Abstract
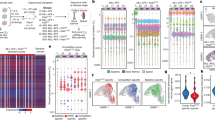
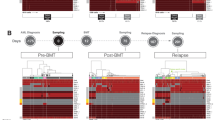
The interplay between tumor heterogeneity and microenvironmental factors is a critical mechanism for clonal selection in leukemia. Evidence of unique clonal capacities to engraft within patient-derived xenograft (PDX) models suggests that intrapatient genetic architecture may be defined by functional differences at the clonal level. However, methods to detect functional differences assigned to genetically defined clones remain limited. Here, we describe a scalable method to directly measure the functional properties of clones within the same leukemia patient by coupling intracellular flow cytometry and next-generation sequencing (NGS). We provide proof of concept utilizing primary chronic myelmonocytic leukemia (CMML) samples and granulocyte–macrophage colony stimulating factor (GM-CSF) to elucidate the interaction between tumor heterogeneity and microenvironmental factors. Mixtures of human leukemia cell lines, with known response to GM-CSF, were used to validate the accuracy of our methodology. Using this approach, we confirm that our method is capable of discriminating GM-CSF sensitive cell lines, identifies somatic variants in primary leukemia samples, and resolves functional clonal architecture in an illustrative patient. Taken together, our data describes a novel method to determine intrapatient functional clonal heterogeneity and provides proof-of-concept for future investigation aimed at elucidating the clinical relevance of functional clonal differences.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Welch JS, Ley TJ, Link DC, Miller CA, Larson DE, Koboldt DC et al. The origin and evolution of mutations in acute myeloid leukemia. Cell 2012; 150: 264–278.
Ding L, Ley TJ, Larson DE, Miller CA, Koboldt DC, Welch JS et al. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 2012; 481: 506–510.
Walter MJ, Shen D, Ding L, Shao J, Koboldt DC, Chen K et al. Clonal architecture of secondary acute myeloid leukemia. N Engl J Med 2012; 366: 1090–1098.
Landau DA, Carter SL, Getz G, Wu CJ . Clonal evolution in hematological malignancies and therapeutic implications. Leukemia 2014; 28: 34–43.
Itzykson R, Solary E . An evolutionary perspective on chronic myelomonocytic leukemia. Leukemia 2013; 27: 1441–1450.
Marusyk A, Tabassum DP, Altrock PM, Almendro V, Michor F, Polyak K . Non-cell-autonomous driving of tumour growth supports sub-clonal heterogeneity. Nature 2014; 514: 54–58.
Kreso A, O'Brien CA, van Galen P, Gan OI, Notta F, Brown AM et al. Variable clonal repopulation dynamics influence chemotherapy response in colorectal cancer. Science 2013; 339: 543–548.
Klco JM, Spencer DH, Miller CA, Griffith M, Lamprecht TL, O'Laughlin M et al. Functional heterogeneity of genetically defined subclones in acute myeloid leukemia. Cancer Cell 2014; 25: 379–392.
Padron E, Painter JS, Kunigal S, Mailloux AW, McGraw K, McDaniel JM et al. GM-CSF-dependent pSTAT5 sensitivity is a feature with therapeutic potential in chronic myelomonocytic leukemia. Blood 2013; 121: 5068–5077.
Ramshaw HS, Bardy PG, Lee MA, Lopez AF . Chronic myelomonocytic leukemia requires granulocyte-macrophage colony-stimulating factor for growth in vitro and in vivo. Exp Hematol 2002; 30: 1124–1131.
Kotecha N, Flores NJ, Irish JM, Simonds EF, Sakai DS, Archambeault S et al. Single-cell profiling identifies aberrant STAT5 activation in myeloid malignancies with specific clinical and biologic correlates. Cancer Cell 2008; 14: 335–343.
Itzykson R, Kosmider O, Renneville A, Morabito M, Preudhomme C, Berthon C et al. Clonal architecture of chronic myelomonocytic leukemias. Blood 2013; 121: 2186–2198.
Itzykson R, Kosmider O, Renneville A, Gelsi-Boyer V, Meggendorfer M, Morabito M et al. Prognostic score including gene mutations in chronic myelomonocytic leukemia. J Clin Oncol 2013; 31: 2428–2436.
Yoshida K, Sanada M, Shiraishi Y, Nowak D, Nagata Y, Yamamoto R et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature 2011; 478: 64–69.
Schmidt E, Pechtl I, McHoull B, Liu M . Streamlining NGS workflows using cancer samples by the application of the DNA Integrity Number (DIN) from the Genomic DNA ScreenTape assay. Cancer Res 2015; 75: 4863.
Schmidt D, Wilson MD, Spyrou C, Brown GD, Hadfield J, Odom DT . ChIP-seq: using high-throughput sequencing to discover protein-DNA interactions. Methods 2009; 48: 240–248.
Robin JD, Ludlow AT, LaRanger R, Wright WE, Shay JW . Comparison of DNA quantification methods for next generation sequencing. Sci Rep 2016; 6: 24067.
Makishima H, Visconte V, Sakaguchi H, Jankowska AM, Kar SA, Jerez A et al. Mutations in the spliceosome machinery, a novel and ubiquitous pathway in leukemogenesis. Blood 2012; 119: 3203–3210.
Merlevede J, Droin N, Qin T, Meldi K, Yoshida K, Morabito M et al. Mutation allele burden remains unchanged in chronic myelomonocytic leukaemia responding to hypomethylating agents. Nat Commun 2016; 7: 10767.
Papaemmanuil E, Cazzola M, Boultwood J, Malcovati L, Vyas P, Bowen D et al. Somatic SF3B1 mutation in myelodysplasia with ring sideroblasts. N Engl J Med 2011; 365: 1384–1395.
Acknowledgements
We thank the Moffitt Cancer Center Core Grant (P30-CA076292), the American Society of Hematology (ASH) Scholar Award, and the MDS Evans Foundation for funding this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Rights and permissions
About this article
Cite this article
Zhang, Q., Ball, M., Zhao, Y. et al. Intrapatient functional clonality deconvoluted by coupling intracellular flow cytometry and next-generation sequencing in human leukemia. Leukemia 32, 532–538 (2018). https://doi.org/10.1038/leu.2017.184
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2017.184