Abstract
Philadelphia chromosome-positive (Ph+) B-cell precursor acute lymphoblastic leukemia (ALL) expressing BCR-ABL1 oncoprotein is a major subclass of ALL with poor prognosis. BCR-ABL1-expressing leukemic cells are highly dependent on double-strand break (DSB) repair signals for their survival. Here we report that a first-in-class HDAC1,2 selective inhibitor and doxorubicin (a hyper-CVAD chemotherapy regimen component) impair DSB repair networks in Ph+ B-cell precursor ALL cells using common as well as distinct mechanisms. The HDAC1,2 inhibitor but not doxorubicin alters nucleosomal occupancy to impact chromatin structure, as revealed by MNase-Seq. Quantitative mass spectrometry of the chromatin proteome along with functional assays showed that the HDAC1,2 inhibitor and doxorubicin either alone or in combination impair the central hub of DNA repair, the Mre11–Rad51–DNA ligase 1 axis, involved in BCR-ABL1-specific DSB repair signaling in Ph+ B-cell precursor ALL cells. HDAC1,2 inhibitor and doxorubicin interfere with DISC (DNA damage-induced transcriptional silencing in cis)) or transcriptional silencing program in cis around DSB sites via chromatin remodeler-dependent and -independent mechanisms, respectively, to further impair DSB repair. HDAC1,2 inhibitor either alone or when combined with doxorubicin decreases leukemia burden in vivo in refractory Ph+ B-cell precursor ALL patient-derived xenograft mouse models. Overall, our novel mechanistic and preclinical studies together demonstrate that HDAC1,2 selective inhibition can overcome DSB repair ‘addiction’ and provide an effective therapeutic option for Ph+ B-cell precursor ALL.
Similar content being viewed by others
Introduction
The Philadelphia (Ph) chromosome resulting from reciprocal t(9;22) translocation was the first reported chromosomal rearrangement linked to a human malignancy.1 The Ph chromosome results in BCR-ABL1 fusion gene, giving rise to the BCR-ABL1 oncoprotein, which drives B-cell precursor acute lymphoblastic leukemia (ALL) and chronic myelogenous leukemia.1, 2 Imatinib (a tyrosine kinase inhibitor of BCR-ABL1 activity) along with hyper-CVAD (cyclophosphamide, vincristine, adriamycin/doxorubicin and dexamethasone) is the standard treatment for Ph+ B-cell precursor ALL.3 However, long-term remission is rare in patients with B-cell precursor ALL compared with chronic myelogenous leukemia, as point mutations in BCR-ABL1 such as the T315I mutation impair drug binding and confer resistance to imatinib and second-generation tyrosine kinase inhibitors.4 Stem cell transplantation along with imatinib is a treatment option with promising potential, but relapse rates and treatment-related deaths are high.5, 6 Additionally, late toxicities and functional impairment are common in long-term survivors and the disease remains incurable in most adults. Therefore, there is a real need for new therapeutics for Ph+ B-cell precursor ALL.
Unlike mismatches and DNA adducts, double-strand breaks (DSBs) are lethal to a cell if left unrepaired.7 BCR-ABL1 was reported to increase DSB repair using non-homologous end joining (NHEJ) and homologous recombination (HR).8, 9, 10, 11 The increase in BCR-ABL1-stimulated DSB repair was attributed to increased expression and/or activity of multiple DSB repair proteins, which confer major survival advantages, including resistance to genotoxic therapies and preventing apoptosis in Ph+ leukemic cells.8, 9, 10, 11 Therefore, an attractive therapeutic approach would be to target the multiple BCR-ABL1-driven aberrantly hyperactive DSB repair signals in Ph+ leukemic cells. However, an inhibitor that directly curtails multiple DNA repair processes to impair BCR-ABL1-mediated DSB repair networks is not available for Ph+ B-cell precursor ALL. Although one could use a cocktail of inhibitors against various DNA repair proteins, an alternative strategy is to use an inhibitor either in isolation or in combination with existing chemotherapy drug(s) to effectively target the various BCR-ABL1-driven aberrant DNA repair signals.
Pan histone deacetylase (HDAC) inhibitors are Food and Drug Administration approved for treating cutaneous T-cell lymphoma, refractory peripheral T-cell lymphoma and multiple myeloma.5, 12, 13, 14 A pan or selective HDAC inhibitor to treat B-cell malignancies is currently not available. Pan HDAC inhibitors exhibit adverse side effects, including cardiac toxicity, due to their targeting of multiple class I and II HDACs with important cellular functions.15, 16 We previously reported an unrecognized genome maintenance function for a subset of class I HDACs, the main targets of pan HDAC inhibitors currently in clinic.17, 18, 19, 20, 21, 22 We showed that HDAC1 and HDAC2 (HDAC1,2)—two class I HDACs—localize to sites of DNA damage in B-cell-derived cancers, and small-molecule inhibition of HDAC1,2 activity induces DSB accumulation,22 implicating a direct role for these enzymes in regulating DSB repair. However, a comprehensive understanding of the DSB repair pathways regulated by HDAC1,2 and the precise mode of HDAC1,2 inhibitor action remained to be elucidated.
Here we report the molecular mechanisms by which HDAC1,2 inhibitor impinges on DSB repair at multiple levels to overcome BCR-ABL1-mediated repair and provide the first evidence for the use of a selective HDAC1,2 inhibitor in treating DNA repair ‘addicted’ cancers. We present a novel mechanism-based strategy wherein combining HDAC1,2 selective inhibitor with a standard-of-care chemotherapy agent doxorubicin targets parallel DNA repair pathways to provide therapeutic benefits for Ph+ B-cell precursor ALL.
Methods
Laser break assay
SupB15 or primary patient Ph+ B-cell precursor ALL cells were treated with either 2 μM HDAC1,2 inhibitors for 46 h or 25 nM doxorubicin for 10 h or a combination of HDAC1,2 inhibitor and doxorubicin for 46 h where doxorubicin was added to cells during the last 10 h. We were unable to use 0.1 μM doxorubicin for these assays, as this concentration of doxorubicin when combined with micro-irradiation caused cells to detach from the chamber dish. The chamber dish was treated with Cell-Tak solution (Corning Inc., Bedford, MA, USA) for 30 min to allow suspension B-cell precursor ALL cells to adhere. Following incubation, the solution was removed and the treated cells were added to the wells prior to laser micro-irradiation. Z-stack images were acquired with confocal microscopy and at least 100 cells were quantified in each experiment. All other methods are described in Supplementary Methods.
Results
First-in-class HDAC1,2 selective inhibitors cause cytotoxicity and reduce DSB repair in Ph+ B-cell precursor ALL cells
Neutral comet assays showed a faster disappearance of DNA damage—indicated by shorter comet tails—in BCR-ABL1+ mouse Ba/F3 cells compared with control cells at various time points postrecovery from exposure to 4 Gy dose of ionizing radiation (Supplementary Figures S1a and b), confirming that BCR-ABL1 promotes hyperactive DNA repair. As HDAC1,2 localize to DSB sites in B cells,22 we set out to test our hypothesis that selective HDAC1,2 inhibition could override BCR-ABL1-driven hyperactive DSB repair and provide a therapeutic strategy in Ph+ B-cell precursor ALL.
We confirmed the localization of HDAC1,2 to DSBs in Ph+ B-cell precursor ALL cells (Supplementary Figure S1c). We confirmed the selectivity of small molecules—ACY957, ACY1035, ACY1071—to inhibit HDAC1,2. In vitro HDAC assays showed they were ~2log-fold more potent toward HDAC1 and HDAC2 compared with HDAC3 (Supplementary Figures S2a and b).22, 23 We observed a robust increase in histone acetylation (H4K5ac) in Ph+ B-cell precursor ALL SupB15 cells treated with 1–2 μM of HDAC1,2 inhibitors (Supplementary Figure S2c). HDAC1,2 selective inhibitor treatment of SupB15 cells caused a significant increase in the dead cell sub-G1 population (Supplementary Figure S3a), γH2AX foci formation (a marker of DNA damage) (Supplementary Figure S3b) and comet tail moment (indicating DNA breaks) (Supplementary Figure S3c). HDAC1,2 selective inhibition did not affect the viability of cell lines derived from solid tumors (U87 and MES-SA) but decreased the viability of NALM6 (a Ph− B-cell precursor ALL line) (Supplementary Figures S4a–c), suggesting that B-cell ALL without BCR-ABL1 are also sensitive to HDAC1,2 inhibition. Nevertheless, we then tested whether HDAC1,2 inhibition impacts DNA repair in BCR-ABL1+ leukemic cells using reporter cell lines to measure the efficiency of NHEJ or HR.24 Owing to the very low frequency of HR events, we were unable to reliably measure its efficiency using the reporter cell line. However, NHEJ repair efficiency was significantly decreased with ACY1035 treatment in the EJ5-GFP reporter cell line (Supplementary Figure S3d). Together, these results showed that BCR-ABL1-mediated hyperactive DSB repair and cell viability of Ph+ B-cell precursor ALL are critically sensitive to HDAC1,2 inhibition.
We then sought a drug that when combined with HDAC1,2 inhibitor could show a synergistic impact on genome maintenance. Doxorubicin, an anthracycline and a component of the multi-agent chemotherapy regimen, is used in B-cell precursor ALL treatment. Doxorubicin intercalates DNA, impairs DNA repair and affects chromatin dynamics at high concentrations.25 However, the precise mechanism of doxorubicin action in these processes is not understood. Given the similarity between the adverse effects of HDAC1,2 inhibitors and doxorubicin on genome stability, we tested the beneficial effects of doxorubicin treatment as single or combination agent with ACY1035 in SupB15 cells. Low-concentration doxorubicin (0.1 μM) treatment for 10 h caused accumulation of DNA damage without causing cell death (Supplementary Figures S5a–c). Hence, we treated Ph+ B-cell precursor ALL cells with 0.1 μM doxorubicin for 10 h for mechanistic studies described below. Prolonged treatment with 0.1 μM doxorubicin for 48 h or combining it with ACY1035 caused accumulation of sub-G1 or dead cells (Supplementary Figure S5d). Staining for SupB15 cell viability showed a significant increase in dead cells following treatment with ACY1035 for 72 h or 0.1 μM doxorubicin for 48 h (Supplementary Figure S5e). Collectively, our results suggested that HDAC1,2 inhibitor in isolation or in combination with doxorubicin, a drug that also impairs DSB repair, triggers apoptosis in Ph+ B-cell precursor ALL cells likely via negating complementary BCR-ABL1-driven DNA repair and genome maintenance signals.
HDAC1,2 inhibitor affects transcription of a small set of DNA repair genes in Ph+ B-cell precursor ALL cells
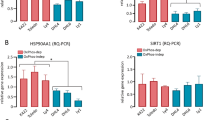
High expression and/or activity of DNA repair factors, including Rad51, Nbs1, Mre11 and DNA ligase 3, was reported in BCR-ABL1+ leukemic cells.8, 9, 10 To examine whether decreased DSB repair upon ACY1035 treatment results from altered expression of DNA repair genes involved in BCR-ABL1-mediated repair signaling, we performed RNA sequencing on SupB15 cells treated with DMSO, ACY1035, doxorubicin or the combination of ACY1035 and doxorubicin. At a threshold of ⩾1.3-fold change, ACY1035 altered the expression of a large number of genes (>2000) compared with doxorubicin (Supplementary Figure S6a). The number of genes altered by combined ACY1035 and doxorubicin treatment was similar to that obtained with ACY1035 alone, suggesting that the doxorubicin concentration used is sufficient to impair DNA repair without altering transcriptional programs. Ingenuity pathway analysis of HDAC1,2 target genes revealed multiple biological processes, including a small set of genes belonging to the ‘DNA damage response’ ontology (Supplementary Figure S6b). This set contained Nbs1 (Nbn), Fanconi anemia genes, SWI/SNF BAF chromatin remodeler complex components (Smarcb1/BAF47, Smarcd3/BRG1) and PB1 (a PBAF remodeling complex component) (Supplementary Figures S6b and c), which are all linked to DNA repair or replication stress-coupled repair.26, 27, 28, 29 Although not identified by pathway analysis, transcript levels for DNA ligase 1 and DNA ligase 3, which are overexpressed in BCR-ABL1+ leukemic cells,8 were also reduced upon HDAC1,2 inhibition (Supplementary Figure S6d). Overall, our RNA-Seq analysis showed that HDAC1,2 inhibition but not doxorubicin decreases the expression of a small set of DNA repair genes in Ph+ B-cell precursor ALL cells.
HDAC1,2 inhibition increases histone acetylation and decreases nucleosomal occupancy to change chromatin landscape
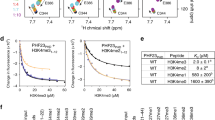
To gain insight into the adverse effects on HDAC1,2 inhibition on DNA repair, we examined bulk histone acetylation in SupB15 cells. HDAC1,2 inhibition but not doxorubicin increased global histone acetylation (Supplementary Figures S7a–c). Using a quantitative chromatin immunoprecipitation (ChIP)-seq or ChIP-Rx approach,30 we then examined the impact of HDAC1,2 inhibition on the genome-wide occupancy of H3K27 acetylation (H3K27ac), a histone mark associated with transcription.31 HDAC1,2 inhibition resulted in increased H3K27ac levels around transcription start sites (TSS), transcription termination sites and enhancer regions in SupB15 cells (Figure 1a). Histone acetylation regulates nucleosome and chromatin structure. In addition to gene transcription, efficient DNA repair is also dependent on proper nucleosome or chromatin structure.32 Therefore, we performed micrococcal nuclease (MNase)-Seq experiments involving high-throughput sequencing of mononucleosomal DNA obtained by MNase digestion of chromatin. Although ACY1035 treatment caused no change in MNase-Seq reads at silent genes and around transcription termination sites, a statistically significant decrease in MNase-Seq reads was observed around TSSs of expressed genes (Figure 1b). Doxorubicin treatment did not alter nucleosomal occupancies at the concentration tested and combined ACY1035 and doxorubicin treatment showed no additive effect (Figure 1b). Moreover, ACY1035 treatment for longer duration (62 h) caused dramatic decrease in nucleosomal occupancies around TSSs of expressed genes (Figure 1c). Overall, these findings indicated that a low concentration of doxorubicin does not alter global chromatin structure, but ACY1035 treatment changes the chromatin landscape during a treatment period when significant death was not observed in SupB15 cells treated with these compounds.
HDAC1,2 inhibition increases H3K27ac genome wide and alters nucleosome occupancy in SupB15 cells. (a) SupB15 cells were treated with DMSO or 2 μM ACY1035 for ChIP-Rx using anti-H3K27ac. DMSO-treated cells were also used for ChIP-Rx with control rabbit IgG alone. Transcription start site (TSS) and transcription termination site (TTS) were ordered from the highest to lowest average TPM (transcripts per million reads) for genes upregulated by ACY1035 treatment compared with control DMSO. CD20 enhancers described previously were used for the analysis.46 Heat maps were generated using Deeptools. Coverage profiles for ChIP-Rx reads are shown on top of the heat maps. Three independent biological replicates were used. (b) Average mean profiles for MNase-Seq reads, indicating nucleosomal occupancy, across 800 bp region upstream (−) or downstream (+) of TSS or TTS of high or low expressed or transcriptionally silent genes are shown. *P-value=0.05 and **P-value=0.06. Data are from two independent control samples and three independent samples treated with 2 μM ACY1035 for 46 h or 0.1 μM doxorubicin for 10 h or 2 μM ACY1035 initially for 36 h followed by an additional 10 h with 0.1 μM doxorubicin, respectively. (c) MNase-Seq analysis with Drosophila S2 cells as an internal spike-in reference genome was performed with SupB15 cells treated with ACY1035 for 62 h. Heat maps and plots were generated as described for (a).
Quantitative proteomic analysis identifies key DSB repair networks targeted by HDAC1,2 inhibitor and doxorubicin
To gain insight into the defective DNA repair, we then examined histone acetylation on chromatin associated with damaged DNA. Immunoprecipitation experiments showed increased histone acetylation in γH2AX-modifed mononucleosomes following ACY1035 treatment (Supplementary Figure S7d) but not with doxorubicin (data not shown). We then asked whether ACY1035 and/or doxorubicin treatments impact the chromatin association of proteins involved in genome maintenance. We isolated chromatin fractions from BCR-ABL1+ Ba/F3 cells treated with DMSO (control), ACY1035, doxorubicin or ACY1035 and doxorubicin. We used a label-free mass spectrometric approach to make quantitative comparisons between these four treatments. Two-dimensional agglomerative clustering of proteins showed highly reproducible peptide counts between the three replicates for each treatment (Figure 2a). We then filtered the data using an analysis of variance P-value⩽0.01 and fold-change in peptide counts was calculated for each treatment vs control comparison (Figure 2b).
HDAC1,2 inhibition and/or doxorubicin treatment changes the chromatin proteome. (a) Mass spectrometry of the chromatin proteome in the indicated control or treated cells. N=3. (b) Venn diagrams were created from the list of proteins with ⩾1.5-fold increase or ⩾1.3-fold decrease in chromatin-bound levels following ACY1035 and/or doxorubicin treatments compared with DMSO control. (c) Proteins showing ⩾1.3-fold decrease in their chromatin association following HDAC1,2 inhibition were subjected to STRING analysis.
ACY1035 treatment caused ⩾1.5-fold increase in the chromatin-bound levels of 156 proteins (Figure 2b and Supplementary Table S1), including cell cycle regulatory and/or transcription factors, such as p53, Runx1, Runx3, Brd2 and programmed cell death-related proteins (granzyme C, granzyme B). HDAC1,2 inhibition caused ⩾1.5-fold decrease in the chromatin-bound levels of 66 proteins, which included 17 chromatin modifiers or remodelers, such as PB1, p400, EPC2, CBX4, CBX8 and TRRAP (Supplementary Table S2 and Supplementary Figure S8a). Four of these chromatin remodelers (PB1, EPC2, CBX4, CBX8) are directly or indirectly associated with polycomb group proteins (PRC1 or PRC2) involved in transcriptional repression.27, 33 Thus these results uncovered a novel link between HDAC1,2 activity and the binding of nucleosome remodelers with chromatin. Setting the threshold to ⩾1.3-fold decrease yielded a larger set of 158 proteins linked to diverse nuclear processes, including cell cycle, DNA damage and DNA repair (Figures 2b and c). We identified 17 DNA repair proteins, including those belonging to the NHEJ and HR repair pathways, namely, Xrcc5/Ku80, and components of the crucial MRN DNA repair protein complex (Mre11, Rad50 and Nbs1/NBN)34 (Figure 2c and Supplementary Figure S8b). Overall, proteome-wide analyses revealed that HDAC1,2 inhibition reduces chromatin association of DNA repair factors and chromatin remodelers regulating various aspects of DSB repair.
Doxorubicin treatment caused ⩾1.5-fold increase in the chromatin-bound levels of 213 proteins, including p53 (Figure 2b), which correlates well with the activation of DNA damage response (Supplementary Figure S5a and b). Doxorubicin treatment decreased the chromatin-bound levels of 287 proteins, including 20 DSB repair proteins, such as the MRN complex components (Mre11, Nbs1, Rad50) that are also reduced by HDAC1,2 inhibition (Figure 2b). In contrast to ACY1035, doxorubicin treatment reduced the chromatin association of 20 core DNA replication proteins, including DNA helicases, Mcm2–7, DNA polymerases and enzymes involved in Okazaki fragment maturation (RNaseH2a and Fen1) (Supplementary Figure S9). Decreased chromatin association for 76 proteins was observed only in the combined ACY1035 and doxorubicin treatment and vast majority of these factors are involved in RNA metabolism (RNA processing and splicing) (Figure 2b and Supplementary Figure S10). Overall, comprehensive analysis of global chromatin proteome showed for the first time that HDAC1,2 inhibitor and/or doxorubicin treatments impinge on common as well as distinct genome maintenance networks.
HDAC1,2 inhibition and doxorubicin override Mre11-dependent DSB repair in Ph+ B-cell precursor ALL cells and primary patient samples
BCR-ABL1+ cells have increased levels and/or activity of DSB repair factors, including Mre11, Nbs1, Rad51, Lig3 and Ligase1 (Lig1).9, 10 Immunoblotting to confirm mass spectrometric results showed no change in Nbs1, Ku70, Ku80 and Sin3A levels, but chromatin-bound levels of DSB repair proteins Mre11, Rad50, Rad51 and DNA Lig1 were substantially decreased following treatment with ACY1035 alone or with ACY1035 and doxorubicin of BCR-ABL1+ mouse Baf/3 cells or SupB15 cells (Supplementary Figures S11a–c).
We then asked whether HDAC1,2 inhibitor and/doxorubicin impact the recruitment of repair factors to DSB sites. We induced DSBs in SupB15 cells using microlaser and monitored recruitment of factors over time using immunofluorescence. We observed a reduction and a 15 min delay in recruitment of Mre11 to break stripes following treatment with ACY1035 or doxorubicin alone or with combined ACY1035 and doxorubicin (Figure 3a). Decreased recruitment of DNA ligase 1 to DSBs was also observed with these treatments (Figure 3b). Although Rad50 and DNA ligase 3 recruitment remained unaffected following ACY1035 and/or doxorubicin treatments (Supplementary Figures S12a and c), a modest but statistically significant decrease in Nbs1 recruitment was observed in combined ACY1035 and doxorubicin treatment (Supplementary Figure S12b). Interestingly, immunoblots showed that doxorubicin treatment increased the chromatin association of Rad51 in wild-type and BCR-ABL1+ mouse BaF/3 cells but not in human SupB15 cells (Supplementary Figure S11), highlighting the species-specific difference in the dynamics of Rad51 recruitment to damaged chromatin. We then used a standard immunofluorescence approach to examine Rad51 localization to DSB sites in SupB15 cells, as we were unable to detect it using the microlaser irradiation approach. To avoid continuous accumulation of DSBs due to ACY1035 and/or doxorubicin treatments, we removed these compounds from the growth medium and monitored Rad51 recruitment to DSBs over time. ACY1035 and doxorubicin combination treatment caused a significant decrease in the number of Rad51 foci co-localizing with DSBs (γH2AX foci) compared with doxorubicin treatment alone (Figure 3c, see 0–2 h recovery time points). Moreover, the percentage of cells with Rad51 co-localizing to DSBs in ACY1035 and the combination treatment reached the levels found in doxorubicin treatment alone only after 4 h postremoval of these compounds (Figure 3c), indicating a delay in Rad51 recruitment to doxorubicin-induced DSB sites in the presence of ACY1035.
HDAC1,2 inhibition and/or doxorubicin treatment impairs the Mre11–DNA ligase1–Rad51 DNA repair axis at DSBs. SupB15 cells were treated with DMSO or ACY1035 (2 μM) for 46 h or Doxorubicin (25 nM) for 10 h. For combined treatment, cells in 2 μM ACY1035 for 36 h were treated for an additional 10 h in the presence of 25 nM Doxorubicin. DSB were generated by 405 nm microlaser-fitted microscope. Immunofluorescence (IF) to examine recruitment of repair factors Mre11 (a) or DNA ligase1 (b) to DSB stripe (marked by γH2AX) was performed 15 or 30 min after microirradiation. 25 nM doxorubicin (Doxo) concentration was used as a higher concentration (0.1 μM) and micro-irradiation compromised cell viability. Percentage of cells with Mre11 or DNA Lig1 co-localizing with γH2AX was quantified from multiple biological replicates. For Mre11, ACY1035 vs DMSO P-value: 0.06; Doxo vs DMSO P-value: 0.02; and combined ACY1035 and Doxo vs DMSO P-value: 0.002; N=4. For DNA Lig1, ACY1035 vs DMSO P-value: 0.06; Doxo vs DMSO P-value: 0.29; and combined ACY1035 and Doxo vs DMSO P-value: 0.04. N=5. (c) SupB15 cells were treated as described in (a and b) prior to growth in a drug-free medium. IF was performed to measure Rad51 localization to γH2AX-marked break sites at the indicated time points after drug removal. 0 h recovery, P-value: 0.09; 1 h recovery, P-value: 0.002; and 2 h recovery, P-value: 0.08. N=5.
Three out of the four primary patient Ph+ B-cell precursor ALL CD34+ progenitor cells showed sensitivity to either HDAC1,2 inhibition alone or the combination treatment, as evidenced by the increased sub-G1 population or dead cells (Figure 4a). These treatments also increased DSBs in primary patient samples (Figure 4b and Supplementary Figure S13). A decrease in Mre11 levels at DSBs was observed following ACY1035 and/or doxorubicin treatments (Figure 4c). Therefore, HDAC1,2 inhibitor and doxorubicin adversely impact the key Mre11-mediated DNA repair network to overcome the BCR-ABL1-mediated DSB repair and survival advantages in Ph+ B-cell precursor ALL cell lines and primary patient cells.
HDAC1,2 inhibitor causes cytotoxicity and impairs DSB repair in primary patient Ph+ B-cell precursor ALL cells. (a) Cell cycle analysis of cultured CD34+ cells isolated from three different Ph+ B-cell precursor ALL patients was performed 72 h following the indicated treatments. (b) Neutral comet assays were performed on CD34+ cells isolated from three Ph+ B-cell precursor ALL patients treated with 2 μM HDAC1,2 selective inhibitor for 46 h or 0.1 μM doxororubicin for 10 h or combination treatment with 2 μM HDAC1,2 selective inhibitor for 36 h and with 0.1 μM doxorubicin for an additional 10 h. Tail moments were calculated from at least 100 nuclei and binned into 7 groups of increasing tail moments. (c) Mre11 localization to microlaser-induced break stripes (γH2AX) in primary Ph+ B-cell precursor ALL CD34+ patient cells treated with either 2 μM ACY1035 for 46 h or 25 nM doxorubicin for 10 h and their combination for a total of 46 h. Percentage of patient cells with Mre11 co-localizing with γH2AX in various treatments were calculated and plotted.
HDAC1,2 inhibitor and doxorubicin inhibit DNA damage-induced transcriptional silencing in cis (DISC) during DSB repair
Agreeing well with HDAC1,2 functions in regulating chromatin structure (Figure 1), only HDAC1,2 inhibition but not doxorubicin decreased the chromatin association of subunits of the PBAF chromatin remodeler complex (Baf180), TRRAP/Tip60/NuA4 chromatin modifier/remodeler complex (p400, Dmap1, Trrap, Brd8 and Epc2) and PcG-PRC1 complex (Cbx4 and Cbx8) (Figures 2b and c and Supplementary Figure S14a). Consistent with our previous findings,20 SMARCA5 chromatin remodeler levels remained unchanged upon HDAC1,2 inhibition. In addition to propagation of DNA repair signaling, a phenomenon termed DISC or shutting down of transcriptional events at chromatin surrounding DSBs is a key process during DNA repair, failure of which results in halting of DSB repair.35, 36 We focused on PB1 (Baf180), a PBAF chromatin-remodeling complex subunit, as it promotes H2AK119 ubiquitination around DSBs to facilitate DISC and allow efficient DNA repair.27 Chromatin-bound Baf180 levels were substantially decreased following treatment with ACY1035 or ACY1035 and doxorubicin but not with doxorubicin alone (Supplementary Figures S14a and b). Additionally, steady-state Baf180 transcript and protein levels were also decreased with ACY1035 treatment (Supplementary Figures S6b and S14c). Baf180 is primarily a nuclear and chromatin-bound protein (Supplementary Figure S14d). Therefore, decreased Baf180 steady-state levels following ACY1035 treatment could be attributed primarily to its reduced chromatin binding. A significant increase in cells with 1–5 γH2AX foci was observed following knockdown of Baf180 (Supplementary Figure S14e), revealing a functional role for Baf180 in DNA repair in Ph+ leukemic cells.
To directly examine whether HDAC1,2 and Baf180 are required for DISC, we used the Fok1 nuclease-based DSB dual reporter system, which enables one to simultaneously visualize nascent transcription (yellow fluorescent protein (YFP) reporter) and DNA damage (mCherry reporter).36 ACY1035 treatment led to a twofold increase in YFP signal compared with control cells (Figure 5a), suggesting a loss in transcriptional repression around DSBs in the absence of HDAC1,2 activity. Knockdown of Baf180 also increased nascent YFP expression, indicating a disruption of DISC (Supplementary Figure S15). Surprisingly, increased YFP signal was also observed following doxorubicin treatment (Figure 5b). H3K27me3 catalyzed by the EZH2-containing PRC2 complex is involved in transcriptional repression around break sites during DSB repair.27 H3K27me3 was decreased at microlaser-induced break sites in SupB15 cells treated with ACY1035 and not doxorubicin (Supplementary Figure S16). Increased transcription around break sites was also observed following ACY1035 and doxorubicin combination treatment (Figure 5c). Collectively, these results showed that HDAC1,2, and possibly doxorubicin-targeted topoisomerase activities, are required to maintain transcriptional repression around DSB sites during DNA repair, in addition to controlling repair signaling promoted by the BCR-ABL1 oncoprotein in Ph+ leukemic cells.
HDAC1,2 inhibitor or doxorubicin impair DISC during DSB repair. DISC was measured using an inducible Fok1 endonuclease-based DSB reporter cell line. Fok1-mcherry indicates the site of DSB and signal for YFP reporter indicates transcriptional activation. Representative nuclei with Fok1-mCherry or YFP or with Fok1-mCherry co-localizing with YFP are shown for 2 μM ACY1035 treatment for 48 or 72 h (a) or 0.1 μM doxorubicin treatment for 10 h N=5 (b) or 2 μM ACY1035 in combination with 0.1 μM doxorubicin for a total of 46 h where doxorubicin was added during the last 10 h of incubation, N=4 (c). *P-value=0.003 and **P-value=0.004. Quantitation in (a) is shown for two independent experiments at 48 and 72 h treatment time points. Txn, Transcription; DSB, double strand break.
HDAC1,2 inhibition in isolation or in combination with doxorubicin decreases leukemia burden in vivo in PDX mouse models
We examined the in vivo efficacy of HDAC1,2 inhibitor treatment using Ph+ B-cell precursor ALL patient-derived xenograft (PDX) mouse models. We used ACY1035 and ACY1071, as they exhibited similar inhibitory effect in vitro and displayed comparable pharmacokinetic characteristics in mice in vivo (Supplementary Figure S17). We created a xenograft mouse model using CD34+ cells obtained from a Ph+ B-cell precursor ALL patient with BCR-ABL1 (T315I) mutation, who had relapsed after imatinib and dasatinib treatments along with autologous stem cell transplantation and ultimately died. This PDX mouse model (PDX#1) was made using mostly bone marrow-derived CD34+ cells obtained from a primary mouse recipient. In this PDX model, leukemia homed primarily to the bone marrow and not to the spleen or peripheral blood during the course of our study—as evidenced from spleen size (Supplementary Figure S18a), immunohistochemical staining with anti-CD19 antibody (Supplementary Figure S18b) and fluorescence-activated cell sorter (FACS) analysis (data not shown). A dramatic response was observed with HDAC1,2 inhibitor treatment alone and in combination with doxorubicin in this PDX model, as seen from the significant reduction in human CD19+ cells in the bone marrow using FACS analysis (Figure 6a). Immunohistochemistry of bone marrow sections using hematoxylin and eosin (H&E) staining or CD19/TdT staining also showed a dramatic reduction in leukemia burden following HDAC1,2 inhibitor treatment (Figure 6a). Bone marrow was repopulated with red blood cells in HDAC1,2 inhibitor- and/or doxorubicin-treated mice, revealing leukemia regression, which is in striking contrast to a pale bone color indicating white blood cell infiltration or leukemia burden in vehicle-treated mice (Figure 6a). Confirming the in vivo efficacy, HDAC1,2 inhibition increased H4K5ac in bone marrow cells (Supplementary Figure S18c). No significant decrease in body weight was observed in PDX#1 mice treated with ACY1035 (Supplementary Figure S18d). Genetic loss of HDAC1,2 was reported to cause cardiac toxicity.37 Fibrosis, an early sign of cardiomyopathy,38 was not present by H&E staining in the hearts of treated mice, indicating that transient selective inhibition of HDAC1,2 does not lead to cardiac toxicity (Supplementary Figure S18e).
HDAC1,2 inhibitor and doxorubicin reduce leukemia burden in primary Ph+ B-cell precursor ALL PDX models. (a) FACS analysis of human CD19 in bone marrow (BM) cells of PDX#1 mouse model treated with vehicle, ACY1035, doxorubicin or ACY1035 plus doxorubicin. *P-value=0.01 and **P-value=0.002. N=9 for vehicle, N=7 for ACY1035, N=4 for doxorubicin and N=5 for ACY1035 plus doxorubicin. Immunohistochemistry of BM sections showing a decrease in TdT and CD19-positive cells following treatment of PDX#1 mice with ACY1035. Images of bones from vehicle or HDAC1,2 inhibitor (ACY1035 or ACY1071) treated PDX#1 mice are shown. Pale color indicates leukemia burden and red color indicates re-establishment of normal hematopoiesis. (b) FACS analysis of human CD19 in bone marrow (BM) or spleen (SPL) cells of PDX#2 model treated with vehicle or doxorubicin (Doxo). N=8 for vehicle and N=6 for Doxo groups. FACS analysis of human CD19 in BM or spleen SPL cells of PDX#2 model treated with vehicle or ACY1071 or ACY1071 plus doxorubicin. (*P-value: 0.08, **P-value: 0.02) N=3 per group. Assessment of leukemia in PDX#2 mice by H&E staining following the indicated treatments. Images of bones from vehicle- or drug-treated PDX#2 mice are shown. (c) FACS analysis of human CD19 in bone marrow (BM) or spleen (SPL) cells of PDX#2 model fed with control chow or ACY957 chow plus doxorubicin. Assessment of leukemia in vehicle or drug combination treated PDX#2 mice by H&E staining. Images of bones from vehicle- or drug-treated PDX#2 mice are also shown. *P-value=0.07 and **P-value=0.02.
We made another model (PDX#2) where immune-compromised NSG mice were injected with cells obtained from a very late stage Ph+ B-ALL patient whose bone marrow had 95% leukemia infiltration at the time of diagnosis, had acquired BCR-ABL1 (T315I) mutation during dasatinib treatment and relapsed later. Leukemic cells in this model were highly resistant to doxorubicin treatment (Figure 6b). Even in this very late stage disease model, a significant response was observed with combined HDAC1,2 inhibitor and doxorubicin treatment compared with HDAC1,2 inhibitor treatment alone, as evidenced by the decrease in leukemia burden using FACS, H&E staining and visual observation of red blood cell infiltration in bone marrow (Figure 6b). A significant increase in H4K5ac in the bone marrow of mice treated with ACY1071 and doxorubicin was observed (Supplementary Figure S18f), along with an absence of body weight loss (Supplementary Figure S18g) and lack of cardiac toxicity (Supplementary Figure S18h). A significant decrease in the leukemia burden was also observed with orally formulated ACY957 chow when used in combination with doxorubicin (Figure 6c) and displayed no cardiac toxicity (Supplementary Figure S18i). Collectively, these results from preclinical studies demonstrate that HDAC1,2 inhibition either alone or in combination with doxorubicin can suppress the growth of BCR-ABL1-expressing leukemic cells in vivo.
Discussion
As summarized in Figure 7, our studies demonstrate how HDAC1,2 inhibitor and doxorubicin adversely impinge on common and distinct genome maintenance networks to overcome BCR-ABL1-mediated survival advantages in Ph+ B-cell precursor ALL cells. Our findings suggest a model wherein inhibition of HDAC1,2 causes an MNase-accessible or ‘permissive’ chromatin structure that provides an unsuitable binding platform resulting in decreased association of chromatin remodelers and other key repair factors and thus adversely impacts many critical events during DSB repair (Figures 1, 2, 3, 4, 5, 6). It is conceivable that a more profound change in chromatin structure upon prolonged absence of HDAC1,2 activity could lead to the disruption of heterochromatic regions and account for the cell death following HDAC1,2 inhibitor treatment. Although Ph−B-ALL cells are also sensitive to HDAC1,2 inhibition when compared with solid tumor cell lines (Supplementary Figure S4), it is imperative to combat the multiple repair signals specifically hijacked by BCR-ABL1 oncoprotein in Ph+ B-cell precursor ALL cells using the potent HDAC1,2 inhibitor in order to achieve optimal clinical efficacy.
The MRN complex, comprised of Mre11–Rad50–Nbs1 subunits, functions as genome guardian by acting as a break sensor that tethers broken DNA ends in preparation for repair.39 Mre11 and Nbs1 levels are upregulated in BCR-ABL1+ cells.10 Mre11 contains exonuclease and endonuclease activities and functions in NHEJ and HR.40 Given these important functions, reduced and delayed binding of Mre11 levels at DSB sites agrees well with the DSB repair defects upon HDAC1,2 inhibition (Figure 3 and Supplementary Figure S3). BCR-ABL1, including the T315I mutant, promotes Rad51-mediated HR repair.9 Rad51, a downstream effector of MRN complex, performs homology searching during HR.41, 42 Reduced and delayed binding of Rad51 to chromatin and DSB sites following HDAC1,2 inhibitor and doxorubicin treatments would further curtail the downstream repair events of Mre11. Although DNA Lig1 is linked to Okazaki fragment maturation during DNA replication,43 it also acts in the final steps of HR.44 Decreased DNA Lig1 at DSB sites following ACY1035/doxorubicin treatment explains how HDAC1,2 inhibition impacts the final steps of DSB repair processes promoted by BCR-ABL1 in Ph+ B-cell precursor ALL cells. Thus one could propose that an HDAC1,2 selective inhibitor obviates the need to use a cocktail of DNA repair factor inhibitors and serves as a ‘pan’ disruptor of DSB repair at various levels. Thus, selective HDAC1,2 inhibition confers a therapeutic option not only for Ph+ B-cell precursor ALL but also potentially for any cancer dependent on or ‘addicted’ to DNA repair. Moreover, key molecular targets identified, such as Mre11 and Rad51, set the stage for their potential use as biomarkers in clinical setting to evaluate the efficacy of HDAC1,2 inhibitor and/or doxorubicin treatments.
We have also demonstrated for the first time that HDACs reprise their ‘classic’ role as transcriptional repressors even during DNA repair, as their activities are required for DISC or transcriptional repression at DSB sites during DNA repair via modulating the functions of chromatin remodeler Baf180 (Figure 5 and Supplementary Figure S14). Thus HDAC1,2 function to overcome the conflict between DNA repair and transcription machineries in order to promote the efficient joining of broken DNA. Our results also allude to the possibility that HDAC1,2 activity controls the chromatin association of other remodelers (TRRAP, CBX4/8, p400) and potentially influence chromatin structure during the access and/or restoration steps of DNA repair,45 which will be investigated in future studies. Interestingly, doxorubicin also interferes with DISC (Figure 5) even though it does not alter nucleosomal occupancy or chromatin association of Baf180. We speculate that altered DNA structure resulting from impaired topoisomerase activity might lead to reduced control of transcriptional repression around break sites, suggesting the presence of another layer of regulation of DNA repair involving DNA topology.
We postulate that HDAC1,2 inhibition and/or doxorubicin could cause the accumulation of DNA damage in two modes: one, they impair DNA replication or transcription, which could culminate in stalled replication forks20 or transcription bubbles (R-loops) to cause DNA damage (Supplementary Figure S19); and second, these compounds impair the subsequent repair of these broken DNA by inhibiting various key steps of DSB repair and lead to further accumulation of DNA damage (Figures 3, 4, 5). In conclusion, transient selective inhibition of HDAC1,2, either as a monotherapy or in combination with a low-dose doxorubicin, can be an excellent strategy with greater clinical efficacy and reduced toxicity for cancers addicted to DNA repair such as Ph+ B-cell precursor ALL.
References
Koretzky GA . The legacy of the Philadelphia chromosome. J Clin Invest 2007; 117: 2030–2032.
Mullighan CG . Molecular genetics of B-precursor acute lymphoblastic leukemia. J Clin Invest 2012; 122: 3407–3415.
Daver N, Thomas D, Ravandi F, Cortes J, Garris R, Jabbour E et al. Final report of a phase II study of imatinib mesylate with hyper-CVAD for the front-line treatment of adult patients with Philadelphia chromosome-positive acute lymphoblastic leukemia. Haematologica 2015; 100: 653–661.
Miller GD, Bruno BJ, Lim CS . Resistant mutations in CML and Ph(+)ALL - role of ponatinib. Biologics 2014; 8: 243–254.
Lee HZ, Kwitkowski VE, Del Valle PL, Ricci MS, Saber H, Habtemariam BA et al. FDA approval: belinostat for the treatment of patients with relapsed or refractory peripheral T-cell lymphoma. Clin Cancer Res 2015; 21: 2666–2670.
Liu-Dumlao T, Kantarjian H, Thomas DA, O'Brien S, Ravandi F . Philadelphia-positive acute lymphoblastic leukemia: current treatment options. Curr Oncol Rep 2012; 14: 387–394.
Podhorecka M, Skladanowski A, Bozko P . H2AX phosphorylation: its role in DNA damage response and cancer therapy. J Nucleic Acids 2010; 2010: 1–9.
Tobin LA, Robert C, Rapoport AP, Gojo I, Baer MR, Tomkinson AE et al. Targeting abnormal DNA double-strand break repair in tyrosine kinase inhibitor-resistant chronic myeloid leukemias. Oncogene 2013; 32: 1784–1793.
Slupianek A, Dasgupta Y, Ren SY, Gurdek E, Donlin M, Nieborowska-Skorska M et al. Targeting RAD51 phosphotyrosine-315 to prevent unfaithful recombination repair in BCR-ABL1 leukemia. Blood 2011; 118: 1062–1068.
Rink L, Slupianek A, Stoklosa T, Nieborowska-Skorska M, Urbanska K, Seferynska I et al. Enhanced phosphorylation of Nbs1, a member of DNA repair/checkpoint complex Mre11-RAD50-Nbs1, can be targeted to increase the efficacy of imatinib mesylate against BCR/ABL-positive leukemia cells. Blood 2007; 110: 651–660.
Skorski T . BCR/ABL regulates response to DNA damage: the role in resistance to genotoxic treatment and in genomic instability. Oncogene 2002; 21: 8591–8604.
Mann BS, Johnson JR, Cohen MH, Justice R, Pazdur R . FDA approval summary: vorinostat for treatment of advanced primary cutaneous T-cell lymphoma. Oncologist 2007; 12: 1247–1252.
Piekarz RL, Robey R, Sandor V, Bakke S, Wilson WH, Dahmoush L et al. Inhibitor of histone deacetylation, depsipeptide (FR901228), in the treatment of peripheral and cutaneous T-cell lymphoma: a case report. Blood 2001; 98: 2865–2868.
Laubach JP, Moreau P, San-Miguel JF, Richardson PG . Panobinostat for the treatment of multiple myeloma. Clin Cancer Res 2015; 21: 4767–4773.
Bates SE, Rosing DR, Fojo T, Piekarz RL . Challenges of evaluating the cardiac effects of anticancer agents. Clin Cancer Res 2006; 12: 3871–3874.
Shah MH, Binkley P, Chan K, Xiao J, Arbogast D, Collamore M et al. Cardiotoxicity of histone deacetylase inhibitor depsipeptide in patients with metastatic neuroendocrine tumors. Clin Cancer Res 2006; 12: 3997–4003.
Bhaskara S . Histone deacetylases 1 and 2 regulate DNA replication and DNA repair: potential targets for genome stability-mechanism-based therapeutics for a subset of cancers. Cell Cycle 2015; 14: 1779–1785.
Bhaskara S, Chyla BJ, Amann JM, Knutson SK, Cortez D, Sun ZW et al. Deletion of histone deacetylase 3 reveals critical roles in S phase progression and DNA damage control. Mol Cell 2008; 30: 61–72.
Bhaskara S, Hiebert SW . Role for histone deacetylase 3 in maintenance of genome stability. Cell Cycle 2011; 10: 727–728.
Bhaskara S, Jacques V, Rusche JR, Olson EN, Cairns BR, Chandrasekharan MB . Histone deacetylases 1 and 2 maintain S-phase chromatin and DNA replication fork progression. Epigenetics Chromatin 2013; 6: 27.
Bhaskara S, Knutson SK, Jiang G, Chandrasekharan MB, Wilson AJ, Zheng S et al. Hdac3 is essential for the maintenance of chromatin structure and genome stability. Cancer Cell 2010; 18: 436–447.
Johnson DP, Spitz GS, Tharkar S, Quayle SN, Shearstone JR, Jones S et al. HDAC1,2 inhibition impairs EZH2- and BBAP-mediated DNA repair to overcome chemoresistance in EZH2 gain-of-function mutant diffuse large B-cell lymphoma. Oncotarget 2015; 6: 4863–4887.
Shearstone JR, Golonzhka O, Chonkar A, Tamang D, van Duzer JH, Jones SS et al. Chemical inhibition of histone deacetylases 1 and 2 induces fetal hemoglobin through activation of GATA2. PLoS One 2016; 11: e0153767.
Wang Z, Yuan H, Roth M, Stark JM, Bhatia R, Chen WY . SIRT1 deacetylase promotes acquisition of genetic mutations for drug resistance in CML cells. Oncogene 2013; 32: 589–598.
Yang F, Teves SS, Kemp CJ, Henikoff S . Doxorubicin, DNA torsion, and chromatin dynamics. Biochim Biophys Acta 2014; 1845: 84–89.
Qi W, Wang R, Chen H, Wang X, Xiao T, Boldogh I et al. BRG1 promotes the repair of DNA double-strand breaks by facilitating the replacement of RPA with RAD51. J Cell Sci 2015; 128: 317–330.
Kakarougkas A, Ismail A, Chambers AL, Riballo E, Herbert AD, Kunzel J et al. Requirement for PBAF in transcriptional repression and repair at DNA breaks in actively transcribed regions of chromatin. Mol Cell 2014; 55: 723–732.
Klochendler-Yeivin A, Picarsky E, Yaniv M . Increased DNA damage sensitivity and apoptosis in cells lacking the Snf5/Ini1 subunit of the SWI/SNF chromatin remodeling complex. Mol Cell Biol 2006; 26: 2661–2674.
Chen YH, Jones MJ, Yin Y, Crist SB, Colnaghi L, Sims RJ 3rd et al. ATR-mediated phosphorylation of FANCI regulates dormant origin firing in response to replication stress. Mol Cell 2015; 58: 323–338.
Orlando DA, Chen MW, Brown VE, Solanki S, Choi YJ, Olson ER et al. Quantitative ChIP-Seq normalization reveals global modulation of the epigenome. Cell Rep 2014; 9: 1163–1170.
Dong X, Weng Z . The correlation between histone modifications and gene expression. Epigenomics 2013; 5: 113–116.
Price BD, D'Andrea AD . Chromatin remodeling at DNA double-strand breaks. Cell 2013; 152: 1344–1354.
Ma RG, Zhang Y, Sun TT, Cheng B . Epigenetic regulation by polycomb group complexes: focus on roles of CBX proteins. J Zhejiang Univ Sci B 2014; 15: 412–428.
Chapman JR, Taylor MR, Boulton SJ . Playing the end game: DNA double-strand break repair pathway choice. Mol Cell 2012; 47: 497–510.
Huen MS, Chen J . ATM creates a veil of transcriptional silence. Cell 2010; 141: 924–926.
Shanbhag NM, Rafalska-Metcalf IU, Balane-Bolivar C, Janicki SM, Greenberg RA . ATM-dependent chromatin changes silence transcription in cis to DNA double-strand breaks. Cell 2010; 141: 970–981.
Montgomery RL, Davis CA, Potthoff MJ, Haberland M, Fielitz J, Qi X et al. Histone deacetylases 1 and 2 redundantly regulate cardiac morphogenesis, growth, and contractility. Genes Dev 2007; 21: 1790–1802.
Ho CY, Lopez B, Coelho-Filho OR, Lakdawala NK, Cirino AL, Jarolim P et al. Myocardial fibrosis as an early manifestation of hypertrophic cardiomyopathy. N Engl J Med 2010; 363: 552–563.
Lamarche BJ, Orazio NI, Weitzman MD . The MRN complex in double-strand break repair and telomere maintenance. FEBS Lett 2010; 584: 3682–3695.
Zha S, Boboila C, Alt FW . Mre11: roles in DNA repair beyond homologous recombination. Nat Struct Mol Biol 2009; 16: 798–800.
Jasin M, Rothstein R . Repair of strand breaks by homologous recombination. Cold Spring Harb Perspect Biol 2013; 5: a012740.
Baumann P, West SC . Role of the human RAD51 protein in homologous recombination and double-stranded-break repair. Trends Biochem Sci 1998; 23: 247–251.
Zheng L, Shen B . Okazaki fragment maturation: nucleases take centre stage. J Mol Cell Biol 2011; 3: 23–30.
Goetz JD, Motycka TA, Han M, Jasin M, Tomkinson AE . Reduced repair of DNA double-strand breaks by homologous recombination in a DNA ligase I-deficient human cell line. DNA Repair 2005; 4: 649–654.
House NC, Koch MR, Freudenreich CH . Chromatin modifications and DNA repair: beyond double-strand breaks. Front Genet 2014; 5: 296.
Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-Andre V, Sigova AA et al. Super-enhancers in the control of cell identity and disease. Cell 2013; 155: 934–947.
Acknowledgements
We thank Dr Roger Greenberg for the U2OS/Fok1 reporter cell line. This work was supported by funds from the University of Utah Research Foundation Seed Grant (51900230), HCI/Radiation Oncology development funds and NIH R01 grant (R01-CA188520) to SB.
Author contributions
STP, DPJ, SEB, EMD and BGB performed the experiments and analyzed the data. SSJ, JRS, SNQ, CM and MJ provided the HDAC1,2 selective inhibitors and performed pharmacological studies. TM performed bioinformatics. RRM performed histopathology. WYC provided the DNA repair reporter cell lines. ADP and MWD provided patient samples, critical comments and clinical perspective. DCS, MBC, KNB and PAZ-M provided valuable suggestions. SB conceived the idea, obtained funding, performed experiments, analyzed the data and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
SNQ, JRS and CM were employees of and owned equity in Acetylon Pharmaceuticals. SSJ and MJ are current employees of Regenacy Pharmaceuticals. The other authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Tharkar-Promod, S., Johnson, D., Bennett, S. et al. HDAC1,2 inhibition and doxorubicin impair Mre11-dependent DNA repair and DISC to override BCR-ABL1-driven DSB repair in Philadelphia chromosome-positive B-cell precursor acute lymphoblastic leukemia. Leukemia 32, 49–60 (2018). https://doi.org/10.1038/leu.2017.174
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2017.174
This article is cited by
-
Genome-wide CRISPR screen identified Rad18 as a determinant of doxorubicin sensitivity in osteosarcoma
Journal of Experimental & Clinical Cancer Research (2022)
-
Novel pH-responsive nanohybrid for simultaneous delivery of doxorubicin and paclitaxel: an in-silico insight
BMC Chemistry (2021)
-
Advances in histone deacetylase inhibitors in targeting glioblastoma stem cells
Cancer Chemotherapy and Pharmacology (2020)