Abstract
Little is known about how leukemia cells alter the bone marrow (BM) niche to facilitate their own growth and evade chemotherapy. Here, we provide evidence that acute myeloid leukemia (AML) blasts remodel the BM niche into a leukemia growth-permissive and normal hematopoiesis-suppressive microenvironment through exosome secretion. Either engrafted AML cells or AML-derived exosomes increased mesenchymal stromal progenitors and blocked osteolineage development and bone formation in vivo. Preconditioning with AML-derived exosomes ‘primed’ the animals for accelerated AML growth. Conversely, disruption of exosome secretion in AML cells through targeting Rab27a, an important regulator involved in exosome release, significantly delayed leukemia development. In BM stromal cells, AML-derived exosomes induced the expression of DKK1, a suppressor of normal hematopoiesis and osteogenesis, thereby contributing to osteoblast loss. Conversely, treatment with a DKK1 inhibitor delayed AML progression and prolonged survival in AML-engrafted mice. In addition, AML-derived exosomes induced a broad downregulation of hematopoietic stem cell-supporting factors (for example, CXCL12, KITL and IGF1) in BM stromal cells and reduced their ability to support normal hematopoiesis. Altogether, this study uncovers novel features of AML pathogenesis and unveils how AML cells create a self-strengthening leukemic niche that promotes leukemic cell proliferation and survival, while suppressing normal hematopoiesis through exosome secretion.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) is a malignant disease characterized by differentiation blockage and proliferation of clonal hematopoietic stem or progenitor cells (HSPCs), which rapidly lead to bone marrow (BM) failure and eventually to death, if left untreated. Even with optimal therapeutic approaches, including intensive chemotherapy and allogeneic hematopoietic stem cell (HSC) transplantation, only a minority of patients are cured. Thus, to design more effective therapeutic approaches and improve clinical outcome, it is critical to identify and pharmacologically target key molecular mechanisms that drive the pathogenesis of this disease.
During leukemogenesis, AML cells (or blasts) progressively occupy and likely alter the BM niche where normal HSCs reside.1 Emerging evidence suggests that leukemia cells induce molecular changes in distinct hematopoietic and non-hematopoietic cell populations in the niche. These changes contribute to the transformation of the normal hematopoietic niche into a ‘leukemia niche’ that becomes permissive of leukemia growth and disrupts normal hematopoiesis.2 In vivo perturbation of the murine BM microenvironment by deletion of the retinoic acid gamma receptor, retinoblastoma protein, Notch signaling or the microRNA (miRNA)-processing enzyme dicer has been shown to promote aberrant myeloproliferation.3, 4, 5, 6 Similarly, forced expression of β-catenin-activating mutants in osteoblasts induces leukemia in mice.7 Furthermore, leukemia cells reduce the niche’s ability to retain HSCs and normal hematopoietic activity.8, 9, 10 However, the fine mechanisms through which leukemia cells co-opt and modify the normal hematopoietic niche remain largely unknown.
Exosomes are small vesicles (30–200 nm) that are secreted by a wide variety of normal and malignant cells,11 and are increasingly recognized as key mediators of cell-to-cell communication.12, 13 Cancer-derived exosomes are capable of supporting cancer growth and disrupting homeostasis of healthy tissue.11, 14, 15, 16, 17, 18 Exosomes released from chronic myeloid leukemia (CML) cells, for example, can stimulate BM stromal cells to produce IL-8, a cytokine that supports leukemia growth.19 Exosomes secreted by AML cell lines and primary AML blasts enter stromal cells and modify their function to enhance leukemia growth.20, 21 However, most of the exosome studies in hematological malignancies are based on in vitro experiments and the direct contributions of AML-derived exosomes to leukemogenesis remain to be fully elucidated in vivo.
Here, we used models of human-to-mouse AML grafts and murine AML leukemia to demonstrate how AML-derived exosomes profoundly transform the BM niche into a microenvironment that permits leukemia growth and suppresses normal hematopoiesis.
Materials and methods
Mice
C57BL/Ka (B6), C57BL/Ka-Thy1.1-CD45.1 (B6-45.1), B6-Rag2−/−γc−/− (DKO) and NOD-scid-γc−/− (NSG) mice were maintained by the Animal Resource Center of City of Hope. Newly born and 8- to 12-week-old DKO or NSG mice of both sexes were sub-lethally irradiated (newborn:150 rad, adult: 250 rad) and used as recipients for AML cell transplantation. MLL-AF9 knockin mice were obtained from Jackson laboratory. B6 or B6-45.1 mice were used as recipients in HSC and whole BM transplantation assays. Mouse care and experimental procedures were performed in accordance with federal guidelines and protocols approved by the Institutional Animal Care and Use Committee at the City of Hope.
Cell lines and patient samples
AML cell lines (KG1A, MV411 and NB4; ATCC) were cultured in IMDM (Gibco, Grand Island, NY, USA) with 10% fetal bovine serum (FBS; Atlanta Biologicals, Flowery Branch, GA, USA). AML patient and control samples (peripheral blood or BM) were obtained under a City of Hope-Institutional Review Board—approved protocol with informed consent from patients or healthy individuals (see Supplementary Table 1 for more information). BM CD34+ cells were enriched using human CD34+ microbeads kit (Miltenyi Biotec, Auburn, CA, USA).
Antibodies
Detail information on antibodies used is available in Supplementary Table 3.
Isolation, culture and analysis of mesenchymal stromal progenitors
Tibias and femurs were excised from 8- to 12-week-old mice. After removing the muscle and connective tissue, the bones were crushed gently with mortar and pestle in PBS buffer. The crushed mass was washed with PBS to remove all the BM cells. The remaining bone fragments were incubated at 37 °C with collagenase I (3 mg/ml; Sigma, St Louis, MO, USA) and gently agitated for 40 min. The digested bones were then filtered with a 40 μm strainer (BD Biosciences, San Jose, CA, USA), pelleted at 200 g at 4 °C and resuspended in PBS. The marrow cells were collected and red blood cells were depleted with ACK lysis buffer. The cells were blocked with anti-CD16/32 antibody and stained with labeled monoclonal antibodies against CD45, CD31, Ter119, Sca1, CD146 and CD166 (Supplementary Figure 1d). The stained cells were then sorted using a FACSAria III sorter (BD Biosciences). The purity of sorted cells was confirmed by FACS analysis to be >99%. Sorted stromal cells from control and AML cells/exosomes-treated mice were cultured in the MEM-alpha medium (Gibco) containing 15% FBS at 37 °C with 5% CO2. For mesenchymal stromal progenitor analysis, at least 3000 events per sample were collected within the CD45-Ter119-CD31-gate and analyzed using FlowJo software (FlowJo, Ashland, OR, USA).
Isolation and analysis of long-term HSCs
HSPCs were enriched from red blood cell-lysed BM cells using c-Kit microbeads kit (Miltenyi Biotec). The enriched HSPCs were then blocked with anti-CD16/32 antibody, stained with labeled antibodies against c-Kit, Sca1, CD150, CD48, CD135 and lineage markers (CD3, CD4, CD8, CD11b, B220, CD19, Gr1 and Ter119) and double-sorted using a FACSAria III to isolate long-term HSCs (LT-HSC) (Supplementary Figure 1c). For LT-HSC analysis, at least 1000 events per sample were collected within the lin-Sca1+c-Kit+ (LSK) gate and analyzed using FlowJo software (FlowJo).
LT-HSC and stromal cells in vitro co-culture assay
Overall, 1000 sorted Sca1+ stromal cells from AML cells per exosomes-treated or control CD45.2+ mice were co-cultured with 500 CD45.1+ mouse LT-HSCs in IMDM medium (Gibco) containing 10% FBS, 5 ng/ml mouse KITL and TPO (Peprotech, Rocky Hill, NJ, USA) for 48 h. Harvested CD45.1+ HSCs were then transplanted into lethally irradiated (1100 rad) CD45.2 recipient mice (100 cells equivalents per mouse) along with 200 000 CD45.2+ helper marrow cells. Peripheral blood was drawn at indicated time points and the contribution of donor-derived cells was analyzed by FACS analysis.
Stromal cell and AML cells/derived exosomes co-culture assay
A total of 10 000 Sca1+ stromal cells were co-cultured with 10 000 AML cells per exosomes (derived from 5 × 106 cells). After 48-h culture, stromal cells were harvested and analyzed for mRNA levels of different niche-associated genes.
Exosome isolation, characterization and labeling
Culture media or FBS was centrifuged at 100 000 g for 10 h to remove bovine exosome contamination from FBS. Centrifuged media was filtered using a 0.2 μm filter to avoid the FBS exosomes pellet, then collected in a flask and used for culturing the cells.
For exosome isolation from leukemic and normal hematopoietic cells, AML cell lines were cultured directly with the centrifuged media. For normal hematopoietic cells or primary AML patient samples the media were supplemented with 2 ng/μl Human TPO and KITL (Peprotech). Supernatants were collected 48 h later, centrifuged at 800 g for 10 min at 4 °C to remove whole cells, centrifuged again at 10 000 g for 10 min at 4 °C to remove cellular debris and filtered through a 40-μm membrane, prior to exosome preparation by centrifugation at 100 000 g for 1 h. The exosome pellet was washed twice in a large volume of PBS (Supplementary Figure 2c).
Nanoparticle tracking analysis was used to determine exosome concentration. The plasma samples were diluted 1:10 and visualized on the NanoSight NS300 nanoparticles detector instrument (Malvern, Westborough, MA, USA). For exosome protein quantification, the exosome pellet was dissolved in RIPA buffer and quantified using BCA Protein Assay Kit (Thermo Fisher Scientific, Rockford, IL, USA). For western blot analysis of exosomes, anti-TSG101, anti-CD63 and anti-β-actin antibodies were used.
For in vitro and in vivo exosome tracing, exosomes were labeled using CFSE (10 μM; Thermo Fisher Scientific) in PBS for 30 min at 37 °C. Labeled exosomes were washed with PBS and centrifuged 100 000 g for 1 h to remove excess dyes.
Tissue section and immunofluorescence staining
Freshly dissected long bones were fixed in 4% paraformaldehyde for 20 min at 4 °C, then decalcified in 10% EDTA at 4 °C for 7 days. The bones were then embedded in 10% gelatin in PBS, frozen on dry ice and stored at −80 °C. Bones were sectioned using the CryoJane taping system (Instrumedics, Ann Arbor, MI, USA). Sections were stained with Masson’s trichrome.
For immunofluorescence staining, sections were blocked with 1% BSA, then probed with primary antibody at 4 °C overnight: anti-Sca1, anti-CD146, anti-MECA32 and/or anti-osteocalcin (OCN). After washing with PBS, the sections were probed with Alexa dye-conjugated antibodies, Invitrogen (Carlsbad, CA, USA) goat anti-rat-555 or anti-rat-488 and anti-rabbit-555, then washed and stained with DAPI and mounted in ProLong Gold anti-fade mounting media (Molecular Probe, Eugene, OR, USA). Sections were analyzed using an Olympus IBX81 microscope (Olympus, Waltham, MA, USA). Slides that passed this initial inspection were scanned using a Zeiss LSM 700 confocal microscope (Zeiss, Thornwood, NY, USA). Zen 2012 analysis software was used to create the maximum intensity projections of confocal z-stacks.
Micro-computed tomography examination
The bones were scanned using an Inveon CT scanner (Siemens, Malvern, PA, USA) under the following specifications: total rotation: 360; magnification: medium-high; binning: 1; transaxial CCD size: 1120 px; axial CCD size: 1408 px; pixel size: 13.60 μm; voltage: 80 kV; current: 500 μA; and exposure time: 4000 ms. The files were then reconstructed using COBRA with no downsampling. Inveon Bone Morphometry Tools (Siemens) were used to analyze the data.
Fluidigm dynamic array
RNA was extracted from 100 double-sorted stromal cells, reverse transcribed and amplified. The resulting amplified cDNAs were analyzed by qPCR using the Biomark Dynamic Array (Fluidigm, South San Francisco, CA, USA). We reviewed the amplification curves and CT data using the Biomark system software suite (Fluidigm). Delta CT values were calculated by subtracting the CT value for a particular gene from the paired β-actin CT value. Fold-change values were calculated in a linear scale based upon the least-squares mean. The TaqMan probes (Applied Biosystems, Carlsbad, CA, USA) used are listed in Supplementary Table 2.
Quantitative real-time PCR
RNA was extracted from sorted cells using the RNeasy RNA isolation kit (Qiagen, Germantown, MD, USA) and was reverse transcribed into cDNA using the Sensiscript RT kit (Qiagen). The cDNA was amplified for specific targets using TaqMan assays on a ViiA 7 Real-Time PCR System (Applied Biosystems). The thermal cycling parameters were 40 cycles of 95 °C for 15 s and 60 °C for 1 min. Data were analyzed using the ViiA 7 software (Applied Biosystems). The TaqMan probes used are listed in Supplementary Table 2.
In vitro lineage differentiation of stroma progenitors
For osteoblast differentiation, freshly sorted Sca1+ cells were cultured in MEM-alpha medium containing 15% FBS with the addition of 10 mm glycerol phosphate, 50 μm ascorbate-2-phosphate and 10−7 m dexamethasone at 37 °C with 2% O2 conditions for 3 weeks. Chondrocyte and adipocyte differentiation were performed using the StemXVivo system (R&D Systems, Minneapolis, MN, USA). Recombinant mouse DKK1 was obtained from R&D systems. DKK1 inhibitor Way-262611 was purchased from Axon Medchem (Reston, VA, USA).
LSK and LT-HSC cell cycle analysis
For cell cycle analysis, double-sorted LSK and LT-HSC cells were fixed in 2% formaldehyde and permeablilized in 70% methanol. Cells were stained with FITC-labeled anti-mouse Ki67 and propidium iodide (Invitrogen). The cutoff range for fluorescence was determined by a negative control sample stained with FITC-labeled isotype control Ig. At least 500 cells per sample were examined for each specimen and analyzed using FlowJo software (FlowJo).
Lentiviral shRNA transduction
Active lentiviral stocks were generated using human Rab27a-specific shRNA constructs (GE Dharmacon, Lafayette, CO, USA; clone ID: V3THS_300918) and control scrambled GFP vector, and used to transduce MV411 cells for exosome release inhibition. The transduced GFP-positive cells were analyzed for Rab27a mRNA expression levels using qPCR to confirm the knockdown.
OCN ELISA
Plasma OCN levels were determined using Quantikine ELISA kit (R&D systems). Frozen plasma samples from AML patients and healthy age-matched individuals were thawed and quantified in duplicate at 450 nm wavelength.
Statistical analysis
Statistical analyses were performed using Prism software v.7 (GraphPad, La Jolla, CA, USA). All data are presented as mean±s.d. Two group comparisons were performed with a two-tailed Student’s t-test; multiple group comparisons were performed with a one-way ANOVA test with multiple comparison option. For Kaplan–Meier survival curve analysis, the comparisons were performed using a log-rank test. A value of P<0.05 was taken as statistically significant. No statistical test was used to determine the sample size. No randomization was used to allocate animals to particular groups; age and sex-matched recipients were used for transplantation experiments. The investigator was not blinded to experimental groups during analysis.
Results
AML engraftment alters bone architecture and cellular composition of the BM microenvironment
To assess AML-induced changes in the BM microenvironment, we first injected human AML cells intrahepatically into newborn (2–3 days old) irradiated immunodeficient RAG2−/−γc−/− (DKO) mice.22 This approach allows for a rapid generation of in vivo AML models to study the interplay of AML blasts and the BM niche in B6 genetic background mice, the main strain used for BM niche component analyses. For these experiments, we injected human primary AML blasts (isolated from AML patients seen at our institution, COH101 or COH103) or cell lines representative of relatively common genetic mutations occurring in human AML: MV411 (FLT3-ITD), KG1A (p53-null) or NB4 (PML-RARα). To confirm our results, we also engrafted AML blasts from the MLL-AF9 knockin AML mouse model23, 24 into 8-week-old irradiated syngeneic recipients. Irradiated, age-matched, vehicle-injected mice or normal human CD34+ BM-injected served as controls. In this series of experiments, mice were killed 3–6 weeks after cell injections, upon evidence of leukemia development, or earlier if the animal was moribund.
All AML recipient mice developed leukemia and exhibited splenomegaly, compared with vehicle- or normal human CD34+ BM-injected controls (Figure 1a; Supplementary Figures 1a–c). We also observed an increased number of endogenous long-term HSCs (LT-HSCs; Lin-Sca1+c-Kit+CD150+CD48−CD135−; Supplementary Figure 1d) in the BM of recipient mice engrafted with human or mouse AML cells, compared with vehicle-injected controls (Figure 1b; Supplementary Figure 1e). The number of endogenous LT-HSCs was not changed in mice engrafted with normal human CD34+ cells compared with vehicle-injected controls (Supplementary Figure 1f).
AML engraftment results in expansion of LT-HSCs, altered stromal compartments, reduced osteoblast number and bone architecture changes. (a) Neonatal DKO mice were injected with vehicle control or 2 million AML cells (MV411, KG1A, NB4 cell lines or COH101, COH103 patient samples) intrahepatically. The litter was killed when transplanted mice were moribund or 4 weeks after transplantation. Spleen weight of control and AML-transplanted (MV411, KG1A, COH101) DKO mice (n=5–7 mice per group in at least three independent experiments). (b) Frequency of the LT-HSC population in marrow of control and AML-transplanted (MV411, KG1A, NB4, COH101) DKO mice (n=5–8 mice per group in at least three independent experiments). (c) Frequency of Sca1+, CD146+ and CD166+ cells in the stromal compartment of control and AML-transplanted DKO mice (n=7–13 mice per group in at least three independent experiments). (d) Trichrome stain of bone sections from control and AML-transplanted (MV411, KG1A) DKO mice. Osteoblasts are marked with yellow arrows. (e) Immunofluorescence staining of bone sections from control or MLL-AF9-transplanted B6 mice. Staining of Sca1 and OCN (left) are shown. Osteoblasts are marked with yellow arrows. Number of Sca1+ cells in the view field determined by immunofluorescent staining (right). (f) Representative μ-CT 2D cross-section of trabecular bone region in femurs of control or AML-transplanted (KG1A, MV411) DKO mice. (g) Femurs of control or AML-transplanted (KG1A, MV411) DKO mice (n=6–8 in two independent experiments). Quantitative μ-CT analysis of cortical wall thickness in compact bone region, relative bone volume (BV/TV), thickness of trabecular bone, number of trabeculae and space between trabeculae in trabecular bone region. *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001.
To analyze the non-hematopoietic compartments of the BM niche, we then enzymatically dissociated bone and marrow and isolated the stromal fraction based on surface markers (CD45−Ter119−CD31−). The stromal fraction was further divided based on CD146, CD166 and Sca1 expression into CD146+CD166− immature osteoprogenitors (CD146+); CD146−CD166+ osteoblasts (CD166+); and CD146−CD166−Sca1+ primitive stromal cells with the ability to differentiate into CD146+, CD166+ or CXCl12-abundant reticular (CAR) cells (Sca1+; Supplementary Figure 1g).25 FACS analyses revealed that Sca1+ and CD146+ cells were significantly increased, and CD166+ cells were significantly decreased, in all models of AML-engrafted mice, compared with vehicle-injected controls (Figure 1c; Supplementary Figure 1h). In contrast, the numbers of CD166+, CD146+ and Sca1+, cells were not significantly changed in mice engrafted with normal human CD34+ cells, compared with vehicle-injected controls (Supplementary Figure 1i).
Histological analysis and immunofluorescence staining of bone sections with anti-OCN antibody confirmed the loss or thinning of the osteoblast layer in AML-engrafted animals, but not in vehicle-injected controls (Figures 1d and e). Immunofluorescence staining of bone sections with anti-Sca1 and anti-CD146 antibodies confirmed the increase of Sca1+ and CD146+ cells (Figure 1e; Supplementary Figure 1j). The number of Sca1+ and CD146+ stromal cells located below the growth plate of the sectioned femurs was also significantly increased in AML-engrafted mice compared with controls (Sca1+: 22.25 vs 7.75 cells per 66 mm2, P=0.0001; CD146+: 6.44 vs 3.50 cells per 102 mm2, P=0.041; Figure 1e; Supplementary Figure 1j). Furthermore, micro-computed tomography (μ-CT) examination showed that cortical wall thickness, as well as trabecular bone volume, decreased in leukemic animals (Figure 1g). Decreased trabecular bone volume resulted in a larger trabecular area in leukemic animals (Figures 1f and g). These data corroborate our FACS stromal compartment analysis (Figure 1c; Supplementary Figure 1h) and together support the notion that AML growth modifies the cellular composition of both the hematopoietic and non-hematopoietic compartments in the BM niche.
AML-derived exosomes target stromal and endothelial cells in the BM niche
Exosomes are increasingly recognized as key mediators of cell-to-cell communication and are aberrantly increased in certain types of cancer.12, 13, 26 To determine whether circulating exosomes were increased in AML patients, we analyzed plasma samples from 43 de novo or secondary AML patients seen at our institution (COH107–COH191; Supplementary Table 1). Using nanoparticle tracking analysis,27 we found that plasma exosomes were significantly increased in AML patients (n=43) compared with healthy controls (n=12; 4.0 vs 1.6 × 109/ml, P=0.0016; Figure 2a). Of note, consistent with our observation that osteoblast activities were reduced in AML-engrafted mice (Figure 1), we found significantly lower plasma levels of OCN, an indicator of bone formation activity, in AML patients than in healthy controls (17.3 vs 6.9 ng/ml, P<0.0001; Figure 2a). Exosome counts were also significantly higher in patients with lower OCN levels (<10 ng/ml) compared to those with higher OCN levels (>10 ng/ml, Figure 2b). These changes were not specific to the FLT3-ITD mutation or risk stratification in the analyzed AML blasts (Supplementary Figures 2a and b).
AML-derived exosomes are elevated and associated with reduced OCN levels in AML patients, are internalized by BM cells, induce splenomegaly, increase LT-HSC population and alter stromal compartment in the BM microenvironment. (a) Plasma exosome numbers measured by NanoSight (left) and plasma OCN levels were determined by ELISA (right) in healthy control (n=12) and AML patients (n=43). (b) Plasma exosome count in AML patients with low (<10 ng/ml, n=29) or high (>10ng/ml, n=14) plasma OCN levels. (c) Overall, 100 μg CFSE-labeled exosomes were co-cultured with BM cells for 4 h. Uptake of CFSE-labeled exosomes by marrow cells was analyzed by fluorescence microscopy. (d) Uptake of CFSE-labeled exosomes by marrow cells was analyzed by FACS analysis for internalization of CFSE-labeled exosomes in indicated BM subpopulations (left). Percent CSFE+ cells within indicated BM subpopulations were summarized (right, n=3 mice in three independent experiments). (e) Six- to eight-week-old B6 mice were injected with either normal human PBMC-derived or AML-derived exosomes. Mice were euthanized 30 days after initial injection. (f) Mouse spleen is weight is summarized (n=5–6 mice per group in at least two independent experiments). (g) Frequency of LT-HSC in BM (n=6–8 mice per group in at least three independent experiments). (h) Frequency of Sca1+, CD146+ and CD166+ cells in the stromal compartment (n=6–8 mice per group in at least three independent experiments). ns, not significant, *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001.
Thus, we hypothesized that AML cells remodel the BM microenvironment partly through exosome secretion. To test this hypothesis, exosomes were prepared from the supernatant fractions of normal human peripheral blood mononuclear cell (PBMC) or AML cell culture by filtration and high-speed centrifugation28 (Supplementary Figure 2c). The purity of the prepared exosomes were then assessed by nanoparticle tracking analysis (Supplementary Figures 2d and e) and the exosome presence was confirmed by western blotting for the exosome markers CD63 and Tsg101 (Supplementary Figure 2f) and by electron microscopy, which documented vesicles of 30–100 nm in diameter (Supplementary Figures 2g and h), consistent with previous reports.13
We labeled the exosomes with the fluorescent dye CFSE and incubated them with whole BM freshly isolated from a wild-type B6 mouse (Figures 2c and d). Internalization of the labeled exosomes into mouse BM cells was demonstrated after 4 h (Figure 2c). FACS analysis revealed that the exosomes were primarily internalized in the BM stromal (CD45−Ter119−CD31−) and endothelial (CD45−Ter119−CD31+) fractions. In contrast, few exosomes were internalized in the hematopoietic (CD45+/Ter119+) fraction, except for a small subset of monocytes (CD11b+Gr1−; Figure 2d). Next, we injected CFSE-labeled exosomes intravenously into B6 mice and harvested their BM cells 10 h later. Again, internalized exosomes were found mainly in the BM stromal and endothelial fractions (Supplementary Figure 2i), suggesting that these cell populations are indeed the preferential BM targets for AML-derived exosome.
AML-derived exosomes induce similar changes to those induced by AML cell engraftment in the BM microenvironment
Having determined the main BM target cells for AML-derived exosomes, next we asked what functional changes were induced by AML-secreted exosomes. Thus, 6- to 8-week-old B6 mice were intravenously injected once/week for four consecutive weeks with 25 μg of exosomes (100 μg in total) isolated from either AML cells or normal human PBMC cultures. The mice were euthanized on day 30, 2 days after the final injection (Figure 2e).
Animals that received AML-derived exosomes showed changes similar to those observed in mice engrafted with AML cells, compared with controls treated with normal PBMC-derived exosomes, including enlarged spleens (Figure 2f; Supplementary Figure 3a) and increased endogenous BM LT-HSC (Figure 2g; Supplementary Figures 3b and c). Like in AML-engrafted mice, FACS analysis showed Sca1+ and CD146+ stromal cells were significantly increased and CD166+ cells were significantly decreased in mice treated with AML-derived exosomes, compared with PBMC-derived exosome-treated controls, suggesting a block in osteoblast differentiation (Figure 2h). Quantitative μ-CT analysis demonstrated loss of trabecular and cortical bone in the femurs of mice treated with AML-derived exosomes, compared with PBMC-derived exosome-treated controls (Supplementary Figure 3d). In vivo PBMC-derived exosome treatment did not induce significant changes in the stromal compartments compared with vehicle-injected controls (Supplementary Figure 3e).
Next, we focused on AML-induced gene expression changes in the BM stromal compartment. Sca1+, CD146+ and CD166+ stromal cells from DKO mice engrafted with AML cell lines (MV411 and KG1A) or primary patients’ blasts (COH101 and COH105) were sorted and analyzed for expression levels of hematopoiesis- and osteogenesis-related genes. Genes that modulate normal hematopoiesis (IGF1, CXCL12, KITL and IL-7) or are implicated in bone development (IGF1, OCN and Col1A1) were downregulated, whereas a suppressor of normal hematopoiesis and osteoblast development (DKK1),29 was upregulated in stromal cells from AML-engrafted vs vehicle-injected control animals (Supplementary Figure 4a). Despite some variations among cell lines and primary samples, a similar trend was observed in all models. These results further validate the observation that osteoblast maturation is disrupted in AML-engrafted mice (Figures 1e–j). Importantly, similar trends of gene expression were also observed in Sca1+, CD146+ and CD166+ stromal cells isolated from mice treated with AML-derived exosomes (Figure 3a).
AML-derived exosomes modulate gene expression in BM stroma and suppress osteogenic differentiation of mesenchymal stromal progenitors; the block is reversed by DKK1 inhibitor. (a) qRT-PCR showing expression levels of indicated genes (n=4–12 in at least three independent experiments). BM Sca1+, CD146+ or CD166+ stromal cells from B6 mice treated with normal PBMC-derived or corresponding AML-derived exosomes. (b, c) Sorted Sca1+ stromal cells from BM of normal B6 mice were treated with normal PBMC-derived or AML (MV411, KG1A)-derived exosomes. Treated cells were divided and cultured in osteogenic, adipogenic or chondrogenic conditions for 21 days. Alizarin Red staining (b) and OCN expression (c) in stroma after osteogenic induction. (d) qRT-PCR of OCN expression of sorted Sca1+ cells from BM of normal B6 mice were cultured with PBMC-derived or AML (MV411)-derived exosomes, with or without DKK1 inhibitor (Way-262611). (e) NSG mice were injected with four doses of DKK1 inhibitor (Way-262611) or DMSO prior to AML transplantation and one dose after. Kaplan–Meier survival curve analysis of treated mice (lower, n=12–13 mice per group in two independent experiments). ns, not significant, *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001.
To validate these results, we treated ex vivo Sca1+, CD146+ and CD166+ cells from B6 mice with AML-derived exosomes. We observed similar downregulation of OCN, Col1A1 and IGF1 as seen in AML-engrafted mice. However, no significant changes were observed in KITL and CXCL12 expression, except for CXCL12 downregulation in CD146+ cells (Supplementary Figure 4b). Levels of genes encoding pro-inflammatory cytokines (that is, IL-6 and CCL3) were also significantly increased in AML exosome-treated CD146+ and CD166+ cells (Supplementary Figure 4b), but not in the stromal cells from leukemic or AML exosome-treated mice (data not shown). Differences between in vivo and ex vivo environments, dosage of AML-derived exosomes (four vs one doses) and time after initial treatments with exosomes (48 h vs 30 days) may account for the differences in outcomes. Similar results were obtained when we cultured AML cell lines (MV411 and KG1A) and primary patient blasts (COH101 and COH103) or AML- or PBMC-derived exosomes with normal human BM stromal cells. IL-6 increased and CXCL12, OCN and Col1A1 similarly decreased in human stromal cells cultured with AML cells or AML-derived exosomes, compared with those cultured with PBMC-derived exosomes or vehicle (Supplementary Figures 4c and d).
Collectively, these data indicate that AML-derived exosomes induced significant changes in the cellular composition of the BM niche similar to those observed in AML, even in the absence of AML cells. This supports our hypothesis that exosomes may mediate the BM changes observed during AML growth.
AML cells and AML-derived exosomes induce DKK1 expression and decrease osteoblast lineage differentiation potential of mesenchymal stromal progenitors
To understand how AML cells and AML-derived exosomes reduce osteoblast formation in the BM niche, we performed in vitro lineage differentiation assays. Sca1+ cells were isolated from DKO mice engrafted with NB4 cells or primary sample COH101 upon evidence of leukemia (~25–30 days after transplantation or earlier if the animals were moribund) and cultured under osteogenic conditions for 21 days. These cells exhibited reduced osteoblast formation compared with Sca1+ cells isolated from vehicle-injected controls as evidenced by Alizarin Red staining (Supplementary Figure 5a) and qRT-PCR for OCN (OCN: 1.03 vs 0.08-fold, P=0.0038; Supplementary Figure 5b).
To assess whether AML-derived exosomes have a similar impact on differentiation of mesenchymal stromal progenitors, we treated Sca1+ cells from B6 mice with AML-derived or PBMC-derived exosomes for 20 h. Treated cells were divided and cultured in osteogenic, adipogenic or chondrogenic conditions for 21 days. Alizarin Red staining and qRT-PCR for OCN, PPARγ (adipogenic marker) and aggrecan (Acan; chondrogenic marker) indicated that Sca1+ cells treated with AML-derived exosomes acquired an increased ability to differentiate into adipocytes, but a decreased ability to differentiate into osteoblasts and chondrocytes, compared with Sca1+ cells treated with PBMC-derived exosomes (OCN: 1.02 vs 0.55-fold, P=0.018; PPARγ: 1.0 vs 4.01-fold, P=0.0122; ACAN: 1.01 vs 0.07-fold, P=0.0233; Figures 3b and c; Supplementary Figures 5c and d).
As DKK1, a potent negative regulator of osteoblast development, was overexpressed in the BM niche of AML-engrafted and AML-derived exosome-treated mice (Figure 3a; Supplementary Figure 4a), we hypothesized that elevated levels of DKK1 may explain the limited osteoblast differentiation seen in the Sca1+ cells from those mice. To test this hypothesis, we cultured ex vivo Sca1+, CD146+ and CD166+ cells from B6 mice with AML-derived or PBMC-derived exosomes. DKK1 expression was upregulated in all BM stromal cells treated with AML-derived exosomes compared to PMBC-derived exosomes (Sca1+: 15.69-fold, P<0.0001; CD146+: 3.8-fold, P=0.0282; CD166+: 11.04-fold, P<0.0001; Supplementary Figure 5e).
Furthermore, Sca1+ cells isolated from B6 mice and cultured under osteogenic conditions with 10 ng/ml of recombinant DKK1 showed no ability to differentiate into osteoblasts (Supplementary Figures 5f and g). To evaluate the ability of DKK1 inhibition to rescue the changes induced by AML cells or AML-derived exosomes, we treated Sca1+ cells from B6 mice with AML-derived or PBMC-derived exosomes in the presence or absence of the DKK1 inhibitor Way-262611 for 20 h. The treated cells were then cultured under osteogenic conditions for 21 days. Osteoblast differentiation was not changed by DKK1 inhibitor in PBMC-exosome-treated Sca1+ cells (OCN: 1.29 vs 0.95-fold, P=0.1546). In contrast, DKK1 inhibitor rescued the decreased osteoblast differentiation induced by AML-derived exosome (OCN: 1.14 vs 0.3-fold, P=0.0018; Figure 3d; Supplementary Figure 5h).
Next, we hypothesized that reversal of the effects of AML cells and AML exosomes on BM niche by in vivo DKK1 inhibition may delay AML progression. Eight- to ten-week-old NSG mice were engrafted with 1 million MV411 AML cells and received four intraperitoneal doses of DKK1 inhibitor (35 mg/kg body weight) or DMSO (vehicle control) pre-engraftment, and one dose post-engraftment. Mice engrafted with AML cells and treated with DKK1 inhibitor survived significantly longer than mice engrafted with AML cells but treated with vehicle only (median survival: 64 vs 50 days, P=0.0067; Figure 3e), suggesting that AML engraftment- or AML-derived exosome-induced DKK1 expression likely suppressed osteoblast differentiation and supported enhanced leukemia growth.
AML-modified stromal cells exhibit reduced ability to support normal hematopoiesis and enhance leukemia growth
One relatively unexpected finding in mice engrafted with AML cells or treated with AML-derived exosomes was the increased percentage of endogenous LT-HSCs (Figures 1b and 2g; Supplementary Figures 3b and c). Thus, we asked whether these LT-HCSs were functionally equivalent to those that support normal hematopoiesis in normal mice. We compared the cell cycle of LT-HSCs from B6 mice treated with AML-derived exosomes with LT-HSCs from mice treated with PBMC-derived exosomes (Supplementary Figures 6a and b). In LT-HSCs from mice treated with AML-derived exosomes, a reduced percentage of cells were in G0 phase (Supplementary Figure 6c), whereas an increased percentage of cells were in S/G2/M phase (Supplementary Figure 6d), compared with LT-HSCs from mice treated with PBMC-derived exosomes.
To assess the hematopoietic repopulating activity in mice treated with AML-derived exosomes, we transplanted whole BM cells from exosome-treated B6-45.1 mice into irradiated B6 (45.2) congenic mice. Mice transplanted with whole BM cells from donors treated with AML (KG1A)-derived exosomes showed lower blood and BM engraftment at 22 weeks after transplant, compared to controls transplanted with whole BM cells from mice treated with PBMC-derived exosomes (Figure 4a). In addition, we used the S phase-specific cytotoxic chemotherapeutic agent 5-fluorouracil (5-FU) to induce myeloablative stress. Mice were treated with three weekly doses of AML- or PBMC-derived exosomes, before receiving weekly injections of 5-FU (150 mg/kg body weight) until death (Supplementary Figure 6e). AML-derived exosome-treated mice showed reduced survival compared with PBMC-derived exosome-treated mice (median survival: 12.5 vs 15.5 days, P=0.0005; Supplementary Figure 6f), thereby indicating a compromised hematopoietic recovery after myeloablative stress.
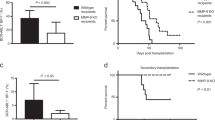
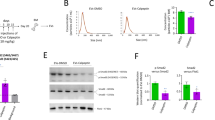
AML-derived exosomes reduce stem cell activities, limit ability of stroma to support normal HSCs and accelerate AML progression; disruption of exosome production delays AML progression. (a) Repopulation ability of marrow from normal PBMC-derived or AML (KG1A)-derived exosome-treated donors was assayed 22 weeks after transplantation into congenic recipients (n=5–6 mice per group in two independent experiments). (b) CD45.1 HSCs were co-cultured with the AML cell/exosome-modified stroma for 48 h and transplanted (100 per mouse) into lethally irradiated WT CD45.2 recipients along with 200 000 unfractionated CD45.2 helper BM cells. Ability of control and AML cell/exosome co-cultured HSCs to repopulate the blood of recipient mice was assayed 19 weeks after transplantation (n=6–8 mice per group in three independent experiments). (c) Three weekly doses of AML-derived or normal PBMC-derived exosomes, or PBS vehicle control, were injected intravenously into 6- to 8-week-old DKO mice. Mice were irradiated and injected with 20 million KG1A cells. AML cell engraftment in peripheral blood was assayed 20 days later (middle). Kaplan–Meier survival analysis of control or treated mice (right; n=8–13 mice in at least two independent experiments). (d) MV411 AML cells were transduced with lentiviral shRNA against Rab27a, a protein involved in exosome release. qRT-PCR of Rab27a levels (left) and total exosome protein quantification (middle). Kaplan–Meier survival curve for mice transplanted with lentiviral shRab27a-transduced MV411 cells (right; n=8–10 mice in two independent experiments). ns, not significant, *P<0.05, **P<0.01, ***P<0.001.
To determine whether AML cells or AML-derived exosomes functionally altered the BM niche’s ability to support normal HSCs, we incubated sorted Sca1+ stromal cells from B6 mice with KG1A and NB4 cells, primary AML patient blasts (COH101), or AML- or PBMC-derived exosomes for 2 days, followed by co-culture with LT-HSCs from B6-45.1 mice for 2 additional days. These LT-HSCs were then transplanted into B6 congenic recipients and hematopoietic activity of the grafts was evaluated (Figure 4b). Mice receiving LT-HSCs co-cultured with AML-treated or AML-derived exosome-treated Sca1+ cells showed a significantly decreased engraftment rate at 16 weeks post-transplantation, compared with controls that received LT-HSCs co-cultured with PBMC-derived exosome-treated or untreated Sca1+ cells (CD45.1: 4.84 vs 17.82 or 19.94% engraftment, P<0.0001; Figure 4b).
Next, we asked whether preconditioning the BM niche using AML-derived exosomes could facilitate AML growth. Mice pretreated with weekly doses of AML-derived exosomes for 3 weeks were then injected with KG1A cells and analyzed for leukemia engraftment. These mice showed a significantly higher rate of leukemia cell engraftment at 20 days (human CD45+: 13.17 vs 1.1 or 1.08% engraftment, P=0.0486 or 0.0363) and a shorter survival than controls pretreated with weekly doses of PBMC-derived exosomes or vehicle before transplant (median survival: 22.5 vs 30 or 35 days, P=0.0083 or 0.0075; Figure 4c). FACS analysis of the stromal compartment showed that populations of Sca1+ and CD146+ stromal cells significantly increased (Sca1+: 4.32 vs 3.17% of total stromal cells, P=0.0113; CD146+: 4.26 vs 3.03% of total stromal cells, P=0.0166) and CD166+ stromal cells significantly decreased (1.91 vs 2.44% of total stromal cells, P=0.0276) in mice that received AML-derived exosomes, compared with mice that received PBMC-derived exosomes (Supplementary Figure 7a).
To test whether AML-derived exosomes are required for leukemia growth, we reduced exosome secretion in AML cells. We used a lentiviral shRNA construct to knockdown Rab27a, a protein implicated in the exosome release process,30 in MV411 AML cells. Exosome secretion was significantly reduced in Rab27a-knockdown cells compared with scrambled shRNA controls (Figure 4d), without any significant changes in the cell cycle (Supplementary Figure 7b) or baseline apoptosis (Supplementary Figure 7c) of the transduced cells. Two million Rab27a-knockdown or scrambled control AML cells were then intrahepatically injected into 2- to 3-day-old DKO mice. The mice that received shRab27a-knockdown AML cells survived longer than mice that received scrambled control AML cells (median survival: 28.5 vs 24 days, P=0.0016; Figure 4d), indicating that reduced exosome secretion in AML cells delayed AML growth.
Discussion
The functional relevance of cancer-derived exosomes to tumor growth, metastasis and treatment response has become increasingly evident.16, 17, 18, 31 Using a combination of in vivo and in vitro approaches, we demonstrate here that AML-derived exosomes are important for transforming the composition and function of the BM niche. Treatment with exosomes derived from a variety of human and mouse AML cells recapitulates changes similar to those observed in mice engrafted with AML cells. Specifically, AML-derived exosomes preferentially target BM stromal and endothelial cells and alter the differentiation potential of mesenchymal progenitors, thereby decreasing osteoblast development. Histologically, these cellular changes result in reduced cortical bone wall thickness and decreased trabecular bone volume and trabeculation. In line with these results, we showed decreased plasma levels of OCN in AML patients. Furthermore, as in AML engraftment experiments, treatment of BM stromal cells with AML-derived exosomes decreases the expression of genes supporting normal hematopoiesis (CXCL12, KITL, IL-7, IGF1) and osteogenesis (OCN, Col1A1, IGF1) and increases expression of genes supporting AML growth (DKK1, IL-6, CCL3). Preconditioning BM with AML-derived exosomes accelerates AML cell engraftment and growth, whereas disruption of exosome secretion in AML cells significantly decreases the ability of these cells to engraft and grow in vivo. Similar to the effect of AML engraftment, treatment with AML-derived exosomes also increases the fraction of normal LT-HSC engaged in cell cycling, and impairs normal hematopoietic reconstitution. Our findings, therefore, indicate that AML cells induce exosome-dependent cellular changes in the hematopoietic and non-hematopoietic compartments of the BM niche that lead to attenuation of normal hematopoiesis while enhancing leukemia growth. Our observations are supported by previous studies that suggest tumor-derived exosomes are key players in vascular leakiness and stromal cell education at metastatic sites.17, 32
The role that AML cells and AML-derived exosomes have in decreasing osteoblast differentiation is particularly intriguing to us, as it may be pharmacologically targetable. Consistent with our data, a recent study indicated suppression of the osteoblast population and osteoblast-associated gene signature through aberrant Notch signaling occurs in acute lymphoblastic leukemia.33 Our findings are also consistent with previous reports demonstrating severe osteoblastic defects in AML mouse models,34 and in AML and MDS patients.35 Genetic ablation of osteoblasts in AML or CML mice was correlated with increased progression of leukemia,35, 36 whereas treatment of AML mice with compounds capable of inhibiting osteoblast loss decreased tumor burden and prolonged survival.35 Hanoun et al.24 reported a similar blockade of osteoblastic differentiation in an MLL-AF9 AML model, which was associated with neuropathic damage in the BM. In contrast, increased trabeculation, trabecular thickening and a massive expansion in osteoblast lineage cells have been observed in a myeloproliferative neoplasm model.9 Krause et al.23 found that activating the parathyroid hormone receptor in osteoblasts caused opposite effects in CML and AML models, with decreased disease for CML mice and increased disease in AML mice. Thus, it is possible that the BM niche is affected differently in distinct types of hematologic malignancies. Although there are only limited reports that adult patients with AML have osteopenia and osteoporosis before allogeneic BM transplantation,37 it is known that bone loss and increased fracture risk are common complications for AML patient after transplantation.38, 39 Mesenchymal progenitors reprogrammed by AML-derived exosomes may contribute to observed bone defects in these patients. Mitigation of osteolineage differentiation defects may provide a novel therapeutic strategy to accelerate patient recovery. More studies are required to determine the exact clinical relevance of our observation.
Gene expression analysis indicates that DKK1, an osteoblast inhibitor and a negative regulator of Wnt signaling that is central to both hematopoiesis and bone development,40 is upregulated in our AML and AML-exosome models. Forced overexpression of DKK1 in osteoblasts results in a phenotype similar to what we observed in AML-engrafted and AML-derived exosome-treated mice—blockage of osteolineage development, disruption of the BM niche and defects in HSC function.41 Using a lineage differentiation assay, we demonstrated that mesenchymal stromal progenitors treated with AML-derived exosomes or from AML mice have an increased adipogenic and decreased osteogenic and chondrogenic differentiation ability. Treatment with DKK1 inhibitors reverses the effect of AML exosomes on stromal lineage differentiation. These findings are in line with previous reports that DKK1 can induce mesenchymal stromal cell proliferation and switch their differentiation pathway from osteogenesis to adipogenesis.42, 43, 44, 45, 46 Moreover, elevated DKK1 levels were also implicated in causing osteolytic phenotypes in multiple myeloma and metastatic breast cancer.29, 47, 48, 49 As Wnt signaling is activated in AML, this may raise concerns that targeting DKK1 overexpression could lead to hyper-activation of the Wnt pathway and enhanced leukemia growth.50, 51, 52 However, our results also indicated that in vivo DKK1 inhibition significantly delays AML progression, rather than accelerates leukemia growth. It was suggested that although AML cells themselves have an endogenous aberrant Wnt activation, they may not be as dependent as normal HSCs on niche-derived Wnt signaling.51 Further studies are required to fully understand the interplay between DKK1 and Wnt signaling in the leukemic niche. Altogether, our results suggest that the elevated DKK1 levels induced by AML-derived exosomes are part of the mechanism employed by AML to alter the lineage potential of BM stromal progenitors and provide a competitive advantage to leukemia cells during AML development. Thus, DKK1 may be a potential new therapeutic target.
Our finding that treating mesenchymal stromal progenitors with AML-derived exosomes increased their adipogenic ability suggests exosome targeted skeletal sites may have increased marrow fat content, which has been associated with enhanced cancer resistance to systemic chemotherapy and radiation.53 Ye et al.54 also indicated that leukemia cells can co-opt adipose tissue to support their metabolism and survival. Recent technology advances have enabled us to simultaneously determine the site-specific marrow fat content, osteoblastic activity and hematopoiesis.55, 56 Improved understanding of bone and marrow composition changes in AML patients may lead to improved therapeutic options.
Altogether, our data support the notion that AML cells remodel the BM niche through exosome secretion, to favor leukemic cell proliferation and survival, whereas suppressing normal hematopoiesis. Although current studies suggest that treatment with certain drugs can reverse the damage caused to the niche by leukemia cells,8 to date, treatments for AML have mainly focused on eradication of leukemia cells, with little consideration that repairing a leukemia-damaged BM niche and eliminating a fertile ground for disease recurrence may be equally important. Thus, novel therapeutic strategies that restore a BM niche that can adequately support normal hematopoiesis over leukemogenesis should be developed. Disrupting exosome production and/or secretion in AML cells or directly targeting factors induced by AML-derived exosomes, such as DKK1, may provide new avenues to prevent AML-induced transformations of BM niche, and enhance the anti-leukemia activity of emerging molecularly-targeted compounds for AML patients.
References
Morrison SJ, Scadden DT . The bone marrow niche for haematopoietic stem cells. Nature 2014; 505: 327–334.
Lane SW, Scadden DT, Gilliland DG . The leukemic stem cell niche: current concepts and therapeutic opportunities. Blood 2009; 114: 1150–1157.
Walkley CR, Olsen GH, Dworkin S, Fabb SA, Swann J, McArthur GA et al. A microenvironment-induced myeloproliferative syndrome caused by retinoic acid receptor gamma deficiency. Cell 2007; 129: 1097–1110.
Walkley CR, Shea JM, Sims NA, Purton LE, Orkin SH . Rb regulates interactions between hematopoietic stem cells and their bone marrow microenvironment. Cell 2007; 129: 1081–1095.
Wang L, Zhang H, Rodriguez S, Cao L, Parish J, Mumaw C et al. Notch-dependent repression of miR-155 in the bone marrow niche regulates hematopoiesis in an NF-kappaB-dependent manner. Cell Stem Cell 2014; 15: 51–65.
Raaijmakers MHGP, Mukherjee S, Guo S, Zhang S, Kobayashi T, Schoonmaker JA et al. Bone progenitor dysfunction induces myelodysplasia and secondary leukaemia. Nature 2010; 464: 852–857.
Kode A, Manavalan JS, Mosialou I, Bhagat G, Rathinam CV, Luo N et al. Leukaemogenesis induced by an activating beta-catenin mutation in osteoblasts. Nature 2014; 506: 240–244.
Zhang B, Ho YW, Huang Q, Maeda T, Lin A, Lee SU et al. Altered microenvironmental regulation of leukemic and normal stem cells in chronic myelogenous leukemia. Cancer Cell 2012; 21: 577–592.
Schepers K, Pietras EM, Reynaud D, Flach J, Binnewies M, Garg T et al. Myeloproliferative neoplasia remodels the endosteal bone marrow niche into a self-reinforcing leukemic niche. Cell Stem Cell 2013; 13: 285–299.
Medyouf H, Mossner M, Jann JC, Nolte F, Raffel S, Herrmann C et al. Myelodysplastic cells in patients reprogram mesenchymal stromal cells to establish a transplantable stem cell niche disease unit. Cell Stem Cell 2014; 14: 824–837.
Skog J, Wurdinger T, van Rijn S, Meijer DH, Gainche L, Sena-Esteves M et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol 2008; 10: 1470–1476.
Kowal J, Tkach M, Thery C . Biogenesis and secretion of exosomes. Curr Opin Cell Biol 2014; 29: 116–125.
Taylor DD, Gercel-Taylor C . Exosomes/microvesicles: mediators of cancer-associated immunosuppressive microenvironments. Semin Immunopathol 2011; 33: 441–454.
Lee TH, D'Asti E, Magnus N, Al-Nedawi K, Meehan B, Rak J . Microvesicles as mediators of intercellular communication in cancer—the emerging science of cellular 'debris'. Semin Immunopathol 2011; 33: 455–467.
Ratajczak J, Miekus K, Kucia M, Zhang J, Reca R, Dvorak P et al. Embryonic stem cell-derived microvesicles reprogram hematopoietic progenitors: evidence for horizontal transfer of mRNA and protein delivery. Leukemia 2006; 20: 847–856.
Melo SA, Sugimoto H, O'Connell JT, Kato N, Villanueva A, Vidal A et al. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell 2014; 26: 707–721.
Peinado H, Aleckovic M, Lavotshkin S, Matei I, Costa-Silva B, Moreno-Bueno G et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat Med 2012; 18: 883–891.
Zhou W, Fong MY, Min Y, Somlo G, Liu L, Palomares MR et al. Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer Cell 2014; 25: 501–515.
Corrado C, Raimondo S, Saieva L, Flugy AM, De Leo G, Alessandro R . Exosome-mediated crosstalk between chronic myelogenous leukemia cells and human bone marrow stromal cells triggers an interleukin 8-dependent survival of leukemia cells. Cancer Lett 2014; 348: 71–76.
Huan J, Hornick NI, Shurtleff MJ, Skinner AM, Goloviznina NA, Roberts CT Jr et al. RNA trafficking by acute myelogenous leukemia exosomes. Cancer Res 2013; 73: 918–929.
Huan J, Hornick NI, Goloviznina NA, Kamimae-Lanning AN, David LL, Wilmarth PA et al. Coordinate regulation of residual bone marrow function by paracrine trafficking of AML exosomes. Leukemia 2015; 29: 2285–2295.
Park CY, Majeti R, Weissman IL . In vivo evaluation of human hematopoiesis through xenotransplantation of purified hematopoietic stem cells from umbilical cord blood. Nat Protoc 2008; 3: 1932–1940.
Krause DS, Fulzele K, Catic A, Sun CC, Dombkowski D, Hurley MP et al. Differential regulation of myeloid leukemias by the bone marrow microenvironment. Nat Med 2013; 19: 1513–1517.
Hanoun M, Zhang D, Mizoguchi T, Pinho S, Pierce H, Kunisaki Y et al. Acute myelogenous leukemia-induced sympathetic neuropathy promotes malignancy in an altered hematopoietic stem cell niche. Cell Stem Cell 2014; 15: 365–375.
Hu X, Garcia M, Weng L, Jung X, Murakami JL, Kumar B et al. Identification of a common mesenchymal stromal progenitor for the adult haematopoietic niche. Nat Commun 2016; 7: 13095.
Zhang X, Yuan X, Shi H, Wu L, Qian H, Xu W . Exosomes in cancer: small particle, big player. J Hematol Oncol 2015; 8: 83.
Sheldon H, Heikamp E, Turley H, Dragovic R, Thomas P, Oon CE et al. New mechanism for Notch signaling to endothelium at a distance by Delta-like 4 incorporation into exosomes. Blood 2010; 116: 2385–2394.
Thery C, Amigorena S, Raposo G, Clayton A . Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr Protoc Cell Biol 2006; Chapter 3: Unit 3.22.
Pinzone JJ, Hall BM, Thudi NK, Vonau M, Qiang YW, Rosol TJ et al. The role of Dickkopf-1 in bone development, homeostasis, and disease. Blood 2009; 113: 517–525.
Ostrowski M, Carmo NB, Krumeich S, Fanget I, Raposo G, Savina A et al. Rab27a and Rab27b control different steps of the exosome secretion pathway. Nat Cell Biol 2010; 12: 19–30.
Camussi G, Deregibus MC, Bruno S, Grange C, Fonsato V, Tetta C . Exosome/microvesicle-mediated epigenetic reprogramming of cells. Am J Cancer Res 2011; 1: 98–110.
Hoshino A, Costa-Silva B, Shen TL, Rodrigues G, Hashimoto A, Tesic Mark M et al. Tumour exosome integrins determine organotropic metastasis. Nature 2015; 527: 329–335.
Wang W, Zimmerman G, Huang X, Yu S, Myers J, Wang Y et al. Aberrant Notch signaling in the bone marrow microenvironment of acute lymphoid leukemia suppresses osteoblast-mediated support of hematopoietic niche function. Cancer Res 2016; 76: 1641–1652.
Frisch BJ, Ashton JM, Xing L, Becker MW, Jordan CT, Calvi LM . Functional inhibition of osteoblastic cells in an in vivo mouse model of myeloid leukemia. Blood 2012; 119: 540–550.
Krevvata M, Silva BC, Manavalan JS, Galan-Diez M, Kode A, Matthews BG et al. Inhibition of leukemia cell engraftment and disease progression in mice by osteoblasts. Blood 2014; 124: 2834–2846.
Bowers M, Zhang B, Ho Y, Agarwal P, Chen CC, Bhatia R . Osteoblast ablation reduces normal long-term hematopoietic stem cell self-renewal but accelerates leukemia development. Blood 2015; 125: 2678–2688.
Massenkeil G, Fiene C, Rosen O, Michael R, Reisinger W, Arnold R . Loss of bone mass and vitamin D deficiency after hematopoietic stem cell transplantation: standard prophylactic measures fail to prevent osteoporosis. Leukemia 2001; 15: 1701–1705.
Ganguly S, Divine CL, Aljitawi OS, Abhyankar S, McGuirk JP, Graves L . Prophylactic use of zoledronic acid to prevent early bone loss is safe and feasible in patients with acute myeloid leukemia undergoing allogeneic stem cell transplantation. Clin Transplant 2012; 26: 447–453.
Drake MT . Osteoporosis and cancer. Curr Osteoporos Rep 2013; 11: 163–170.
Mao B, Wu W, Li Y, Hoppe D, Stannek P, Glinka A et al. LDL-receptor-related protein 6 is a receptor for Dickkopf proteins. Nature 2001; 411: 321–325.
Fleming HE, Janzen V, Lo Celso C, Guo J, Leahy KM, Kronenberg HM et al. Wnt signaling in the niche enforces hematopoietic stem cell quiescence and is necessary to preserve self-renewal in vivo. Cell Stem Cell 2008; 2: 274–283.
Gustafson B, Eliasson B, Smith U . Thiazolidinediones increase the wingless-type MMTV integration site family (WNT) inhibitor Dickkopf-1 in adipocytes: a link with osteogenesis. Diabetologia 2010; 53: 536–540.
Christodoulides C, Laudes M, Cawthorn WP, Schinner S, Soos M, O'Rahilly S et al. The Wnt antagonist Dickkopf-1 and its receptors are coordinately regulated during early human adipogenesis. J Cell Sci 2006; 119 (Pt 12): 2613–2620.
Cheng SL, Shao JS, Cai J, Sierra OL, Towler DA . Msx2 exerts bone anabolism via canonical Wnt signaling. J Biol Chem 2008; 283: 20505–20522.
Granchi D, Baglio SR, Amato I, Giunti A, Baldini N . Paracrine inhibition of osteoblast differentiation induced by neuroblastoma cells. International journal of cancer. J Int Cancer 2008; 123: 1526–1535.
Gregory CA, Singh H, Perry AS, Prockop DJ . The Wnt signaling inhibitor dickkopf-1 is required for reentry into the cell cycle of human adult stem cells from bone marrow. J Biol Chem 2003; 278: 28067–28078.
Tian E, Zhan F, Walker R, Rasmussen E, Ma Y, Barlogie B et al. The role of the Wnt-signaling antagonist DKK1 in the development of osteolytic lesions in multiple myeloma. N Engl J Med 2003; 349: 2483–2494.
Bu G, Lu W, Liu CC, Selander K, Yoneda T, Hall C et al. Breast cancer-derived Dickkopf1 inhibits osteoblast differentiation and osteoprotegerin expression: implication for breast cancer osteolytic bone metastases. International journal of cancer. J Int Cancer 2008; 123: 1034–1042.
Voorzanger-Rousselot N, Goehrig D, Journe F, Doriath V, Body JJ, Clezardin P et al. Increased Dickkopf-1 expression in breast cancer bone metastases. Br J Cancer 2007; 97: 964–970.
Wang Y, Krivtsov AV, Sinha AU, North TE, Goessling W, Feng Z et al. The Wnt/beta-catenin pathway is required for the development of leukemia stem cells in AML. Science 2010; 327: 1650–1653.
Lane SW, Wang YJ, Lo Celso C, Ragu C, Bullinger L, Sykes SM et al. Differential niche and Wnt requirements during acute myeloid leukemia progression. Blood 2011; 118: 2849–2856.
Yeung J, Esposito MT, Gandillet A, Zeisig BB, Griessinger E, Bonnet D et al. beta-Catenin mediates the establishment and drug resistance of MLL leukemic stem cells. Cancer Cell 2010; 18: 606–618.
Conklin KA . Dietary polyunsaturated fatty acids: impact on cancer chemotherapy and radiation. Altern Med Rev 2002; 7: 4–21.
Ye H, Adane B, Khan N, Sullivan T, Minhajuddin M, Gasparetto M et al. Leukemic stem cells evade chemotherapy by metabolic adaptation to an adipose tissue niche. Cell Stem Cell 2016; 19: 23–37.
Frost ML, Fogelman I, Blake GM, Marsden PK, Cook G Jr . Dissociation between global markers of bone formation and direct measurement of spinal bone formation in osteoporosis. J Bone Miner Res 2004; 19: 1797–1804.
Yagi M, Arentsen L, Shanley RM, Rosen CJ, Kidder LS, Sharkey LC et al. A dual-radioisotope hybrid whole-body micro-positron emission tomography/computed tomography system reveals functional heterogeneity and early local and systemic changes following targeted radiation to the murine caudal skeleton. Calcif Tissue Int 2014; 94: 544–552.
Acknowledgements
We thank Dr Sarah Wilkinson for critical reading of the manuscript. Donna Isbell, Stacey Sparks and Danielle Kim of the Animal Resource Center for animal care. Lucy Brown of the Analytical Cytometry Core for flow cytometry and sorting support. Dr Marcia Miller and Zhou Li of the Electron Microscopy Core for assitance in exosomes measurement and imaging. Dr David Colcher and Junie Chea of the Small Animal Imaging Core for μ-CT analysis. Research reported in this publication included work performed in the Small Animal Imaging Core supported by the National Cancer Institute of the National Institutes of Health under award number P30CA33572. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. This study was supported in part by grants to C-CC from the Children Leukemia Research Association, the ThinkCure! Foundation, the Margaret E. Early Medical Research Trust, the Tim Nesvig Lymphoma Research Fund, the STOP Cancer Foundation, the American Cancer Society (Grant 128766-RSG-15-162) and the National Institutes of Health (R00 HL087936). XH was supported by a California Institute for Regenrative Medicine (CIRM) training grant. JLM was supported by a Parsons Foundation Fellowship and NIH fellowship (1F31HL114393-01A1).
Author contributions
BK and MG conducted the experiments; LW, XJ, JLM and XH assisted with data analysis. TM, AL, GM, DLD, ASS, VP and RB obtained and processed AML patient and healthy control samples. ARK provided the MLL-AF9 mouse model. BK, MG, VP, RB and C-CC designed the experiments. BK, MG, SKH, NC, Y-HK, GM and C-CC wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Kumar, B., Garcia, M., Weng, L. et al. Acute myeloid leukemia transforms the bone marrow niche into a leukemia-permissive microenvironment through exosome secretion. Leukemia 32, 575–587 (2018). https://doi.org/10.1038/leu.2017.259
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2017.259
This article is cited by
-
A bispecific nanosystem activates endogenous natural killer cells in the bone marrow for haematologic malignancies therapy
Nature Nanotechnology (2024)
-
CXCR4 as a therapeutic target in acute myeloid leukemia
Leukemia (2024)
-
Harnessing upregulated E-selectin while enhancing SDF-1α sensing redirects infused NK cells to the AML-perturbed bone marrow
Leukemia (2024)
-
ActivinA modulates B-acute lymphoblastic leukaemia cell communication and survival by inducing extracellular vesicles production
Scientific Reports (2024)
-
Crosstalk between autophagy and metabolism: implications for cell survival in acute myeloid leukemia
Cell Death Discovery (2024)