Abstract
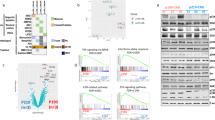
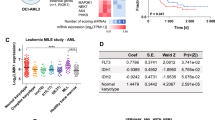
PTPN11 encodes the Shp2 non-receptor protein-tyrosine phosphatase implicated in several signaling pathways. Activating mutations in Shp2 are commonly associated with juvenile myelomonocytic leukemia but are not as well defined in other neoplasms. Here we report that Shp2 mutations occur in human acute myeloid leukemia (AML) at a rate of 6.6% (6/91) in the ECOG E1900 data set. We examined the role of mutated Shp2 in leukemias harboring MLL translocations, which co-occur in human AML. The hyperactive Shp2E76K mutant, commonly observed in leukemia patients, significantly accelerated MLL-AF9-mediated leukemogenesis in vivo. Shp2E76K increased leukemic stem cell frequency and affords MLL-AF9 leukemic cells IL3 cytokine hypersensitivity. As Shp2 is reported to regulate anti-apoptotic genes, we investigated Bcl2, Bcl-xL and Mcl1 expression in MLL-AF9 leukemic cells with and without Shp2E76K. Although the Bcl2 family of genes was upregulated in Shp2E76K cells, Mcl1 showed the highest upregulation in MLL-AF9 cells in response to Shp2E76K. Indeed, expression of Mcl1 in MLL-AF9 cells phenocopies expression of Shp2E76K, suggesting Shp2 mutations cooperate through activation of anti-apoptotic genes. Finally, we show Shp2E76K mutations reduce sensitivity of AML cells to small-molecule-mediated Mcl1 inhibition, suggesting reduced efficacy of drugs targeting MCL1 in patients with hyperactive Shp2.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Raimondi SC, Chang MN, Ravindranath Y, Behm FG, Gresik MV, Steuber CP et al. Chromosomal abnormalities in 478 children with acute myeloid leukemia: clinical characteristics and treatment outcome in a cooperative pediatric oncology group study-POG 8821. Blood 1999; 94: 3707–3716.
Rubnitz JE, Link MP, Shuster JJ, Carroll AJ, Hakami N, Frankel LS et al. Frequency and prognostic significance of HRX rearrangements in infant acute lymphoblastic leukemia: a Pediatric Oncology Group study. Blood 1994; 84: 570–573.
Huret JL, Minor SL, Dorkeld F, Dessen P, Bernheim A . Atlas of genetics and cytogenetics in oncology and haematology, an interactive database. Nucleic Acids Res 2000; 28: 349–351.
Bitoun E, Oliver PL, Davies KE . The mixed-lineage leukemia fusion partner AF4 stimulates RNA polymerase II transcriptional elongation and mediates coordinated chromatin remodeling. Hum Mol Genet 2007; 16: 92–106.
Lin C, Smith ER, Takahashi H, Lai KC, Martin-Brown S, Florens L et al. AFF4, a component of the ELL/P-TEFb elongation complex and a shared subunit of MLL chimeras, can link transcription elongation to leukemia. Mol Cell 2010; 37: 429–437.
Yokoyama A, Lin M, Naresh A, Kitabayashi I, Cleary ML . A higher-order complex containing AF4 and ENL family proteins with P-TEFb facilitates oncogenic and physiologic MLL-dependent transcription. Cancer Cell 2010; 17: 198–212.
Armstrong SA, Staunton JE, Silverman LB, Pieters R, den Boer ML, Minden MD et al. MLL translocations specify a distinct gene expression profile that distinguishes a unique leukemia. Nat Genet 2002; 30: 41–47.
Mullighan CG, Goorha S, Radtke I, Miller CB, Coustan-Smith E, Dalton JD et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 2007; 446: 758–764.
Balgobind BV, Zwaan CM, Pieters R, Van den Heuvel-Eibrink MM . The heterogeneity of pediatric MLL-rearranged acute myeloid leukemia. Leukemia 2011; 25: 1239–1248.
Gilliland DG, Griffin JD . The roles of FLT3 in hematopoiesis and leukemia. Blood 2002; 100: 1532–1542.
Ono R, Nakajima H, Ozaki K, Kumagai H, Kawashima T, Taki T et al. Dimerization of MLL fusion proteins and FLT3 activation synergize to induce multiple-lineage leukemogenesis. J Clin Invest 2005; 115: 919–929.
Stubbs MC, Kim YM, Krivtsov AV, Wright RD, Feng Z, Agarwal J et al. MLL-AF9 and FLT3 cooperation in acute myelogenous leukemia: development of a model for rapid therapeutic assessment. Leukemia 2008; 22: 66–77.
Kim WI, Matise I, Diers MD, Largaespada DA .. RAS oncogene suppression induces apoptosis followed by more differentiated and less myelosuppressive disease upon relapse of acute myeloid leukemia. Blood 2009; 113: 1086–1096.
Mohi MG, Neel BG . The role of Shp2 (PTPN11) in cancer. Curr Opin Genet Dev 2007; 17: 23–30.
Chan G, Kalaitzidis D, Neel BG . The tyrosine phosphatase Shp2 (PTPN11) in cancer. Cancer Meta Rev 2008; 27: 179–192.
Tartaglia M, Mehler EL, Goldberg R, Zampino G, Brunner HG, Kremer H et al. Mutations in PTPN11, encoding the protein tyrosine phosphatase SHP-2, cause Noonan syndrome. Nat Genet 2001; 29: 465–468.
Tartaglia M, Niemeyer CM, Fragale A, Song X, Buechner J, Jung A et al. Somatic mutations in PTPN11 in juvenile myelomonocytic leukemia, myelodysplastic syndromes and acute myeloid leukemia. Nat Genet 2003; 34: 148–150.
Paulsson K, Horvat A, Strombeck B, Nilsson F, Heldrup J, Behrendtz M et al. Mutations of FLT3, NRAS, KRAS, and PTPN11 are frequent and possibly mutually exclusive in high hyperdiploid childhood acute lymphoblastic leukemia. Genes Chromosomes Cancer 2008; 47: 26–33.
Tartaglia M, Martinelli S, Cazzaniga G, Cordeddu V, Iavarone I, Spinelli M et al. Genetic evidence for lineage-related and differentiation stage-related contribution of somatic PTPN11 mutations to leukemogenesis in childhood acute leukemia. Blood 2004; 104: 307–313.
Lawrence MS, Stojanov P, Mermel CH, Robinson JT, Garraway LA, Golub TR et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 2014; 505: 495–501.
Welch JS, Ley TJ, Link DC, Miller CA, Larson DE, Koboldt DC et al. The origin and evolution of mutations in acute myeloid leukemia. Cell 2012; 150: 264–278.
Neel BG, Gu H, Pao L . The 'Shp'ing news: SH2 domain-containing tyrosine phosphatases in cell signaling. Trends Biochem Sci 2003; 28: 284–293.
Zhu HH, Ji K, Alderson N, He Z, Li S, Liu W et al. Kit-Shp2-Kit signaling acts to maintain a functional hematopoietic stem and progenitor cell pool. Blood 2011; 117: 5350–5361.
Li L, Modi H, McDonald T, Rossi J, Yee JK, Bhatia R .. A critical role for SHP2 in STAT5 activation and growth factor-mediated proliferation, survival, and differentiation of human CD34+ cells. Blood 2011; 118: 1504–1515.
Chan G, Cheung LS, Yang W, Milyavsky M, Sanders AD, Gu S et al. Essential role for Ptpn11 in survival of hematopoietic stem and progenitor cells. Blood 2011; 117: 4253–4261.
Xu R, Yu Y, Zheng S, Zhao X, Dong Q, He Z et al. Overexpression of Shp2 tyrosine phosphatase is implicated in leukemogenesis in adult human leukemia. Blood 2005; 106: 3142–3149.
Nabinger SC, Li XJ, Ramdas B, He Y, Zhang X, Zeng L et al. The protein tyrosine phosphatase, Shp2, positively contributes to FLT3-ITD-induced hematopoietic progenitor hyperproliferation and malignant disease in vivo. Leukemia 2013; 27: 398–408.
Hof P, Pluskey S, Dhe-Paganon S, Eck MJ, Shoelson SE . Crystal structure of the tyrosine phosphatase SHP-2. Cell 1998; 92: 441–450.
Stein-Gerlach M, Wallasch C, SHP-2 Ullrich A .. SH2-containing protein tyrosine phosphatase-2. Int J Biochem Cell Biol 1998; 30: 559–566.
Barford D, Neel BG . Revealing mechanisms for SH2 domain mediated regulation of the protein tyrosine phosphatase SHP-2. Structure 1998; 6: 249–254.
Schubbert S, Lieuw K, Rowe SL, Lee CM, Li X, Loh ML et al. Functional analysis of leukemia-associated PTPN11 mutations in primary hematopoietic cells. Blood 2005; 106: 311–317.
Xu D, Liu X, Yu WM, Meyerson HJ, Guo C, Gerson SL et al. Non-lineage/stage-restricted effects of a gain-of-function mutation in tyrosine phosphatase Ptpn11 (Shp2) on malignant transformation of hematopoietic cells. J Exp Med 2011; 208: 1977–1988.
Chan G, Kalaitzidis D, Usenko T, Kutok JL, Yang W, Mohi MG et al. Leukemogenic Ptpn11 causes fatal myeloproliferative disorder via cell-autonomous effects on multiple stages of hematopoiesis. Blood 2009; 113: 4414–4424.
Mohi MG, Williams IR, Dearolf CR, Chan G, Kutok JL, Cohen S et al. Prognostic, therapeutic, and mechanistic implications of a mouse model of leukemia evoked by Shp2 (PTPN11) mutations. Cancer Cell 2005; 7: 179–191.
Xu D, Wang S, Yu WM, Chan G, Araki T, Bunting KD et al. A germline gain-of-function mutation in Ptpn11 (Shp-2) phosphatase induces myeloproliferative disease by aberrant activation of hematopoietic stem cells. Blood 2010; 116: 3611–3621.
Yu ZH, Xu J, Walls CD, Chen L, Zhang S, Zhang R et al. Structural and mechanistic insights into LEOPARD syndrome-associated SHP2 mutations. J Biol Chem 2013; 288: 10472–10482.
Patel JP, Gonen M, Figueroa ME, Fernandez H, Sun Z, Racevskis J et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N Engl J Med 2012; 366: 1079–1089.
ElSharawy A, Warner J, Olson J, Forster M, Schilhabel MB, Link DR et al. Accurate variant detection across non-amplified and whole genome amplified DNA using targeted next generation sequencing. BMC Genomics 2012; 13: 500.
Tewhey R, Warner JB, Nakano M, Libby B, Medkova M, David PH et al. Microdroplet-based PCR enrichment for large-scale targeted sequencing. Nat Biotechnol 2009; 27: 1025–1031.
Muntean AG, Chen W, Jones M, Granowicz EM, Maillard I, Hess JL . MLL fusion protein-driven AML is selectively inhibited by targeted disruption of the MLL-PAFc interaction. Blood 2013, 30.
Chan RJ, Leedy MB, Munugalavadla V, Voorhorst CS, Li Y, Yu M et al. Human somatic PTPN11 mutations induce hematopoietic-cell hypersensitivity to granulocyte-macrophage colony-stimulating factor. Blood 2005; 105: 3737–3742.
Zuber J, McJunkin K, Fellmann C, Dow LE, Taylor MJ, Hannon GJ et al. Toolkit for evaluating genes required for proliferation and survival using tetracycline-regulated RNAi. Nat Biotechnol 2011; 29: 79–83.
Glaser SP, Lee EF, Trounson E, Bouillet P, Wei A, Fairlie WD et al. Anti-apoptotic Mcl-1 is essential for the development and sustained growth of acute myeloid leukemia. Genes Dev 2012; 26: 120–125.
Muntean AG, Giannola D, Udager AM, Hess JL . The PHD fingers of MLL block MLL fusion protein-mediated transformation. Blood 2008; 112: 4690–4693.
Tan J, Jones M, Koseki H, Nakayama M, Muntean AG, Maillard I et al. CBX8, a polycomb group protein, is essential for MLL-AF9-induced leukemogenesis. Cancer Cell 2011; 20: 563–575.
Hu Y, Smyth GK . ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J Immunol Methods 2009; 347: 70–78.
Fernandez HF, Sun Z, Yao X, Litzow MR, Luger SM, Paietta EM et al. Anthracycline dose intensification in acute myeloid leukemia. N Engl J Med 2009; 361: 1249–1259.
Inaba T, Inukai T, Yoshihara T, Seyschab H, Ashmun RA, Canman CE et al. Reversal of apoptosis by the leukaemia-associated E2A-HLF chimaeric transcription factor. Nature 1996; 382: 541–544.
Huang H, Woo AJ, Waldon Z, Schindler Y, Moran TB, Zhu HH et al. A Src family kinase-Shp2 axis controls RUNX1 activity in megakaryocyte and T-lymphocyte differentiation. Genes Dev 2012; 26: 1587–1601.
Tse C, Shoemaker AR, Adickes J, Anderson MG, Chen J, Jin S et al. ABT-263: a potent and orally bioavailable Bcl-2 family inhibitor. Cancer Res 2008; 68: 3421–3428.
Abulwerdi F, Liao C, Liu M, Azmi AS, Aboukameel A, Mady AS et al. A novel small-molecule inhibitor of mcl-1 blocks pancreatic cancer growth in vitro and in vivo. Mol Cancer Ther 2014; 13: 565–575.
Abulwerdi FA, Liao C, Mady AS, Gavin J, Shen C, Cierpicki T et al. 3-substituted-N-(4-hydroxynaphthalen-1-yl)arylsulfonamides as a novel class of selective Mcl-1 inhibitors: structure-based design, synthesis, SAR, and biological evaluation. J Med Chem 2014; 57: 4111–4133.
Dobson CL, Warren AJ, Pannell R, Forster A, Lavenir I, Corral J et al. The mll-AF9 gene fusion in mice controls myeloproliferation and specifies acute myeloid leukaemogenesis. EMBO J 1999; 18: 3564–3574.
Lavau C, Szilvassy SJ, Slany R, Cleary ML . Immortalization and leukemic transformation of a myelomonocytic precursor by retrovirally transduced HRX-ENL. EMBO J 1997; 16: 4226–4237.
Zhou P, Qian L, Bieszczad CK, Noelle R, Binder M, Levy NB et al. Mcl-1 in transgenic mice promotes survival in a spectrum of hematopoietic cell types and immortalization in the myeloid lineage. Blood 1998; 92: 3226–3239.
Okamoto T, Coultas L, Metcalf D, van Delft MF, Glaser SP, Takiguchi M et al. Enhanced stability of Mcl1, a prosurvival Bcl2 relative, blunts stress-induced apoptosis, causes male sterility, and promotes tumorigenesis. Proc Natl Acad Sci USA 2014; 111: 261–266.
Campbell CJ, Lee JB, Levadoux-Martin M, Wynder T, Xenocostas A, Leber B et al. The human stem cell hierarchy is defined by a functional dependence on Mcl-1 for self-renewal capacity. Blood 2010; 116: 1433–1442.
Acknowledgements
We thank Dr Jay Hess for helpful discussion. This work was supported by NIH grants R01-CA149442 (ZN-C), R00 CA158136 (AGM) and an American Society of Hematology Scholar Award (AGM)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Chen, L., Chen, W., Mysliwski, M. et al. Mutated Ptpn11 alters leukemic stem cell frequency and reduces the sensitivity of acute myeloid leukemia cells to Mcl1 inhibition. Leukemia 29, 1290–1300 (2015). https://doi.org/10.1038/leu.2015.18
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2015.18
This article is cited by
-
Divergent leukaemia subclones as cellular models for testing vulnerabilities associated with gains in chromosomes 7, 8 or 18
Scientific Reports (2021)
-
The Clinical impact of PTPN11 mutations in adults with acute myeloid leukemia
Leukemia (2021)
-
The role of EVI1 gene quantification in AML patients with 11q23/MLL rearrangement after allogeneic hematopoietic stem cell transplantation
Bone Marrow Transplantation (2021)
-
Integrated analysis of patient samples identifies biomarkers for venetoclax efficacy and combination strategies in acute myeloid leukemia
Nature Cancer (2020)
-
Necroinflammation emerges as a key regulator of hematopoiesis in health and disease
Cell Death & Differentiation (2019)