Abstract
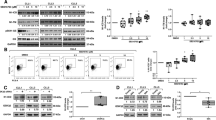
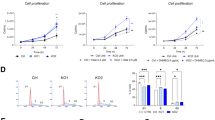
In chronic lymphocytic leukemia (CLL), NOTCH1 mutations have been associated with clinical resistance to the anti-CD20 rituximab, although the mechanisms behind this peculiar behavior remain to be clarified. In a wide CLL series (n=692), we demonstrated that CLL cells from NOTCH1-mutated cases (87/692) were characterized by lower CD20 expression and lower relative lysis induced by anti-CD20 exposure in vitro. Consistently, CD20 expression by CLL cells was upregulated in vitro by γ-secretase inhibitors or NOTCH1-specific small interfering RNA and the stable transfection of a mutated (c.7541-7542delCT) NOTCH1 intracellular domain (NICD-mut) into CLL-like cells resulted in a strong downregulation of both CD20 protein and transcript. By using these NICD-mut transfectants, we investigated protein interactions of RBPJ, a transcription factor acting either as activator or repressor of NOTCH1 pathway when respectively bound to NICD or histone deacetylases (HDACs). Compared with controls, NICD-mut transfectants had RBPJ preferentially complexed to NICD and showed higher levels of HDACs interacting with the promoter of the CD20 gene. Finally, treatment with the HDAC inhibitor valproic acid upregulated CD20 in both NICD-mut transfectants and primary CLL cells. In conclusion, NOTCH1 mutations are associated with low CD20 levels in CLL and are responsible for a dysregulation of HDAC-mediated epigenetic repression of CD20 expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hallek M, Cheson BD, Catovsky D, Caligaris-Cappio F, Dighiero G, Dohner H et al. Guidelines for the diagnosis and treatment of chronic lymphocytic leukemia: a report from the International Workshop on Chronic Lymphocytic Leukemia updating the National Cancer Institute-Working Group 1996 guidelines. Blood 2008; 111: 5446–5456.
Hallek M . Chronic lymphocytic leukemia: 2013 update on diagnosis, risk stratification and treatment. Am J Hematol 2013; 88: 803–816.
Fabbri G, Rasi S, Rossi D, Trifonov V, Khiabanian H, Ma J et al. Analysis of the chronic lymphocytic leukemia coding genome: role of NOTCH1 mutational activation. J Exp Med 2011; 208: 1389–1401.
Puente XS, Pinyol M, Quesada V, Conde L, Ordonez GR, Villamor N et al. Whole-genome sequencing identifies recurrent mutations in chronic lymphocytic leukaemia. Nature 2011; 475: 101–105.
Rossi D, Rasi S, Spina V, Bruscaggin A, Monti S, Ciardullo C et al. Integrated mutational and cytogenetic analysis identifies new prognostic subgroups in chronic lymphocytic leukemia. Blood 2013; 121: 1403–1412.
Quesada V, Conde L, Villamor N, Ordonez GR, Jares P, Bassaganyas L et al. Exome sequencing identifies recurrent mutations of the splicing factor SF3B1 gene in chronic lymphocytic leukemia. Nat Genet 2012; 44: 47–52.
Wang L, Lawrence MS, Wan Y, Stojanov P, Sougnez C, Stevenson K et al. SF3B1 and other novel cancer genes in chronic lymphocytic leukemia. N Engl J Med 2011; 365: 2497–2506.
Del Poeta G, Dal Bo M, Del Principe MI, Pozzo F, Rossi FM, Zucchetto A et al. Clinical significance of c.7544-7545 delCT NOTCH1 mutation in chronic lymphocytic leukaemia. Br J Haematol 2013; 160: 415–418.
Del Giudice I, Rossi D, Chiaretti S, Marinelli M, Tavolaro S, Gabrielli S et al. NOTCH1 mutations in +12 chronic lymphocytic leukemia (CLL) confer an unfavorable prognosis, induce a distinctive transcriptional profiling and refine the intermediate prognosis of +12 CLL. Haematologica 2012; 97: 437–441.
Rossi D, Rasi S, Fabbri G, Spina V, Fangazio M, Forconi F et al. Mutations of NOTCH1 are an independent predictor of survival in chronic lymphocytic leukemia. Blood 2012; 119: 521–529.
Lobry C, Oh P, Aifantis I . Oncogenic and tumor suppressor functions of Notch in cancer: it’s NOTCH what you think. J Exp Med 2011; 208: 1931–1935.
Yuan JS, Kousis PC, Suliman S, Visan I, Guidos CJ . Functions of notch signaling in the immune system: consensus and controversies. Annu Rev Immunol 2010; 28: 343–365.
Castel D, Mourikis P, Bartels SJ, Brinkman AB, Tajbakhsh S, Stunnenberg HG . Dynamic binding of RBPJ is determined by Notch signaling status. Genes Dev 2013; 27: 1059–1071.
Davis RL, Turner DL . Vertebrate hairy and enhancer of split related proteins: transcriptional repressors regulating cellular differentiation and embryonic patterning. Oncogene 2001; 20: 8342–8357.
Hsieh JJ, Hayward SD . Masking of the CBF1/RBPJ kappa transcriptional repression domain by Epstein-Barr virus EBNA2. Science 1995; 268: 560–563.
Iso T, Kedes L, Hamamori Y . HES and HERP families: multiple effectors of the Notch signaling pathway. J Cell Physiol 2003; 194: 237–255.
Jarriault S, Brou C, Logeat F, Schroeter EH, Kopan R, Israel A . Signalling downstream of activated mammalian Notch. Nature 1995; 377: 355–358.
Kato H, Taniguchi Y, Kurooka H, Minoguchi S, Sakai T, Nomura-Okazaki S et al. Involvement of RBP-J in biological functions of mouse Notch1 and its derivatives. Development 1997; 124: 4133–4141.
Klinakis A, Szabolcs M, Politi K, Kiaris H, Artavanis-Tsakonas S, Efstratiadis A . Myc is a Notch1 transcriptional target and a requisite for Notch1-induced mammary tumorigenesis in mice. Proc Natl Acad Sci USA 2006; 103: 9262–9267.
Weng AP, Millholland JM, Yashiro-Ohtani Y, Arcangeli ML, Lau A, Wai C et al. c-Myc is an important direct target of Notch1 in T-cell acute lymphoblastic leukemia/lymphoma. Genes Dev 2006; 20: 2096–2109.
Rosati E, Sabatini R, Rampino G, Tabilio A, Di IM, Fettucciari K et al. Constitutively activated Notch signaling is involved in survival and apoptosis resistance of B-CLL cells. Blood 2009; 113: 856–865.
Paganin M, Ferrando A . Molecular pathogenesis and targeted therapies for NOTCH1-induced T-cell acute lymphoblastic leukemia. Blood Rev 2011; 25: 83–90.
Weng AP, Ferrando AA, Lee W, Morris JP, Silverman LB, Sanchez-Irizarry C et al. Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 2004; 306: 269–271.
Sportoletti P, Baldoni S, Cavalli L, Del PB, Bonifacio E, Ciurnelli R et al. NOTCH1 PEST domain mutation is an adverse prognostic factor in B-CLL. Br J Haematol 2010; 151: 404–406.
Arruga F, Gizdic B, Serra S, Vaisitti T, Ciardullo C, Coscia M et al. Functional impact of NOTCH1 mutations in chronic lymphocytic leukemia. Leukemia 2014; 28: 1060–1070.
Stilgenbauer S, Schnaiter A, Paschka P, Zenz T, Rossi M, Dohner K et al. Gene mutations and treatment outcome in chronic lymphocytic leukemia: results from the CLL8 trial. Blood 2014; 123: 3247–3254.
Matutes E, Owusu-Ankomah K, Morilla R, Garcia MJ, Houlihan A, Que TH et al. The immunological profile of B-cell disorders and proposal of a scoring system for the diagnosis of CLL. Leukemia 1994; 8: 1640–1645.
Gattei V, Bulian P, Del Principe MI, Zucchetto A, Maurillo L, Buccisano F et al. Relevance of CD49d protein expression as overall survival and progressive disease prognosticator in chronic lymphocytic leukemia. Blood 2008; 111: 865–873.
Zucchetto A, Vaisitti T, Benedetti D, Tissino E, Bertagnolo V, Rossi D et al. The CD49d/CD29 complex is physically and functionally associated with CD38 in B-cell chronic lymphocytic leukemia cells. Leukemia 2012; 26: 1301–1312.
Rossi D, Spina V, Bomben R, Rasi S, Dal-Bo M, Bruscaggin A et al. Association between molecular lesions and specific B-cell receptor subsets in chronic lymphocytic leukemia. Blood 2013; 121: 4902–4905.
Balatti V, Bottoni A, Palamarchuk A, Alder H, Rassenti LZ, Kipps TJ et al. NOTCH1 mutations in CLL associated with trisomy 12. Blood 2012; 119: 329–331.
Bomben R, Gobessi S, Dal BM, Volinia S, Marconi D, Tissino E et al. The miR-17 approximately 92 family regulates the response to Toll-like receptor 9 triggering of CLL cells with unmutated IGHV genes. Leukemia 2012; 26: 1584–1593.
Saborit-Villarroya I, Vaisitti T, Rossi D, D'Arena G, Gaidano G, Malavasi F et al. E2A is a transcriptional regulator of CD38 expression in chronic lymphocytic leukemia. Leukemia 2011; 25: 479–488.
Sugimoto T, Tomita A, Hiraga J, Shimada K, Kiyoi H, Kinoshita T et al. Escape mechanisms from antibody therapy to lymphoma cells: downregulation of CD20 mRNA by recruitment of the HDAC complex and not by DNA methylation. Biochem Biophys Res Commun 2009; 390: 48–53.
Bray SJ . Notch signalling: a simple pathway becomes complex. Nat Rev Mol Cell Biol 2006; 7: 678–689.
Dohner H, Stilgenbauer S, Benner A, Leupolt E, Krober A, Bullinger L et al. Genomic aberrations and survival in chronic lymphocytic leukemia. N Engl J Med 2000; 343: 1910–1916.
Tam CS, Otero-Palacios J, Abruzzo LV, Jorgensen JL, Ferrajoli A, Wierda WG et al. Chronic lymphocytic leukaemia CD20 expression is dependent on the genetic subtype: a study of quantitative flow cytometry and fluorescent in-situ hybridization in 510 patients. Br J Haematol 2008; 141: 36–40.
Tedder TF, Streuli M, Schlossman SF, Saito H . Isolation and structure of a cDNA encoding the B1 (CD20) cell-surface antigen of human B lymphocytes. Proc Natl Acad Sci USA 1988; 85: 208–212.
Hiraga J, Tomita A, Sugimoto T, Shimada K, Ito M, Nakamura S et al. Down-regulation of CD20 expression in B-cell lymphoma cells after treatment with rituximab-containing combination chemotherapies: its prevalence and clinical significance. Blood 2009; 113: 4885–4893.
Shimizu R, Kikuchi J, Wada T, Ozawa K, Kano Y, Furukawa Y . HDAC inhibitors augment cytotoxic activity of rituximab by upregulating CD20 expression on lymphoma cells. Leukemia 2010; 24: 1760–1768.
Bo MD, Del Principe MI, Pozzo F, Ragusa D, Bulian P, Rossi D et al. NOTCH1 mutations identify a chronic lymphocytic leukemia patient subset with worse prognosis in the setting of a rituximab-based induction and consolidation treatment. Ann Hematol 2014; 93: 1765–1774.
Golay J, Lazzari M, Facchinetti V, Bernasconi S, Borleri G, Barbui T et al. CD20 levels determine the in vitro susceptibility to rituximab and complement of B-cell chronic lymphocytic leukemia: further regulation by CD55 and CD59. Blood 2001; 98: 3383–3389.
Sportoletti P, Baldoni S, Del PB, Aureli P, Dorillo E, Ruggeri L et al. A revised NOTCH1 mutation frequency still impacts survival while the allele burden predicts early progression in chronic lymphocytic leukemia. Leukemia 2014; 28: 436–439.
Andersson ER, Lendahl U . Therapeutic modulation of Notch signalling—are we there yet? Nat Rev Drug Discov 2014; 13: 357–378.
Minucci S, Pelicci PG . Histone deacetylase inhibitors and the promise of epigenetic (and more) treatments for cancer. Nat Rev Cancer 2006; 6: 38–51.
Ropero S, Esteller M . The role of histone deacetylases (HDACs) in human cancer. Mol Oncol 2007; 1: 19–25.
Acknowledgements
This study was supported in part by the Associazione Italiana Ricerca Cancro (AIRC), Investigator Grant IG-13227; Progetto Ricerca Finalizzata I.R.C.C.S. n. RF-2009-1469205, n. RF-2010-2307262, Progetto Giovani Ricercatori n. GR-2009-1475467, n. GR-2010-2317594, n. GR-2011-02347441, n. GR-2011-02346826, Ministero della Salute, Rome, Italy; Fondazione Cariplo (grant 2012-0689); Associazione Italiana contro le Leucemie, linfomi e mielomi (AIL), Venezia Section, Pramaggiore Group, Italy; Fondazione per la Vita di Pordenone, Italy; Ricerca Scientifica Applicata, Regione Friuli Venezia Giulia (‘Linfonet’ Project), Trieste, Italy; and ‘5x1000 Intramural Program’, Centro di Riferimento Oncologico, Aviano, Italy. FA is supported by a Beat-Leukemia fellowship.
Author Contributions
FP contributed to write the manuscript, analyzed the data and performed the research. TB performed the research. FA, PB, PM, ET, BG, FMR, RB, AZ, DB and MD contributed to perform the research. GDA, AC, FZ, GP, DR, GG., GDP and SD provided well-characterized biological samples and contributed to write the manuscript. VG and MDB designed the study, interpreted data and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Pozzo, F., Bittolo, T., Arruga, F. et al. NOTCH1 mutations associate with low CD20 level in chronic lymphocytic leukemia: evidence for a NOTCH1 mutation-driven epigenetic dysregulation. Leukemia 30, 182–189 (2016). https://doi.org/10.1038/leu.2015.182
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2015.182
This article is cited by
-
High-resolution Melting Analysis for NOTCH1 c.7541-7542delCT Mutation in Chronic Lymphocytic Leukemia: Prognostic Significance in Egyptian Patients
Indian Journal of Hematology and Blood Transfusion (2022)
-
Recent progress of prognostic biomarkers and risk scoring systems in chronic lymphocytic leukemia
Biomarker Research (2020)
-
Diminished interaction between mutant NOTCH1 and the NuRD corepressor complex upregulates CCL17 in chronic lymphocytic leukemia
Leukemia (2019)
-
Chronic lymphocytic leukaemia: from genetics to treatment
Nature Reviews Clinical Oncology (2019)
-
Advances in Epigenetics and Epigenomics in Chronic Lymphocytic Leukemia
Current Genetic Medicine Reports (2019)