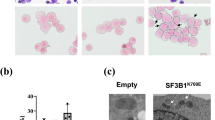
Abstract
The splicing factor SF3B1 is the most frequently mutated gene in myelodysplastic syndromes (MDS), and is strongly associated with the presence of ring sideroblasts (RS). We have performed a systematic analysis of cryptic splicing abnormalities from RNA sequencing data on hematopoietic stem cells (HSCs) of SF3B1-mutant MDS cases with RS. Aberrant splicing events in many downstream target genes were identified and cryptic 3′ splice site usage was a frequent event in SF3B1-mutant MDS. The iron transporter ABCB7 is a well-recognized candidate gene showing marked downregulation in MDS with RS. Our analysis unveiled aberrant ABCB7 splicing, due to usage of an alternative 3′ splice site in MDS patient samples, giving rise to a premature termination codon in the ABCB7 mRNA. Treatment of cultured SF3B1-mutant MDS erythroblasts and a CRISPR/Cas9-generated SF3B1-mutant cell line with the nonsense-mediated decay (NMD) inhibitor cycloheximide showed that the aberrantly spliced ABCB7 transcript is targeted by NMD. We describe cryptic splicing events in the HSCs of SF3B1-mutant MDS, and our data support a model in which NMD-induced downregulation of the iron exporter ABCB7 mRNA transcript resulting from aberrant splicing caused by mutant SF3B1 underlies the increased mitochondrial iron accumulation found in MDS patients with RS.
Similar content being viewed by others
Introduction
The myelodysplastic syndromes (MDS) are a heterogeneous group of clonal hematopoietic stem cell (HSC) malignancies characterized by ineffective hematopoiesis leading to peripheral blood cytopenias, and show increasing bone marrow blasts.1 The MDS show frequent progression (approximately 40% of patients) to acute myeloid leukemia. Several genes involved in pre-messenger RNA splicing, including SF3B1, U2AF1, SRSF2 and ZRSR2,2, 3, 4 have been shown to be mutated in over 50% of MDS patients, revealing a new leukemogenic pathway involving spliceosomal dysfunction.
The splicing factor SF3B1 is the most frequently mutated gene in patients with MDS (20–28% of all cases).5, 6 Mutations of SF3B1 occur in a high proportion (>80%) of MDS patients in whom the presence of ring sideroblasts (RS) is a characteristic disease feature, namely the refractory anemia with RS (RARS) and refractory cytopenia with multilineage dysplasia and RS (RCMD-RS) subtypes.5, 7 In the recent 2016 revision of the World Health Organization (WHO) classification for MDS, if a patient harbors an SF3B1 mutation, a diagnosis of MDS with RS (MDS-RS) may be made if 5–14% RS are present in the bone marrow.8 SF3B1 mutations are closely associated with the presence of RS, suggesting a causal relationship and making SF3B1 the first gene showing a strong association with a particular morphological feature in MDS.5 RS are erythroblasts with excessive mitochondrial iron accumulation,9 and RARS patients with SF3B1 mutation have altered iron distribution characterized by coarse iron deposits in comparison with RARS patients without SF3B1 mutation.10 SF3B1 mutations occur more frequently in low-risk MDS cases and are independent predictors of favorable survival in MDS.5 The clinical consequences of mutations in SF3B1 are well documented in MDS, however the functional consequences of SF3B1 mutations in human hematopoietic cells are not fully understood.
A well-recognized candidate gene for MDS with the RS phenotype is the iron transporter ABCB7. Our group first reported marked downregulation of ABCB7 in MDS patients with RARS subtype.11 Hereditary X-linked sideroblastic anemia with ataxia is caused by partial loss-of-function mutations of ABCB7, which inhibit heme biosynthesis.12 Moreover, knockdown of ABCB7 in HeLa cells resulted in an iron-deficient phenotype with mitochondrial iron accumulation.13 Conditional gene targeting in mice has shown that ABCB7 is essential for hematopoiesis.12
SF3B1 is a core component of the U2-small nuclear ribonucleoprotein complex and is involved in stabilizing the interaction of the U2-small nuclear ribonucleoprotein with the branch point (BP),14 upstream of the 3′ splice site. SF3B1 also interacts with other spliceosomal proteins such as U2AF2, which binds the polypyrimidine tract (PPT) downstream of the BP.15, 16 Base-pairing of U2 snRNA with the pre-messenger RNA bulges out the BP adenosine, specifying it as the site to initiate the nucleophilic attack in the first step of splicing. The binding of the SF3B complex proteins around the BP prevents the premature activity at the site before the fully active spliceosome is assembled.17 The role of SF3B1 and the U2-small nuclear ribonucleoprotein in recognizing and binding the BP suggest that SF3B1 mutations may alter BP and/or 3′ splice site selection.
The splicing factor genes found to be mutated in MDS code for proteins that have a role in the recognition of 3′ splice sites during processing of pre-messenger RNAs.3 Altered RNA splicing has been suggested as the mechanism underlying the observed phenotypic changes concomitant to splicing factor gene mutations, including SF3B1, and the identification of aberrantly spliced target genes in the hematopoietic cells of SF3B1-mutant MDS cases is important.
A number of studies to date have used RNA sequencing (RNA-Seq) on unfractionated bone marrow mononuclear cells from a small number of SF3B1-mutant MDS patients.10, 18, 19 MDS is a disorder of the HSC and we thus studied the transcriptome of CD34+ cells from MDS patients with SF3B1 mutations using RNA-Seq. We have recently identified many genes significantly differentially expressed at the transcript and/or exon level in bone marrow CD34+ cells of SF3B1-mutant MDS compared with wild-type and healthy control cases.20
Recently, SF3B1 mutations have been identified in various tumor types, suggesting that somatic mutations in spliceosome genes have an important role in tumorigenesis.21, 22, 23, 24 SF3B1 mutations have been shown to occur in chronic lymphocytic leukemia, uveal melanoma, breast cancer and pancreatic cancer.24, 25 SF3B1 mutations have clear mutational hotspots and are considered to be gain-of-function/neomorphic mutations.2, 3, 26, 27 The codons most commonly affected by SF3B1 mutations in other cancers that harbor this mutation, including chronic lymphocytic leukemia, uveal melanoma, breast cancer and pancreatic cancer, are the same as the ones affected in MDS (K700, R625 and K666). Recent studies of chronic lymphocytic leukemia, breast cancer and uveal melanoma using RNA-Seq have shown that SF3B1 mutations are associated with differential exon usage and induce cryptic alternative 3′ splice site selection in these cancers.28 However, a systematic analysis of cryptic splicing abnormalities has not been performed in MDS HSCs.
In this study, we have performed an analysis of RNA-Seq data on HSCs of SF3B1-mutant MDS cases to identify aberrant/cryptic splicing events. The identification of the splicing aberrations induced by SF3B1 mutation in the HSCs of MDS patients will shed light on the downstream effects that lead to the MDS phenotype and may allow for the identification of new therapeutic targets in this disease.
Materials and methods
Samples and RNA-Seq
RNA-Seq data were obtained from CD34+ cells isolated from bone marrow samples of eight MDS patients (four RARS and four RCMD-RS) with SF3B1 mutation (four K700E, one E622D, one R625L, one H662Q and one K666R; 45–52% SF3B1-mutant allele expression range), four MDS cases (all refractory cytopenia with multilineage dysplasia) without mutations in the splicing factor genes SF3B1, SRSF2, U2AF1 and ZRSR2, and five healthy individuals.20 CD34+ cells were isolated from bone marrow samples of the 12 MDS patients and five healthy controls using magnetic-activated cell sorting columns (Miltenyi Biotec, Bergisch Gladbach, Germany), according to the manufacturer’s recommendations.
DNase-treated (Invitrogen, Carlsbad, CA, USA) total RNA was purified using XP beads (Beckman Coulter, High Wycombe, UK), and library preparation was performed using the NEBNext Ultra directional RNA Library prep kit (NEB, Hitchin, UK) following the manufacturer’s recommendations. Custom indexes were used and samples were purified using XP beads (Beckman Coulter) instead of size selection. Sequencing was performed on an Illumina HiSeq2000 instrument (Illumina, San Diego, CA, USA).
RNA-Seq data analysis
Following QC analysis with the fastQC package (http://www.bioinformatics.babraham.ac.uk/projects/fastqc), reads were aligned using STAR29 against the human genome assembly (NCBI build37 (hg19) UCSC transcripts). Non-uniquely mapped reads and reads that were identified as PCR duplicates using Samtools30 were discarded. The aligned reads were reconstructed into transcripts using Cufflinks31 and were then merged into a single assembly, along with known isoforms from the NCBI build37 (hg19) UCSC transcripts. This reference-guided assembly was then used as the transcripts annotation by rMATS.
Alternative 3′ and 5′ splice sites, skipped exons, mutually exclusive exons and retained introns were quantified using rMATS32 with the assembly produced from Cufflinks. The default parameters were used for the comparison of the samples. The results were filtered using a false discovery rate<0.05 and inclusion level difference values >0.3 or <−0.3. The alternative spliced events were then plotted using the sashimi plots of the MISO software.33 The results were visualized and filtered using the data visualization tool Zegami (http://zegami.com/).
Gene ontology analysis
Analysis of 3′ splice site properties
End point RT-PCR validation of aberrant splicing isoforms
Six genes (including ABCB7) with cryptic 3′ splice sites were selected from the rMATS data analysis for end point reverse transcriptase-PCR (RT-PCR) validation. See Supplementary Methods for details.
Erythroblast cell culture and cycloheximide treatment
Bone marrow CD34+ cells from healthy individuals were purchased from Lonza (Basel, Switzerland). Bone marrow samples were obtained and CD34+ cells were isolated from two MDS patients with SF3B1 K700E mutation. CD34+ cells from MDS patients and from healthy controls were cultured as previously described.34, 35 On day 11 and day 14, an aliquot of cells were treated with 100 μg/ml cycloheximide for 4 h and subsequently collected for RNA extraction. Total RNA was reverse-transcribed using High capacity cDNA reverse transcription kit (Applied Biosystems, Foster City, CA, USA). The expression of aberrantly spliced ABCB7 was determined by RT-PCR as described in the Supplementary Methods.
Generation of SF3B1-mutant K562 cells by CRISPR/Cas9 and cycloheximide treatment
Pancreatic cell line Panc 05.04 culture and cycloheximide treatment
Results
Cryptic splicing events in HSCs of SF3B1-mutant MDS
We have analyzed RNA-Seq data obtained from the CD34+ cells from eight MDS cases harboring SF3B1 mutations (SF3B1-mutant, all with >15% RS), four MDS patients without splicing factor gene mutations (wild type) and five healthy individuals (control)20 using rMATS, a bioinformatics pipeline designed to detect alternative (including cryptic) splicing events involving two isoforms from an alternatively spliced region.32 These events are categorized as alternative 3′ splice site (A3SS) usage, alternative 5′ splice site (A5SS) usage, exon skipping, mutually exclusive exons or retained introns.
When comparing SF3B1-mutant to wild type, we identified 126 significant splicing events (92 genes), of which 42 were A3SS, 6 A5SS, 8 mutually exclusive exons, 13 skipped exons and 57 retained introns (Tables 1 and 2, Supplementary Table S1). When comparing SF3B1-mutant with controls, 213 significant splicing events (164 genes) were identified, of which 62 were A3SS, 10 A5SS, 12 mutually exclusive exon, 12 skipped exons and 117 retained intron (Tables 1 and 3, Supplementary Table S2). Top-ranking significant genes showing at least one cryptic splicing event in both of these comparisons include TMEM14C, ENOSF1, SEPT6, DYNLL1, HINT2 and ABCC5.
We performed gene ontology analysis on the lists of significant genes showing aberrant splicing events identified by the rMATS pipeline using GOseq. The significant main ontology themes for the comparison of SF3B1-mutant with wild type and controls include ‘RNA processing’ and ‘RNA splicing’ (Supplementary Tables S3 and S4).
Properties of misregulated alternative 3′ splice sites
Compared with all alternative splicing events (regulated and unregulated) detected in the data by rMATS, we found a significant overrepresentation of regulated A3SS (P<1 × E-08, χ2-test with Yates's continuity correction), but no significant overrepresentation of alternative 5′ splice sites, in the comparison of SF3B1-mutant with both wild type and controls (Table 1). Indeed, 15/20 (75%) most significant aberrant splicing events in the comparison of SF3B1-mutant with wild type (Table 2) and 13/20 (65%) most significant aberrant splicing events in the comparison of SF3B1-mutant with controls (Table 3) were A3SS. These data are in accord with the known function of SF3B1 in the recognition of BPs and 3′ splice sites. In addition to A3SS, retained introns were also significantly overrepresented, while cassette and mutually exclusive exons were underrepresented.
The majority of the regulated A3SS events involved use of an A3SS upstream of the canonical 3′ splice site (Table 1). Analysis of the sequences of upstream and downstream cryptic 3′ splice sites, along with their associated canonical 3′ splice sites revealed distinct sequence features. Both sets of canonical 3′ splice sites had extensive 18–20 nt PPTs with enrichment of uridines at most positions (Figure 1a, Supplementary Figure S1A). In contrast, the upstream cryptic sites had a shorter PPT of ~8 nt with a stretch of 4–5A residues 13–17 nt upstream. Downstream cryptic sites in the comparison of SF3B1-mutant with wild type also had more purine interruptions (Figure 1a), but this was not evident in the comparison of SF3B1-mutant with control (Supplementary Figure S1A). Analysis of the distance separating pairs of A3SS revealed a very distinct pattern for upstream cryptic 3′ splice sites, with a strong peak at 15nt (Figure 1b, Supplementary Figure S1B, blue trace; 15nt=3.9 on log2 scale). In contrast, downstream cryptic 3′ splice sites and A3SS unregulated by the SF3B1 mutation showed a much broader distribution of spacing with most pairs of A3SS being much more widely spaced (red and green traces in Figure 1b and Supplementary Figure S1B). One interesting exception was a peak at 3 nt (1.6 on log2 scale) for the unregulated A3SS, which corresponds to the so-called NAGNAG class of A3SS,36 selection of which occurs at step 2 of splicing37 consistent with a lack of effect of SF3B1 mutation. Comparison of 3′ splice site strength showed that both upstream and downstream cryptic 3′ splice sites were weaker than their associated canonical sites in the comparison of SF3B1-mutant with wild type (Figure 1c), but in the comparison of SF3B1-mutant with controls, only the upstream cryptic sites were weaker (Supplementary Figure S1C). This is consistent with the adenosine interruptions of the PPT in the upstream cryptic sites (Figure 1a, Supplementary Figure S1A). We used SVM-BP finder38 to predict the location and strength of the top-scoring predicted BPs associated with each A3SS. Although the predicted BP scores did not differ significantly between sets of regulated cryptic 3′ splice site and their associated canonical sites (Figure 1d, Supplementary Figure S1D), top-scoring BPs for upstream cryptic sites were closer (median 18nt, BP-AG distance) than those for their associated canonical sites (median 27nt) or unregulated A3SS (median 26nt) (Figure 1e, Supplementary Figure S1E). Taken together, our data show that MDS-associated SF3B1 mutations result in widespread use of cryptic 3′ splice sites a short distance upstream of canonical sites and could be consistent either with use of a common BP in association with upstream cryptic and canonical splice site pairs28 or, more likely, with shifted use of both BP and consequently 3′ splice site in the SF3B1-mutant,39, 40 as shown in recent reports in other cancers.
Properties of A3SS misregulated in SF3B1-mutant compared with wild-type SF3B1 MDS HSCs. (a) Sequence logos for upstream and downstream cryptic 3′ splice sites along with their associated canonical sites. (b) Density plot showing distance (log2) between pairs of 3′ splice sites. Blue line: upstream cryptic. Red line: downstream cryptic. Green line: A3SS unaffected by SF3B1 mutation. (c) 3′ splice site strengths (maximum entropy) for upstream and downstream control (unregulated) A3SS, upstream cryptic sites and their associated canonical sites, and downstream cryptic sites and their associated canonical sites. (d) BP scores. (e) Distance of highest scoring predicted BP from associated 3′ splice site. (*P<0.05, **P<0.01).
Validation of cryptic 3′ splice site events
Several of the misregulated A3SS events identified by the rMATS pipeline were validated using RT-PCR in patient and healthy control samples. We chose TMEM14C, SEPT6, HINT2, DYNLL1 and ENOSF1 for validation. These genes all showed upstream cryptic 3′ splice sites in the rMATS analysis with the cryptic AG site located between 11 and 17 nucleotides upstream of the canonical site (Supplementary Figure S2). Aberrant splicing of TMEM14C and DYNLL1 introduces an addition of 14 base pairs in the 5′UTR region of these genes. The cryptic splice site event in ENOSF1 leads to the addition of 15 base pairs (encoding five amino acids) to the coding sequence. Aberrant splicing of SEPT6 and HINT2 introduces an addition of 17 and 11 base pairs, respectively, leading to a frameshift in SEPT6 and to the rise of a premature termination codon in the first 3 base pairs of exon 5 in HINT2. The RT-PCR results confirmed the cryptic splicing events identified by RNA-Seq in these five genes in MDS patient samples with SF3B1 mutation (Figures 2a and b).
Validation and semi-quantification of cryptic 3′ splice site usage in five selected genes from the rMATS analysis in SF3B1-mutant MDS HSC samples. (a) PCR products of the five genes (TMEM14C, ENOSF1, DYNLL1, SEPT6 and HINT2) were amplified from SF3B1-mutant MDS samples (Mut), wild-type MDS samples (WT) and healthy control samples (HC) run on an Agilent 2100 Bioanalyzer instrument using the DNA 1000 kit. The aberrant transcripts (indicated by the higher band) were observed in the SF3B1-mutant MDS samples. (b) Semi-quantification of the PCR bands showed high cryptic to canonical isoform ratio in the SF3B1-mutant MDS samples.
Cryptic splicing of ABCB7 in HSCs of SF3B1-mutant MDS
We previously reported marked downregulation of the iron transporter ABCB7 in MDS patients with RS.11 In this study, we identified a significant A3SS event (false discovery rate=0.006, inclusion level difference=−0.184) in the ABCB7 gene in the comparison of SF3B1-mutant with controls. Aberrant splicing introduces an addition of 21 base pairs from the intronic region between exons 8 and 9 causing an addition of seven amino acids to the protein sequence with the last 3 base pairs proximal to the canonical exon 9 giving rise to a premature termination codon (Figures 3a and b). This result was confirmed by RT-PCR in patient and healthy control HSC samples: the aberrant ABCB7 transcript was observed in all SF3B1-mutant MDS samples analyzed (n=7), but not in any of the samples from wild-type patients (n=4) or from healthy controls (n=5) (Figure 4a). Sanger sequencing of gel extract bands confirmed the presence of the addition of 21 base pairs in the aberrant ABCB7 transcript (Figure 3b). These data demonstrate that aberrant splicing of ABCB7 observed is specific to MDS cases carrying mutation of SF3B1.
Cryptic 3′ splice site usage of ABCB7 in SF3B1-mutant MDS HSCs. (a) Visualization of RNA-Seq traces for the ABCB7 gene (intron 8-exon 9 junction) in two healthy controls, two MDS patients with no known splicing factor mutations and two SF3B1-mutant MDS cases, using Integrative Genomics Viewer (IGV). A cryptic 3′ splice site is observed in the ABCB7 gene between exon 8 and exon 9 in the SF3B1-mutant MDS cases, leading to an addition of 21 base pairs to the coding sequence causing a premature termination codon at the seventh amino acid (indicated with *). (b) Sanger sequencing traces of gel extracted bands corresponding to the canonical ABCB7 transcript and to the aberrant ABCB7 transcript containing the additional 21 base pairs (highlighted in blue).
Aberrant splicing of ABCB7 in HSCs and cultured erythroblast from SF3B1-mutant MDS samples, in the K562-SF3B1K700E cell line and a SF3B1-mutant pancreatic cell line. (a) RT-PCR confirmation of aberrant splicing of ABCB7 in SF3B1-mutant MDS HSC samples. The higher 174 bp band corresponds to the aberrant ABCB7 transcript and the lower 153 bp band corresponds to the canonical ABCB7 transcript. The aberrant ABCB7 transcript (higher 174 bp band) was observed in the SF3B1-mutant MDS samples (Mut), but not in samples from wild-type MDS patients (WT) or in samples from healthy controls (HC). (b) Aberrant splicing of ABCB7 in cultured erythroblast from SF3B1-mutant MDS patients. Each panel shows data from one different SF3B1-mutant MDS patient (Mut) and from one different healthy control (HC). RT-PCR for ABCB7 was performed on cultured erythroblasts at day 11 and day 14 of culture treated with and without the NMD inhibitor cycloheximide (CHX). The higher 174 bp band corresponds to the aberrant ABCB7 transcript and the lower 153 bp band corresponds to the canonical ABCB7 transcript. The RT-PCR results showed an increase in the product corresponding to the aberrant ABCB7 transcript (higher 174 bp band) in the SF3B1-mutant patient samples treated with CHX compared with untreated samples. No aberrant ABCB7 splicing was seen in the samples from healthy controls. (c) Aberrant splicing of ABCB7 in K562-SF3B1K700E. RT-PCR for ABCB7 was performed on cultured K562-SF3B1K700E and K562-SF3B1WT cells treated with and without the NMD inhibitor CHX. The higher 174 bp band corresponds to the aberrant ABCB7 transcript and the lower 153 bp band corresponds to the canonical ABCB7 transcript. The RT-PCR results showed an increase in the product corresponding to the aberrant ABCB7 transcript (higher 174 bp band) in the K562-SF3B1K700E cells treated with cycloheximide (CHX) compared with the untreated cells. No aberrant ABCB7 splicing was seen in the K562-SF3B1WT cells untreated or treated with CHX. (d) Aberrant splicing of ABCB7 in Panc 05.04 cells with SF3B1 mutation (K700E). RT-PCR for ABCB7 was performed on cultured Panc 05.04 cells treated with and without the NMD inhibitor CHX. The higher 174 bp band corresponds to the aberrant ABCB7 transcript and the lower 153 bp band corresponds to the canonical ABCB7 transcript. The RT-PCR results showed an increase in the product corresponding to the aberrant ABCB7 transcript (higher 174 bp band) in the Panc 05.04 cells treated with CHX compared with the untreated cells.
NMD targets the aberrantly spliced ABCB7 transcript in SF3B1-mutant MDS erythroblasts, isogenic K562-SF3B1K700E and a SF3B1-mutant pancreatic cell line
The presence of premature termination codons can lead to degradation of mRNA transcripts by nonsense-mediated RNA decay (NMD).41 We thus investigated whether the aberrantly spliced ABCB7 transcript containing a premature termination codon that we identified in SF3B1-mutant MDS is affected by NMD. CD34+ cells from two SF3B1-mutant MDS RARS patients and from two healthy controls were cultured using a method developed to study the generation of erythroblasts.34, 35 Cells were collected at day 11 and day 14 of erythroid culture and treated with cycloheximide (an inhibitor of protein biosynthesis known to impair NMD42), and subjected to RT-PCR for ABCB7. The RT-PCR results showed an increase in the product corresponding to the aberrant ABCB7 transcript in the SF3B1-mutant patient samples treated with cycloheximide compared with untreated samples (Figure 4b), indicating that NMD targets the aberrantly spliced ABCB7 transcript and underlies the downregulation of ABCB7 observed in SF3B1-mutant MDS patients.
In addition, we performed RT-PCR for ABCB7 in K562-SF3B1K700E and SF3B1WT isogenic cells obtained using CRISPR/Cas9 gene editing and the pancreatic cell line Panc 05.04, which has a heterozygous SF3B1 K700E mutation, and we observed the same ABCB7 cryptic 3′ splice site event (Figures 4c and d), which introduces a stop codon as identified in the CD34+ cells and cultured erythroblasts of SF3B1-mutant MDS patients. Treatment of K562-SF3B1K700E and Panc 05.04 cells with cycloheximide resulted in an increase of the aberrantly spliced form of the ABCB7 transcript (Figures 4c and d).
Discussion
The splicing factor SF3B1 is the most frequently mutated gene found in MDS, and is strongly associated with the RS phenotype.5, 6 It is still unknown how SF3B1 mutations lead to the formation of RS in MDS. Given the critical functions of SF3B1 on 3′ splice site recognition, the probable consequence of this spliceosome mutation is aberrant splicing of various downstream target genes in MDS. The MDS arise in the HSC population in the bone marrow and SF3B1 is a founder mutation.43, 44 It is important to study the impact of SF3B1 mutation on the transcriptome in the cell of origin. We sought to identify the aberrant/cryptic mRNA splicing events associated with the SF3B1 mutation in the HSCs of MDS patients. We performed an analysis of RNA-Seq data that we generated from CD34+ cells isolated from bone marrow samples of SF3B1-mutant MDS patients with RS, MDS patients with no splicing factor gene mutation and from healthy individuals.20
The recent identification of A3SS usage in other malignancies with SF3B1 mutation22, 28 highlights the importance of interrogating RNA-seq data using a method that allows the identification of not only annotated alternative splicing events, but also of de novo cryptic splicing events. In this study, we used de novo transcriptome reconstruction and the rMATS pipeline to identify unannotated alternative splicing events, including A3SS and retained introns. We have found that SF3B1 mutations are associated with various aberrant splicing events in the HSCs of MDS patients. Interestingly, we identified significant cryptic 3′ splice site usage affecting many genes when comparing the transcriptome of SF3B1-mutant MDS cases with that of MDS wild-type and control cases, in accord with the known role of SF3B1 in the recognition of 3′ splice sites. A3SS events were significantly overrepresented in both comparisons, but no significant overrepresentation of alternative 5′ splice sites was observed. Indeed, the majority (65–75%) of the most significant aberrant splicing events in the comparison of SF3B1-mutant with both wild type and controls were A3SS. Furthermore, the number of genes with A3SS was sixfold to sevenfold higher than the number of genes with alternative 5′ splice sites. These data show that aberrant 3′ splice site selection is a frequent and important event associated with SF3B1 mutations in the HSCs of MDS patients.
The most distinct group of misregulated events in SF3B1-mutant MDS involved the use of cryptic 3′ splice sites located 15–20 nt upstream of the canonical 3′ splice site, consistent with recent reports in other cancers.28, 39, 40 This is an unusual location as the typical 3′ splice site arrangement comprises a BP typically 18–40 nt upstream of the 3′ splice site, with an optimal separation of 19–23 nt45 and a minimal separation of ~12 nt.37 The 15 nt separation of A3SS could therefore be consistent with a single BP-PPT unit from which either of two 3′ splice sites can be used,28 similar to Drosophila SXL exon 3.46 Alternatively, SF3B1 mutations could lead to a shift in BP selection a short distance upstream leading to altered 3′ splice site selection.39, 40 For the six events that we investigated, the predicted and mapped BP locations support the second scenario in which altered BP selection drives the change in 3′ splice site selection in MDS (Supplementary Figure S2). This would be consistent with the recruitment of U2-small nuclear ribonucleoprotein to positions 5′ of the BP in response to SF3B1 targeting drugs.47 Emerging data suggest that myeloid malignancies with splicing factor gene mutations are preferentially susceptible to additional splicing perturbations induced by splicing factor inhibitors48 and this may also represent a therapeutic approach in SF3B1-mutant MDS.
In our study, we found at least one cryptic splicing event in TMEM14C, ENOSF1, DYNLL1, SEPT6 and HINT2 (validated by RT-PCR) when comparing SF3B1-mutant to wild type and to controls. Importantly, cryptic splicing events affecting TMEM14C, ENOSF1, DYNLL1 and HINT2 have been associated with SF3B1 mutation in other cancers such as chronic lymphocytic leukemia, uveal melanoma and breast cancer.21, 22, 23 Emerging evidence thus suggests that there are several common downstream target genes in SF3B1-mutant malignancies, which may have implications for the design of new therapies for this group of cancers.
It is most probable that several target genes showing cryptic splicing contribute to the phenotype observed in MDS patients with the SF3B1 mutation. We note for example that TMEM14C has an important role in the terminal steps of the heme synthesis pathway.49
MDS patients with RARS suffer from a refractory anemia and show erythroid hyperplasia and ineffective erythropoiesis as a result of increased apoptosis in the bone marrow.50 RS are characterized by excessive iron accumulation in the mitochondria of erythroid progenitors.9 The close association between SF3B1 mutations and RS is consistent with a causal relationship, and makes this the first gene to be strongly associated with a specific feature of MDS.5 How SF3B1 mutations affect formation of RS is still unknown.
Cryptic splicing of genes involved in iron homeostasis and/or hemoglobin synthesis could have a role in the ineffective erythropoiesis observed in MDS patients with SF3B1 mutation and RS.51 We reported some years ago a strong relationship between an increasing percentage of bone marrow RS in MDS patients and decreasing expression levels of the iron transporter ABCB7.11 The ABCB7 gene is the functional ortholog of the yeast Atm1p gene,52 which has been shown to be required for mitochondrial iron homeostasis53 and is involved in the transport of a component required for the maturation of iron-sulfur cluster proteins from the mitochondria to the cytosol.12 Functional studies showing that forced expression of ABCB7 can restore erythroid growth and survival of RARS progenitors while decreasing the expression of aberrant mitochondrial ferritin (a marker for aberrant iron accumulation) subsequently implicated ABCB7 in the phenotype of acquired sideroblastic anemia (RARS).54 In a recent study, we performed an integrative analysis in MDS and the strongest association found was between the presence of SF3B1 mutations and marked downregulation of ABCB7.55 Given the strong correlation between SF3B1 mutations and the presence of RS,5 our data suggested a three-way association among SF3B1 mutation, ABCB7 downregulation and the occurrence of RS. Therefore, the marked downregulation of ABCB7 observed in MDS patients with RARS has been recognized as an important finding for several years; however, the mechanism underlying the downregulation of this gene in MDS has remained a mystery.
In the current study, the use of a pipeline that can detect cryptic splicing events has enabled the identification of A3SS usage of ABCB7 in the HSC of SF3B1-mutant MDS patients. Importantly, this event leads to aberrant splicing of the ABCB7 mRNA transcript, resulting in the addition of seven amino acids, including a premature termination codon, to the protein sequence in patient samples. It is recognized that the presence of premature termination codons can lead to degradation of the mRNA transcript by NMD,41 and we hypothesized that this event underlies the marked downregulation of ABCB7 observed in the HSCs of SF3B1-mutant MDS patients. This mechanism is strongly supported by our data on cultured SF3B1-mutant MDS erythroblasts treated with the NMD inhibitor cycloheximide, showing that the aberrantly spliced ABCB7 transcript is targeted by NMD in erythroid cells. Importantly, we next showed that the aberrantly spliced ABCB7 transcript was present in the myeloid cell line K562 in which the SF3B1 mutation was introduced using CRISPR/Cas9 gene editing, and in the pancreatic cell line Panc 05.04 (which is mutant for SF3B1). Treatment of these two cell lines with cycloheximide resulted in an increase of the aberrantly spliced form of the ABCB7 transcript. These data provide strong evidence that the SF3B1 mutation leads to aberrant ABCB7 splicing and downregulation via NMD in human myeloid cells and other cancer cells. Aberrant splicing of ABCB7 has been reported recently in an isogenic SF3B1-mutant pre-B ALL cell line in another study.39 We suggest that downregulation of the iron exporter ABCB7 resulting from aberrant splicing of the mRNA transcript leading to NMD underlies the increased mitochondrial iron accumulation found in MDS patients with RS. Our data provide an important link between inherited and acquired forms of sideroblastic anemia.
It is possible that the detection of aberrantly spliced target genes, in particular ABCB7, by RT-PCR could form the basis of a new diagnostic test for SF3B1-mutated MDS and may provide valuable information in cases with suspected MDS.
Our study is the first to describe the cryptic splicing events that occur in the hematopoietic progenitor cells of SF3B1-mutant MDS. These data illuminate the downstream target genes that may have a role in the development of the MDS phenotype, and further our understanding of the effect of SF3B1 mutations on splicing in malignancy. We demonstrate a mechanism linking the presence of SF3B1 mutation in MDS RARS patients and the NMD-induced marked downregulation of the iron transporter ABCB7, and provide strong evidence supporting a critical role of ABCB7 in the development of the RS phenotype. ABCB7 might represent a therapeutic target in MDS with RS.
References
Heaney ML, Golde DW . Myelodysplasia. N Engl J Med 1999; 340: 1649–1660.
Papaemmanuil E, Cazzola M, Boultwood J, Malcovati L, Vyas P, Bowen D et al. Somatic SF3B1 mutation in myelodysplasia with ring sideroblasts. N Engl J Med 2011; 365: 1384–1395.
Yoshida K, Sanada M, Shiraishi Y, Nowak D, Nagata Y, Yamamoto R et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature 2011; 478: 64–69.
Papaemmanuil E, Gerstung M, Malcovati L, Tauro S, Gundem G, Van Loo P et al. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood 2013; 122: 3616–3627, quiz 3699.
Malcovati L, Papaemmanuil E, Bowen DT, Boultwood J, Della Porta MG, Pascutto C et al. Clinical significance of SF3B1 mutations in myelodysplastic syndromes and myelodysplastic/myeloproliferative neoplasms. Blood 2011; 118: 6239–6246.
Thol F, Kade S, Schlarmann C, Loffeld P, Morgan M, Krauter J et al. Frequency and prognostic impact of mutations in SRSF2, U2AF1, and ZRSR2 in patients with myelodysplastic syndromes. Blood 2012; 119: 3578–3584.
Malcovati L, Karimi M, Papaemmanuil E, Ambaglio I, Jadersten M, Jansson M et al. SF3B1 mutation identifies a distinct subset of myelodysplastic syndrome with ring sideroblasts. Blood 2015; 126: 233–241.
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM et al. The 2016 revision to the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia. Blood 2016; 127: 2391–2405.
Cazzola M, Invernizzi R, Bergamaschi G, Levi S, Corsi B, Travaglino E et al. Mitochondrial ferritin expression in erythroid cells from patients with sideroblastic anemia. Blood 2003; 101: 1996–2000.
Visconte V, Rogers HJ, Singh J, Barnard J, Bupathi M, Traina F et al. SF3B1 haploinsufficiency leads to formation of ring sideroblasts in myelodysplastic syndromes. Blood 2012; 120: 3173–3186.
Boultwood J, Pellagatti A, Nikpour M, Pushkaran B, Fidler C, Cattan H et al. The role of the iron transporter ABCB7 in refractory anemia with ring sideroblasts. PLoS One 2008; 3: e1970.
Pondarre C, Campagna DR, Antiochos B, Sikorski L, Mulhern H, Fleming MD . Abcb7, the gene responsible for X-linked sideroblastic anemia with ataxia, is essential for hematopoiesis. Blood 2007; 109: 3567–3569.
Cavadini P, Biasiotto G, Poli M, Levi S, Verardi R, Zanella I et al. RNA silencing of the mitochondrial ABCB7 transporter in HeLa cells causes an iron-deficient phenotype with mitochondrial iron overload. Blood 2007; 109: 3552–3559.
Schellenberg MJ, Dul EL, MacMillan AM . Structural model of the p14/SF3b155. branch duplex complex. RNA 2011; 17: 155–165.
Gozani O, Potashkin J, Reed R . A potential role for U2AF-SAP 155 interactions in recruiting U2 snRNP to the branch site. Mol Cell Biol 1998; 18: 4752–4760.
Wang C, Chua K, Seghezzi W, Lees E, Gozani O, Reed R . Phosphorylation of spliceosomal protein SAP 155 coupled with splicing catalysis. Genes Dev 1998; 12: 1409–1414.
Lardelli RM, Thompson JX, Yates JR 3rd, Stevens SW . Release of SF3 from the intron branchpoint activates the first step of pre-mRNA splicing. RNA 2010; 16: 516–528.
Makishima H, Visconte V, Sakaguchi H, Jankowska AM, Abu Kar S, Jerez A et al. Mutations in the spliceosome machinery, a novel and ubiquitous pathway in leukemogenesis. Blood 2012; 119: 3203–3210.
Visconte V, Avishai N, Mahfouz R, Tabarroki A, Cowen J, Sharghi-Moshtaghin R et al. Distinct iron architecture in SF3B1-mutant myelodysplastic syndrome patients is linked to an SLC25A37 splice variant with a retained intron. Leukemia 2015; 29: 188–195.
Dolatshad H, Pellagatti A, Fernandez-Mercado M, Yip BH, Malcovati L, Attwood M et al. Disruption of SF3B1 results in deregulated expression and splicing of key genes and pathways in myelodysplastic syndrome hematopoietic stem and progenitor cells. Leukemia 2015; 29: 1092–1103.
Ferreira PG, Jares P, Rico D, Gomez-Lopez G, Martinez-Trillos A, Villamor N et al. Transcriptome characterization by RNA sequencing identifies a major molecular and clinical subdivision in chronic lymphocytic leukemia. Genome Res 2014; 24: 212–226.
Furney SJ, Pedersen M, Gentien D, Dumont AG, Rapinat A, Desjardins L et al. SF3B1 mutations are associated with alternative splicing in uveal melanoma. Cancer Discov 2013; 3: 1122–1129.
Maguire SL, Leonidou A, Wai P, Marchio C, Ng CK, Sapino A et al. SF3B1 mutations constitute a novel therapeutic target in breast cancer. J Pathol 2015; 235: 571–580.
Scott LM, Rebel VI . Acquired mutations that affect pre-mRNA splicing in hematologic malignancies and solid tumors. J Natl Cancer Inst 2013; 105: 1540–1549.
Biankin AV, Waddell N, Kassahn KS, Gingras MC, Muthuswamy LB, Johns AL et al. Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature 2012; 491: 399–405.
Abdel-Wahab O, Levine R . The spliceosome as an indicted conspirator in myeloid malignancies. Cancer Cell 2011; 20: 420–423.
Damm F, Kosmider O, Gelsi-Boyer V, Renneville A, Carbuccia N, Hidalgo-Curtis C et al. Mutations affecting mRNA splicing define distinct clinical phenotypes and correlate with patient outcome in myelodysplastic syndromes. Blood 2012; 119: 3211–3218.
DeBoever C, Ghia EM, Shepard PJ, Rassenti L, Barrett CL, Jepsen K et al. Transcriptome sequencing reveals potential mechanism of cryptic 3' splice site selection in SF3B1-mutated cancers. PLoS Comput Biol 2015; 11: e1004105.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 2013; 29: 15–21.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009; 25: 2078–2079.
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 2010; 28: 511–515.
Shen S, Park JW, Lu ZX, Lin L, Henry MD, Wu YN et al. rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc Natl Acad Sci USA 2014; 111: E5593–E5601.
Katz Y, Wang ET, Airoldi EM, Burge CB . Analysis and design of RNA sequencing experiments for identifying isoform regulation. Nat Methods 2010; 7: 1009–1015.
Caceres G, McGraw K, Yip BH, Pellagatti A, Johnson J, Zhang L et al. TP53 suppression promotes erythropoiesis in del(5q) MDS, suggesting a targeted therapeutic strategy in lenalidomide-resistant patients. Proc Natl Acad Sci USA 2013; 110: 16127–16132.
Pellagatti A, Jadersten M, Forsblom AM, Cattan H, Christensson B, Emanuelsson EK et al. Lenalidomide inhibits the malignant clone and up-regulates the SPARC gene mapping to the commonly deleted region in 5q- syndrome patients. Proc Natl Acad Sci USA 2007; 104: 11406–11411.
Hiller M, Huse K, Szafranski K, Jahn N, Hampe J, Schreiber S et al. Widespread occurrence of alternative splicing at NAGNAG acceptors contributes to proteome plasticity. Nat Genet 2004; 36: 1255–1257.
Smith CW, Chu TT, Nadal-Ginard B . Scanning and competition between AGs are involved in 3' splice site selection in mammalian introns. Mol Cell Biol 1993; 13: 4939–4952.
Corvelo A, Hallegger M, Smith CW, Eyras E . Genome-wide association between branch point properties and alternative splicing. PLoS Comput Biol 2010; 6: e1001016.
Darman RB, Seiler M, Agrawal AA, Lim KH, Peng S, Aird D et al. Cancer-associated SF3B1 hotspot mutations induce cryptic 3' splice site selection through use of a different branch point. Cell Rep 2015; 13: 1033–1045.
Alsafadi S, Houy A, Battistella A, Popova T, Wassef M, Henry E et al. Cancer-associated SF3B1 mutations affect alternative splicing by promoting alternative branchpoint usage. Nat Commun 2016; 7: 10615.
Lewis BP, Green RE, Brenner SE . Evidence for the widespread coupling of alternative splicing and nonsense-mediated mRNA decay in humans. Proc Natl Acad Sci USA 2003; 100: 189–192.
Ishigaki Y, Li X, Serin G, Maquat LE . Evidence for a pioneer round of mRNA translation: mRNAs subject to nonsense-mediated decay in mammalian cells are bound by CBP80 and CBP20. Cell 2001; 106: 607–617.
Nilsson L, Astrand-Grundstrom I, Anderson K, Arvidsson I, Hokland P, Bryder D et al. Involvement and functional impairment of the CD34(+)CD38(-)Thy-1(+) hematopoietic stem cell pool in myelodysplastic syndromes with trisomy 8. Blood 2002; 100: 259–267.
Woll PS, Kjallquist U, Chowdhury O, Doolittle H, Wedge DC, Thongjuea S et al. Myelodysplastic syndromes are propagated by rare and distinct human cancer stem cells in vivo. Cancer Cell 2014; 25: 794–808.
Chua K, Reed R . An upstream AG determines whether a downstream AG is selected during catalytic step II of splicing. Mol Cell Biol 2001; 21: 1509–1514.
Penalva LO, Lallena MJ, Valcarcel J . Switch in 3' splice site recognition between exon definition and splicing catalysis is important for sex-lethal autoregulation. Mol Cell Biol 2001; 21: 1986–1996.
Corrionero A, Minana B, Valcarcel J . Reduced fidelity of branch point recognition and alternative splicing induced by the anti-tumor drug spliceostatin A. Genes Dev 2011; 25: 445–459.
Lee SC, Dvinge H, Kim E, Cho H, Micol JB, Chung YR et al. Modulation of splicing catalysis for therapeutic targeting of leukemia with mutations in genes encoding spliceosomal proteins. Nat Med e-pub ahead of print 2 May 2016.
Yien YY, Robledo RF, Schultz IJ, Takahashi-Makise N, Gwynn B, Bauer DE et al. TMEM14C is required for erythroid mitochondrial heme metabolism. J Clin Invest 2014; 124: 4294–4304.
Hellstr+om-Lindberg E, Schmidt-Mende J, Forsblom AM, Christensson B, Fadeel B, Zhivotovsky B . Apoptosis in refractory anaemia with ringed sideroblasts is initiated at the stem cell level and associated with increased activation of caspases. Br J Haematol 2001; 112: 714–726.
Conte S, Katayama S, Vesterlund L, Karimi M, Dimitriou M, Jansson M et al. Aberrant splicing of genes involved in haemoglobin synthesis and impaired terminal erythroid maturation in SF3B1 mutated refractory anaemia with ring sideroblasts. Br J Haematol 2015; 171: 478–490.
Shimada Y, Okuno S, Kawai A, Shinomiya H, Saito A, Suzuki M et al. Cloning and chromosomal mapping of a novel ABC transporter gene (hABC7), a candidate for X-linked sideroblastic anemia with spinocerebellar ataxia. J Hum Genet 1998; 43: 115–122.
Kispal G, Csere P, Guiard B, Lill R . The ABC transporter Atm1p is required for mitochondrial iron homeostasis. FEBS Lett 1997; 418: 346–350.
Nikpour M, Scharenberg C, Liu A, Conte S, Karimi M, Mortera-Blanco T et al. The transporter ABCB7 is a mediator of the phenotype of acquired refractory anemia with ring sideroblasts. Leukemia 2013; 27: 889–896.
Gerstung M, Pellagatti A, Malcovati L, Giagounidis A, Porta MG, Jadersten M et al. Combining gene mutation with gene expression data improves outcome prediction in myelodysplastic syndromes. Nat Commun 2015; 6: 5901.
Acknowledgements
This study was supported by Bloodwise (UK). FGL and KIS were supported by a grant from the MRC (MR/K018965/1), and ML by a grant from the Wellcome Trust (092900).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Dolatshad, H., Pellagatti, A., Liberante, F. et al. Cryptic splicing events in the iron transporter ABCB7 and other key target genes in SF3B1-mutant myelodysplastic syndromes. Leukemia 30, 2322–2331 (2016). https://doi.org/10.1038/leu.2016.149
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2016.149
This article is cited by
-
Studying the connection between SF3B1 and four types of cancer by analyzing networks constructed based on published research
Scientific Reports (2023)
-
Mutation in SF3B1 gene promotes formation of polyploid giant cells in Leukemia cells
Medical Oncology (2022)
-
Complex landscape of alternative splicing in myeloid neoplasms
Leukemia (2021)
-
The biological function and clinical significance of SF3B1 mutations in cancer
Biomarker Research (2020)
-
Somatic SF3B1 hotspot mutation in prolactinomas
Nature Communications (2020)