel. Therefore, subnanometer-resolved TERS imaging of closely packed species remains to be demonstrated. Furthermore, understanding how to extract useful chemical information from such TERS images becomes important and increases the demand for advanced statistical analysis methods.
In this paper, by using a scanning tunneling microscope (STM)-controlled, plasmon-enhanced Raman scattering technique under low temperature and ultrahigh vacuum, we demonstrate high-resolution TERS imaging of molecular domains and molecular chains composed of different porphyrin molecules. By developing an advanced multivariate analysis method based on full-spectrum fingerprints (rather than single peaks), we achieve an unbiased panoramic view of the chemical information and spatial distribution of different molecules on surfaces and are even able to resolve submolecular features and differences in molecular adsorption configurations.
Materials and methods
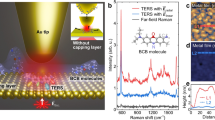
Our experiments were performed using a custom low-temperature ultrahigh-vacuum STM (Unisoku) at ~80 K under a base pressure of ~1 × 10–10 Torr, equipped with a side-illumination confocal optical system47, as shown in Figure 1a and the Supplementary Information (see Supplementary Section S1 for more details). Two different porphyrin molecules, that is, metal-centered zinc-5,10,15,20-tetraphenyl-porphyrin (ZnTPP) and free-base meso-tetrakis (3,5-di-tertiarybutyl-phenyl)-porphyrin (H2TBPP) (upper-right corner of Figure 1a), were used to build different molecular nanostructures on metal surfaces via thermal evaporation. Both ZnTPP and H2TBPP molecules were thermally sublimed at ~580 K onto Ag(111) sequentially. The vacuum pressure in the preparation chamber was <2 × 10−9 Torr during the molecular deposition. The coverage of molecules on the substrate was controlled using different deposition times. Silver (Ag) was used as the tip and substrate material because the Ag nanogap can offer strong plasmonic enhancement. Atomically flat Ag(111) substrates were prepared via cycles of argon ion sputtering and annealing in ultrahigh vacuum. Electrochemically etched Ag tips were used for both STM imaging and TERS measurements, operating in the constant-current-topographic mode with the sample biased48.
TERS imaging of molecular domains adsorbed at silver terraces. (a) Schematic of STM-controlled TERS. The molecular structures of the porphyrin molecules under study (ZnTPP and H2TBPP) are shown in the upper-right corner. (b) STM image of two adjacent porphyrin molecular domains (–1 V, 5 pA). (c) STM image simultaneously acquired during TERS imaging of the area denoted by the dashed square in b (–0.1 V, 1 nA, 7 × 7 nm2, 32 × 32 pixels, 1 s per pixel). The boundary between the molecular domains is highlighted by a yellow line. (d) TERS spectra, averaged over the blue and green squares (3 × 3 pixels) shown in c, extracted from the datacube for ZnTPP and H2TBPP molecules, respectively. (e, f) TERS images reconstructed based on single-peak analysis for the Raman peaks at ~701 cm–1 (e, integrated over 687–736 cm–1) and ~918 cm–1 (f, integrated over 890–959 cm–1).
The laser excitation, Raman photon collection and detection systems are described elsewhere (Supplementary Section S1). A continuous-wave green laser at 532 nm was used as a Raman pumping source with a relatively small photon flux of ∼100 W cm−2 over the junction area. The thermal drift at ∼80 K was ∼150 pm per min. High-resolution Raman spectral imaging was achieved by adopting a plasmon-enhanced STM-controlled TERS29, 30. To generate large enhancement, the nanocavity plasmon mode was tuned to invoke a double-resonance scheme29, 30, with its resonance matching the Raman emission of the molecules particularly well. The acquisition time for each pixel in the TERS mapping process was kept at 1 s in the present work to create a balance between the SNR of the spectra and the thermal drift.
Results and discussion
TERS imaging of molecular domains
Figure 1b shows an STM image of two different porphyrin molecular islands that are packed close to each other, that is, within the van der Waals contact distance. A 7 × 7 nm2 area, denoted by the dashed square in Figure 1b, across the molecular boundary was selected for TERS imaging with a pixel resolution of 0.22 nm per pixel, in which a TERS spectrum was recorded at each pixel during raster scanning. A simultaneously acquired STM image is shown in Figure 1c, in which the H2TBPP molecules appear brighter than the ZnTPP molecules due to the presence of eight additional tertiary butyl groups (see Supplementary Section S2 for more details). During the TERS mapping, the tip was very close to the molecular layer, and the local order of the molecular layer could be disturbed. Fortunately, owing to the stable feedback control in our ultrahigh-vacuum, low-temperature STM system, there was hardly any modification to the ZnTPP molecular layer despite some disturbances on the lobes of the H2TBPP molecules.
Figure 1d shows the TERS spectra, averaged over the blue and green squares (3 × 3 pixels) shown in Figure 1c, extracted from the datacube for the ZnTPP and H2TBPP molecules, respectively. Because of the limited acquisition time and resultant weak Raman signals, the averaging procedure was used to improve the SNR of these TERS spectra to obtain the molecular vibrational fingerprints, which indeed agree with those of previous reports29, 30. As shown in Figure 1d, the fingerprint features of the TERS spectra for the ZnTPP and H2TBPP molecules are evidently different from each other. However, when we use conventional analysis based on single Raman peaks, the optical contrasts of the reconstructed TERS images are still limited, even for the strong characteristic Raman peaks (for example, ~701 cm–1 for ZnTPP30 and ~918 cm–1 for H2TBPP29) (Figure 1e and 1f), because of the low SNR of the Raman spectra at individual pixels. Although the slightly brighter area in Figure 1e (Figure 1f) can be generally attributed to the ZnTPP (H2TBPP) islands, a clear and conclusive identification is still hard to achieve without a priori knowledge about the molecules under study. The chemical identification becomes even more difficult for TERS images based on the analysis of weak or overlapped spectral peaks (see Supplementary Section S2 for more details). Therefore, it is highly desirable to have an advanced statistical analysis method that can take into account the full-spectrum fingerprint information and provide an unbiased panoramic view of the spatial distribution of different molecular species on surfaces.
Multivariate analysis of the TERS data set over complex molecular domains
Multivariate analysis has been widely used in hyperspectral imaging, from fluorescence to Raman and reflectance imaging34, 39, 49, 50, 51, 52. Here we develop an optimized multivariate analysis pipeline to extract chemical information from the subnanometer-resolved TERS images based on the analysis of full-spectrum fingerprints rather than single peaks. The analysis pipeline is composed of several steps, including principal component analysis (PCA)39, hierarchical clustering analysis (HCA)53 and vertex component analysis (VCA)34, 54, as detailed below.
The datacube of a TERS image is first preprocessed via total variation-constrained denoising55, 56, baseline correction with asymmetric least squares57 and vector normalization (the integral of each spectrum is set equal to 1) to improve the signal SNR and to make the spectra at each pixel comparable, as shown in Figure 2a (see Supplementary Section S3 for more details). Next, the datacube is processed by conducting PCA to identify the most significant spectral features. In the PCA space, each spectrum in the datacube can be considered as a single point in this multidimensional space, as shown in Figure 2b. PCA is a common first-step process in multivariate data analysis and is advantageous in that uncorrelated noise is distributed equally throughout the principal components (PCs), whereas important signal variations are concentrated in the first few PCs. Therefore, projecting the data into a lower-dimensional PC space (the first 10 principal components are used in our multivariate fingerprint analysis) has the effect of preserving the important spectral features in the data set while removing noise.
Data processing and clustering in the multivariate analysis for the TERS imaging of molecular domains. (a) Spectra preprocessed via denoising and baseline correction. (b) TERS datacube in PCA subspace. (c) Dendrogram illustrating the clustering of the data set based on the pairwise distances defined in b. The red and gray lines indicate the merge distance values that divide the entire data set into 4 clusters and 30 clusters, respectively. (d) Dependence of the merge distance on the number of clusters.
Then, HCA is applied to the dimension-reduced data set, which is briefly described as follows. In the PCA space with the first 10 PCs, the pairwise distance between two spectral points can be defined as the Euclidean distance (that is, spectral distance), as shown in Figure 2b (for visualization purposes, only the first three PCs are shown). With such a definition, the spectral distance provides a quantitative evaluation of the degree of spectral similarity between two spectra. That is, the longer the spectral distance, the greater the spectral difference. Thus, similar spectra are clustered near each other, whereas dissimilar spectra (those with large pairwise distances) are spatially separated in the PCA space. A dendrogram of the data set is constructed by repeatedly merging two clusters to form a larger cluster based on these pairwise distances, as shown in Figure 2c, which depicts an example application to the TERS image of molecular domains presented in Figure 1 (see Supplementary Section S3 for more details). In our method, Ward’s criterion58, 59 is used for clustering the data, where at each step in the clustering, the two clusters chosen for merging are those whose merging leads to the smallest increase in within-cluster variance.
To guide the selection of the appropriate number of clusters for our data set, we can evaluate the change in merge distance versus the number of clusters, as plotted in Figure 2d. (For visualization purposes, we truncate Figure 2d at 30 clusters, corresponding to the gray line in Figure 2c). Evidently, the merge distance decreases rapidly when the number of clusters is increased from two to three, indicating that the largest difference in the data set is between clusters one and two. The change in merge distance becomes much smaller for larger numbers of clusters, indicating much subtler spectral differences between clusters. The occurrence of a unique and large merge-distance difference between two and three clusters strongly suggests that the most likely number of chemically distinct components is 2, which is consistent with the real number of molecular species for the molecular domain sample in Figure 1.
We chose two clusters, which is the most likely number of clusters, as suggested by the analysis above, to generate an HCA image for TERS imaging of the molecular domain sample presented in Figure 1. The HCA image can be generated by coloring each cluster with a particular color for all pixels. As shown in Figure 3a, the delineation of different molecular domains can be unambiguously observed, being much clearer than in the images presented in Figure 1e and 1f, which are based on single-peak analysis. The right panel shows the corresponding distinctive TERS spectra that are averaged over all the spectra in each cluster, with an improved SNR of ~30-fold compared with the raw single-pixel data.
HCA and VCA for the TERS imaging of molecular domains. (a, b) HCA image and corresponding cluster average spectra when the number of clusters is set to two and four, respectively. (c, d) VCA image and corresponding endmember spectra when the number of endmembers is set to two and four, respectively.
According to the dendrogram in Figure 2c, each cluster corresponding to different molecular domains can be further divided into two subclusters, giving rise to four clusters. In this case, as shown in Figure 3b, although the spectral differences between the subclusters in each domain are relatively small, the HCA image seems to exhibit certain submolecular features. However, such patch-like patterns appear randomly and are difficult to correlate with the relatively ordered molecular domains shown in Figure 1b. Such inconsistency is due to the drawback of HCA, which assumes that each spectrum at each pixel must be assigned to one cluster or another. However, in reality, due to the finite spatial confinement of the nanocavity plasmon, each spectrum at each pixel (0.22 nm per pixel) can still contain contributions from different subclusters at adjacent pixels, which violates the assumption underlying HCA.
To overcome this limitation of HCA, another multivariate analysis technique called VCA is used, which assumes that each spectrum is a linear combination of contributions from several underlying components (called ‘endmembers’) (see Supplementary Section S4 for more details). The number of endmembers can be estimated based on the dendrogram and the associated analysis provided by the HCA described above.
In the simplest case, that is, a two-component mixture, the data set is projected into a one-dimensional space (the PC1 axis), as shown in Figure 2b, in which the ‘purest’ pixels are the points lying at the extrema of this axis. The pixels near the center could be defined as mixtures of these two purest pixels. Non-negative least squares fitting is used to determine the amount that each endmember contributes to each pixel, which yields the VCA abundance image shown in Figure 3c. The VCA image clearly shows how the TERS spectra gradually transition from one molecular domain to another. By analyzing the line profiles across the domain boundary in the VCA abundance image, the spatial resolution of TERS imaging for chemical identification can be estimated down to ~0.4 nm (see Supplementary Section S5 for more details). The origin of the subnanometer resolution is believed to correlate with the highly confined plasmonic field inside the STM junction29, 30, 60, 61, 62. Although the exact mechanism is not yet completely clear, some recent theoretical studies suggest that the subnanometer resolution may involve additional effects such as field gradients63, molecular self-interaction64 and even the chemical effect associated with the interaction with a single atom at the tip apex65.
Such subnanometer-resolved spatial resolution (~0.4 nm) implies that the TERS images can provide submolecular information. Indeed, when we analyze the TERS imaging data set of molecular domains with four VCA endmembers, the generated VCA image (Figure 3d) reveals fine structures in both molecular domains. As mentioned before, the molecular lobes in the H2TBPP domain can be disturbed slightly by the tip during the TERS mapping process. As a result, we focus on the analysis and discussion of the TERS imaging pattern in the intact ZnTPP molecular domain, even though some irregular periodic-like patterns appear in the H2TBPP domain (Figure 3d). The TERS-VCA image for the ZnTPP domain shows relatively periodic patterns, which can be divided into blue and red regions with two different spectral profiles in terms of relative intensities among different Raman peaks. As shown in the solid (dashed) line profile of Figure 4a, the periodicity of the blue (red) region along the green dashed line in the inset VCA image is ~1.4 nm, which is in good agreement with one of the lattice constants (n) measured based on a high-resolution STM topograph for an ordered self-assembled ZnTPP monolayer (Figure 4b). The periodic pattern for the ZnTPP domain in the TERS-VCA image also resembles the unit-cell symmetry of the ordered ZnTPP packing structure.
Chemical origin of the submolecular-resolved TERS spectral feature in the ZnTPP molecular domain. (a) Abundance profiles for two ZnTPP endmembers along the green line in the inset VCA image (same as Figure 3d). (b) High-resolution STM image (−1 V, 100 pA) of a ZnTPP monolayer acquired by using a special tip, with its tip apex modified by a molecule. (c) Top view (left panel) and side view (right panel) of the absorption configuration of a ZnTPP molecule on Ag(111) used for theoretical simulations. See Supplementary Section S6 for details. (d, e) Simulated VCA abundance image (d) and associated endmember spectra (thick lines) for a ZnTPP molecular domain, with experimental endmember spectra (thin lines) also plotted for comparison (e).
To determine the chemical origin of the red and blue regions in the VCA abundance map, we carried out density functional theory calculations based on a previously established molecular configuration30 (Figure 4c) and simulated site-specific TERS spectra by treating the tip as a highly localized plasmonic field scanning over the ZnTPP molecular domain, as detailed in Supplementary Section S6. A VCA procedure similar to that described above was then applied to these simulated TERS spectra by selecting two endmembers for the ZnTPP domain. The simulated VCA abundance image and related endmember spectra are shown in Figure 4d and 4e, respectively, which reproduce the periodic patterns and the spectral difference for the ZnTPP domain shown in Figure 3d fairly well. As illustrated in Figure 4d, when the tip is positioned above the molecular lobes (the red region), the corresponding spectra exhibit a strong Raman peak at ~700 cm−1 (the red curve in Figure 4e) that is associated with the out-of-plane vibrational mode of the porphyrin core and phenyl ring (see Supplementary Movie for details); when the tip is positioned above the molecular core areas (the blue region), the corresponding spectra exhibit strong Raman peaks at ~1200 cm−1 and above (the blue curve in Figure 4e) due to the stronger involvement of the porphyrin core vibrations (see Supplementary Movie for details). Therefore, by combining with the theoretical simulations, we can assign the red region in the experimentally observed TERS-VCA image of Figure 3d to the ZnTPP molecular lobe areas and the blue region in Figure 3d to the ZnTPP molecular core areas. Such ability to distinguish submolecular features in molecular domains illustrates the power of the combined TERS-VCA method to map local vibrations and associated local structural units within a single molecule. We note that TERS signals also show up in the pinhole between two molecular domains, which are caused by TERS signals from the surrounding molecules because of the small size of the pinhole (comparable to the spatial extent of the localized plasmonic field) and the inability of the tip to reach down to the bare metal surface when scanning over the pinhole.
Multivariate analysis of TERS imaging of molecular chains
The PCA–HCA–VCA analysis pipeline can also be used to study randomly distributed molecular mixtures. In the following, we demonstrate an example of the multivariate analysis applied to a mixed molecular chain. Figure 5a shows an STM image of two molecular chains adsorbed on silver step edges. A 16 × 16 nm2 area denoted by the dashed square in Figure 5a, containing a mixture of ZnTPP and H2TBPP molecules, was selected for TERS imaging, with a simultaneously acquired STM image shown in Figure 5b.
VCA for TERS imaging of a mixed molecular chain adsorbed at silver step edges. (a) STM topograph of molecular chains adsorbed at silver step edges (–1 V, 10 pA). (b) STM image simultaneously acquired during TERS imaging of the area denoted by the dashed square in a (–0.3 V, 0.8 nA, 16 × 16 nm2, 32 × 32 pixels, 0.5 nm per pixel, 1 s per pixel). (c) Dendrogram of the data set. The gray and red lines indicate the merge distance values that divide the whole data set into three clusters and four clusters, respectively. (d, e) TERS-VCA image and corresponding endmember spectra for three endmembers. (f, g) TERS-VCA image and corresponding endmember spectra for four endmembers. See Supplementary Sections 7 and 8 for more details.
The TERS spectra for ZnTPP and H2TBPP molecules adsorbed at the step edges are shown to have more similarities than those adsorbed on the terraces (see Supplementary Section S7 for more details), presumably due to different adsorption configurations and orientations. In this case, it is more difficult to distinguish between different molecular species via single-peak analysis (see Supplementary Section S7 for more details). However, the subtle spectral differences can still be resolved via the multivariate analysis based on the full-spectrum fingerprints, as demonstrated below.
Figure 5c shows the dendrogram of the hyperspectral data set, obtained according to the procedure described above. At first glance, there are two evidently different components, which corresponds very well to the co-existence of both a Raman-silent bare metal surface and Raman-active organic molecules (see Supplementary Section S8 for more details). If we further examine the molecular branch in the dendrogram, we can divide the entire data set into three clusters (indicated by the gray line). In this case, two types of porphyrin molecules (ZnTPP and H2TBPP) as well as their locations in the molecular chain can be clearly distinguished in the generated VCA image (Figure 5d) based on their chemical fingerprints (Figure 5e). The blue-dominant areas represent the ZnTPP molecules, and the green-dominant areas represent the H2TBPP molecules, which correlates very well with the STM image presented in Figure 5a.
Interestingly, as shown in Figure 5f and 5g, the ZnTPP areas can be further divided into red and blue regions if the endmember number is chosen to be four (indicated by the red dashed line in Figure 5c). Around the step edges, single ZnTPP molecules adsorbed at the lower terrace (for example, the one highlighted by the small arrow in Figure 5a and 5f) or the portions of ZnTPP molecules overhanging and leaning toward the lower terrace but adsorbed at the upper terrace are identified as the red endmember. The spectral differences between the two ZnTPP endmembers in Figure 5g revealed by the analysis reflect the differences in molecular adsorption configurations and the influence of the local environment30. However, because of the complicated plasmonic field distribution at the step edge, it is still challenging to theoretically simulate the TERS-VCA abundance image experimentally observed in Figure 5g around the step edge. In this sense, our environment-sensitive experimental results may provide fine spectral features that can help to confirm more sophisticated theoretical models or test theoretical predictions. We note that further division into more clusters is not meaningful because the associated merge distance enters the noise range indicated by the merge distance value (~0.1) for the Raman-silent bare metal surface, as illustrated in Figure 5c.
Conclusions
In summary, by combining subnanometer-resolved TERS imaging with an advanced multivariate analysis method, we have demonstrated an unbiased method for mapping the chemical identity and spatial distribution of different molecules on surfaces, which can even resolve submolecular features and the differences in molecular adsorption configurations. Our analysis clearly highlights the power of multivariate fingerprint analysis to unlock the full potential of even a noisy TERS hyperspectral data set. The spatial distribution of different species in TERS imaging can easily be obtained using HCA images. Thanks to the intelligent assumption of VCA regarding the natural mixing of spectra in a data set (which is lost during HCA), further analysis via VCA can provide a better estimate of the spectra of chemical species and yield ‘natural images’ with submolecular features that can correlate with local molecular vibrations and structural units within a single molecule. In short, treating the images by multivariate analysis allows easier identification of the chemical composition in molecular structures on a surface and can reveal further details not readily observed when using single-peak analysis. Our results can help to promote TERS as a routine analytical technique for the analysis of complex nanostructures and lay the foundation for the exploration of nanomaterials, nanodevices, surface chemistry and biology.
References
Hartschuh A . Tip-enhanced near-field optical microscopy. Angew Chem Int Ed 2008; 47: 8178–8191.
Yeo BS, Stadler J, Schmid T, Zenobi R, Zhang WH . Tip-enhanced Raman spectroscopy — its status, challenges and future directions. Chem Phys Lett 2009; 472: 1–13.
Berweger S, Raschke MB . Signal limitations in tip-enhanced Raman scattering: the challenge to become a routine analytical technique. Anal Bioanal Chem 2010; 396: 115–123.
Pettinger B, Schambach P, Villagómez CJ, Scott N . Tip-enhanced Raman spectroscopy: near-fields acting on a few molecules. Annu Rev Phys Chem 2012; 63: 379–399.
Stadler J, Schmid T, Zenobi R . Developments in and practical guidelines for tip-enhanced Raman spectroscopy. Nanoscale 2012; 4: 1856–1870.
Schmid T, Opilik L, Blum C, Zenobi R . Nanoscale chemical imaging using tip-enhanced Raman spectroscopy: a critical review. Angew Chem Int Ed 2013; 52: 5940–5954.
Kurouski D . Advances of tip-enhanced Raman spectroscopy (TERS) in electrochemistry, biochemistry, and surface science. Vib Spectrosc 2017; 91: 3–15.
Pozzi EA, Goubert G, Chiang N, Jiang N, Chapman CT et al. Ultrahigh-vacuum tip-enhanced Raman spectroscopy. Chem Rev 2017; 117: 4961–4982.
Zhang ZL, Sheng SX, Wang RM, Sun MT . Tip-enhanced Raman spectroscopy. Anal Chem 2016; 88: 9328–9346.
Hartschuh A, Sánchez EJ, Xie XS, Novotny L . High-resolution near-field Raman microscopy of single-walled carbon nanotubes. Phys Rev Lett 2003; 90: 095503.
Anderson N, Hartschuh A, Cronin S, Novotny L . Nanoscale vibrational analysis of single-walled carbon nanotubes. J Am Chem Soc 2005; 127: 2533–2537.
Yano T, Verma P, Saito Y, Ichimura T, Kawata S . Pressure-assisted tip-enhanced Raman imaging at a resolution of a few nanometres. Nat Photonics 2009; 3: 473–477.
Yano T, Ichimura T, Kuwahara S, H'Dhili F, Uetsuki K et al. Tip-enhanced nano-Raman analytical imaging of locally induced strain distribution in carbon nanotubes. Nat Commun 2013; 4: 2592.
Chen C, Hayazawa N, Kawata S . A 1.7 nm resolution chemical analysis of carbon nanotubes by tip-enhanced Raman imaging in the ambient. Nat Commun 2014; 5: 3312.
Liao MH, Jiang S, Hu CR, Zhang R, Kuang YM et al. Tip-enhanced Raman spectroscopic imaging of individual carbon nanotubes with subnanometer resolution. Nano Lett 2016; 16: 4040–4046.
Domke KF, Zhang D, Pettinger B . Toward Raman fingerprints of single dye molecules at atomically smooth Au(111). J Am Chem Soc 2006; 128: 14721–14727.
Jiang N, Foley ET, Klingsporn JM, Sonntag MD, Valley NA et al. Observation of multiple vibrational modes in ultrahigh vacuum tip-enhanced Raman spectroscopy combined with molecular-resolution scanning tunneling microscopy. Nano Lett 2012; 12: 5061–5067.
Klingsporn JM, Jiang N, Pozzi EA, Sonntag MD, Chulhai D et al. Intramolecular insight into adsorbate-substrate interactions via low-temperature, ultrahigh-vacuum tip-enhanced Raman spectroscopy. J Am Chem Soc 2014; 136: 3881–3887.
Chiang NH, Jiang N, Chulhai DV, Pozzi EA, Hersam MC et al. Molecular-resolution interrogation of a porphyrin monolayer by ultrahigh vacuum tip-enhanced Raman and fluorescence spectroscopy. Nano Lett 2015; 15: 4114–4120.
Zhong JH, Jin X, Meng LY, Wang X, Su HS et al. Probing the electronic and catalytic properties of a bimetallic surface with 3 nm resolution. Nat Nanotechnol 2016; 12: 132–136.
Domke KF, Zhang D, Pettinger B . Tip-enhanced Raman spectra of picomole quantities of DNA nucleobases at Au(111). J Am Chem Soc 2007; 129: 6708–6709.
Bailo E, Deckert V . Tip-enhanced Raman spectroscopy of single RNA strands: towards a novel direct-sequencing method. Angew Chem Int Ed 2008; 47: 1658–1661.
Treffer R, Lin XM, Bailo E, Deckert-Gaudig T, Deckert V . Distinction of nucleobases–a tip-enhanced Raman approach. Beilstein J Nanotechnol 2011; 2: 628–637.
Deckert-Gaudig T, Kämmer E, Deckert V . Tracking of nanoscale structural variations on a single amyloid fibril with tip-enhanced Raman scattering. J Biophotonics 2012; 5: 215–219.
Lipiec E, Sekine R, Bielecki J, Kwiatek WM, Wood BR . Molecular characterization of DNA double strand breaks with tip-enhanced Raman scattering. Angew Chem Int Ed 2014; 53: 169–172.
Najjar S, Talaga D, Schué L, Coffinier Y, Szunerits S et al. Tip-enhanced Raman spectroscopy of combed double-stranded DNA bundles. J Phys Chem C 2014; 118: 1174–1181.
Liu J, Yin DY, Wang SS, Chen H-Y, Liu Z . Probing low-copy-number proteins in a single living cell. Angew Chem Int Ed 2016; 55: 13215–13218.
Bonhommeau S, Talaga D, Hunel J, Cullin C, Lecomte S . Tip-enhanced Raman spectroscopy to distinguish toxic oligomers from Aβ1-42 fibrils at the nanometer scale. Angew Chem Int Ed 2017; 56: 1771–1774.
Zhang R, Zhang Y, Dong ZC, Jiang S, Zhang C et al. Chemical mapping of a single molecule by plasmon-enhanced Raman scattering. Nature 2013; 498: 82–86.
Jiang S, Zhang Y, Zhang R, Hu C, Liao M et al. Distinguishing adjacent molecules on a surface using plasmon-enhanced Raman scattering. Nat Nanotechnol 2015; 10: 865–869.
Chiang N, Chen X, Goubert G, Chulhai DV, Chen X et al. Conformational contrast of surface-mediated molecular switches yields Ångstrom-scale spatial resolution in ultrahigh vacuum tip-enhanced Raman spectroscopy. Nano Lett 2016; 16: 7774–7778.
Steidtner J, Pettinger B . Tip-enhanced Raman spectroscopy and microscopy on single dye molecules with 15 nm resolution. Phys Rev Lett 2008; 100: 236101.
Stadler J, Schmid T, Zenobi R . Nanoscale chemical imaging using top-illumination tip-enhanced Raman spectroscopy. Nano Lett 2010; 10: 4514–4520.
Richter M, Hedegaard M, Deckert-Gaudig T, Lampen P, Deckert V . Laterally resolved and direct spectroscopic evidence of nanometer-sized lipid and protein domains on a single cell. Small 2011; 7: 209–214.
Stadler J, Schmid T, Opilik L, Kuhn P, Dittrich PS et al. Tip-enhanced Raman spectroscopic imaging of patterned thiol monolayers. Beilstein J Nanotechnol 2011; 2: 509–515.
Paulite M, Blum C, Schmid T, Opilik L, Eyer K et al. Full spectroscopic tip-enhanced Raman imaging of single nanotapes formed from β-Amyloid(1-40) peptide fragments. ACS Nano 2013; 7: 911–920.
Pozzi EA, Sonntag MD, Jiang N, Klingsporn JM, Hersam MC et al. Tip-enhanced Raman imaging: an emergent tool for probing biology at the nanoscale. ACS Nano 2013; 7: 885–888.
Picardi G, Królikowska A, Yasukuni R, Chaigneau M, Escude M et al. Exchange of methyl- and azobenzene-terminated alkanethiols on polycrystalline gold studied by tip-enhanced Raman mapping. ChemPhysChem 2014; 15: 276–282.
Rusciano G, Zito G, Isticato R, Sirec T, Ricca E et al. Nanoscale chemical imaging of Bacillus subtilis spores by combining tip-enhanced Raman scattering and advanced statistical tools. ACS Nano 2014; 8: 12300–12309.
Lin W-I, Shao F, Stephanidis B, Zenobi R . Tip-enhanced Raman spectroscopic imaging shows segregation within binary self-assembled thiol monolayers at ambient conditions. Anal Bioanal Chem 2015; 407: 8197–8204.
Opilik L, Payamyar P, Szczerbiński J, Schütz AP, Servalli M et al. Minimally invasive characterization of covalent monolayer sheets using tip-enhanced Raman spectroscopy. ACS Nano 2015; 9: 4252–4259.
Deckert-Gaudig T, Kurouski D, Hedegaard MAB, Singh P, Lednev IK et al. Spatially resolved spectroscopic differentiation of hydrophilic and hydrophobic domains on individual insulin amyloid fibrils. Sci Rep 2016; 6: 33575.
Dill TJ, Rozin MJ, Palani S, Tao AR . Colloidal nanoantennas for hyperspectral chemical mapping. ACS Nano 2016; 10: 7523–7531.
Jiang N, Chiang N, Madison LR, Pozzi EA, Wasielewski MR et al. Nanoscale chemical imaging of a dynamic molecular phase boundary with ultrahigh vacuum tip-enhanced Raman spectroscopy. Nano Lett 2016; 16: 3898–3904.
Touzalin T, Dauphin AL, Joiret S, Lucas IT, Maisonhaute E . Tip enhanced Raman spectroscopy imaging of opaque samples in organic liquid. Phys Chem Chem Phys 2016; 18: 15510–15513.
Xiao LF, Schultz ZD . Targeted-TERS detection of integrin receptors on human cancer cells. Cancer Cell Microenviron 2016; 3: e1419.
Chen LG, Zhang C, Zhang R, Zhang XL, Dong ZC . Note: optical optimization for ultrasensitive photon mapping with submolecular resolution by scanning tunneling microscope induced luminescence. Rev Sci Instrum 2013; 84: 066106.
Zhang C, Gao B, Chen LG, Meng QS, Yang H et al. Fabrication of silver tips for scanning tunneling microscope induced luminescence. Rev Sci Instrum 2011; 82: 083101.
Gowen AA, O'Donnell CP, Cullen PJ, Downey G, Frias JM . Hyperspectral imaging - an emerging process analytical tool for food quality and safety control. Trends Food Sci Technol 2007; 18: 590–598.
Raicu V, Stoneman MR, Fung R, Melnichuk M, Jansma DB et al. Determination of supramolecular structure and spatial distribution of protein complexes in living cells. Nat Photonics 2009; 3: 107–113.
Davis BM, Hemphill AJ, Maltaş DC, Zipper MA, Wang P et al. Multivariate hyperspectral Raman imaging using compressive detection. Anal Chem 2011; 83: 5086–5092.
Huang CK, Ando M, Hamaguchi H, Shigeto S . Disentangling dynamic changes of multiple cellular components during the yeast cell cycle by in vivo multivariate Raman imaging. Anal Chem 2012; 84: 5661–5668.
Diem M, Romeo M, Boydston-White S, Miljković M, Matthäus C . A decade of vibrational micro-spectroscopy of human cells and tissue (1994-2004). Analyst 2004; 129: 880–885.
Nascimento JMP, Dias JMB . Vertex component analysis: a fast algorithm to unmix hyperspectral data. IEEE Trans Geosci Remote Sens 2005; 43: 898–910.
Goldstein T, Osher S . The split Bregman method for L1-regularized problems. SIAM J Imaging Sci 2009; 2: 323–343.
Chu KQ, McMillan PJ, Smith ZJ, Yin J, Atkins J et al. Image reconstruction for structured-illumination microscopy with low signal level. Opt Express 2014; 22: 8687–8702.
Boelens HFM, Dijkstra RJ, Eilers PHC, Fitzpatrick F, Westerhuis JA . New background correction method for liquid chromatography with diode array detection, infrared spectroscopic detection and Raman spectroscopic detection. J Chromatogr A 2004; 1057: 21–30.
Ward JH Jr . Hierarchical grouping to optimize an objective function. J Am Stat Assoc 1963; 58: 236–244.
Szekely GJ, Rizzo ML . Hierarchical clustering via joint between-within distances: extending Ward's minimum variance method. J Classif 2005; 22: 151–183.
Duan S, Tian GJ, Ji YF, Shao JS, Dong ZC et al. Theoretical modeling of plasmon-enhanced Raman images of a single molecule with subnanometer resolution. J Am Chem Soc 2015; 137: 9515–9518.
Barbry M, Koval P, Marchesin F, Esteban R, Borisov AG et al. Atomistic near-field nanoplasmonics: reaching atomic-scale resolution in nanooptics. Nano Lett 2015; 15: 3410–3419.
Trautmann S, Aizpurua J, Götz I, Undisz A, Dellith J et al. A classical description of subnanometer resolution by atomic features in metallic structures. Nanoscale 2017; 9: 391–401.
Meng LY, Yang ZL, Chen JN, Sun MT . Effect of electric field gradient on sub-nanometer spatial resolution of tip-enhanced Raman spectroscopy. Sci Rep 2015; 5: 9240.
Zhang C, Chen B-Q, Li Z-Y . Optical origin of subnanometer resolution in tip-enhanced Raman mapping. J Phys Chem C 2015; 119: 11858–11871.
Latorre F, Kupfer S, Bocklitz T, Kinzel D, Trautmann S et al. Spatial resolution of tip-enhanced Raman spectroscopy - DFT assessment of the chemical effect. Nanoscale 2016; 8: 10229–10239.
Acknowledgements
This work is supported by the National Natural Science Foundation of China, the National Basic Research Program of China, and the Strategic Priority Research Program of the Chinese Academy of Sciences. ZJS gratefully acknowledges funding support from the NSFC’s 1000 Young Talents Recruitment Plan for Global Experts.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Note: Supplementary Information for this article can be found on the Light: Science & Applications’ website.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Jiang, S., Zhang, X., Zhang, Y. et al. Subnanometer-resolved chemical imaging via multivariate analysis of tip-enhanced Raman maps. Light Sci Appl 6, e17098 (2017). https://doi.org/10.1038/lsa.2017.98
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/lsa.2017.98
Keywords
This article is cited by
-
Tip-enhanced Raman scattering
Nature Reviews Methods Primers (2024)
-
Surface potential modulation as a tool for mitigating challenges in SERS-based microneedle sensors
Scientific Reports (2022)