Abstract
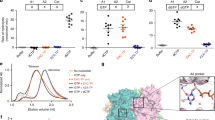
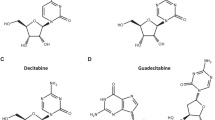
The three DNA methyltransferase (DNMT)-inhibiting cytosine nucleoside analogues, azacitidine, decitabine and zebularine, which are currently studied as nonintensive therapy for myelodysplastic syndromes and acute myeloid leukemia (AML), differ in structure and metabolism, suggesting that they may have differential molecular activity. We investigated cellular and molecular effects of the three substances relative to cytarabine in Kasumi-1 AML blasts. Under in vitro conditions mimicking those used in clinical trials, the DNMT inhibitors inhibited proliferation and triggered apoptosis but did not induce myeloid differentiation. The DNMT inhibitors showed no interference with cell-cycle progression whereas cytarabine treatment resulted in an S-phase arrest. Quantitative methylation analysis of hypermethylated gene promoters and of genome-wide LINE1 fragments using bisulfite sequencing and MassARRAY suggested that the hypomethylating potency of decitabine was stronger than that of azacitidine; zebularine showed no hypomethylating activity. In a comparative gene expression analysis, we found that the effects of each DNMT inhibitor on gene transcription were surprisingly different, involving several genes relevant to leukemogenesis. In addition, the gene methylation and expression analyses suggested that the effects of DNMT-inhibiting cytosine nucleoside analogues on the cellular transcriptome may, in part, be unrelated to direct promoter DNA hypomethylation, as previously shown by others.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lübbert M, Minden M . Decitabine in acute myeloid leukemia. Semin Hematol 2005; 42: S38–S42.
Silverman LR, McKenzie DR, Peterson BL, Holland JF, Backstrom JT, Beach CL et al. Further analysis of trials with azacitidine in patients with myelodysplastic syndrome: studies 8421, 8921, and 9221 by the Cancer and Leukemia Group B. J Clin Oncol 2006; 24: 3895–3903.
Scott SA, Lakshimikuttysamma A, Sheridan DP, Sanche SE, Geyer CR, DeCoteau JF . Zebularine inhibits human acute myeloid leukemia cell growth in vitro in association with p15INK4B demethylation and reexpression. Exp Hematol 2007; 35: 263–273.
Kuendgen A, Lübbert M . Current status of epigenetic treatment in myelodysplastic syndromes. Ann Hematol 2008; 87: 601–611.
Hurd PJ, Whitmarsh AJ, Baldwin GS, Kelly SM, Waltho JP, Price NC et al. Mechanism-based inhibition of C5-cytosine DNA methyltransferases by 2-H pyrimidinone. J Mol Biol 1999; 286: 389–401.
Jones PA, Baylin SB . The fundamental role of epigenetic events in cancer. Nat Rev Genet 2002; 3: 415–428.
Yoo CB, Jones PA . Epigenetic therapy of cancer: past, present and future. Nat Rev Drug Discov 2006; 5: 37–50.
Berg T, Guo Y, Abdelkarim M, Fliegauf M, Lübbert M . Differential p15/INK4B reactivation and growth inhibition induced by 5-aza-2′-deoxycytidine in AML1/ETO-positive and negative myeloid cells. Leuk Res 2007; 31: 497–506.
Asou H, Tashiro S, Hamamoto K, Otsuji A, Kita K, Kamada N . Establishment of a human acute myeloid leukemia cell line (Kasumi-1) with 8;21 chromosome translocation. Blood 1991; 77: 2031–2036.
Kozu T, Miyoshi H, Shimizu K, Maseki N, Kaneko Y, Asou H et al. Junctions of the AML1/MTG8(ETO) fusion are constant in t(8;21) acute myeloid leukemia detected by reverse transcription polymerase chain reaction. Blood 1993; 82: 1270–1276.
Li C, Hung WW . Model-based analysis of oligonucleotide arrays: model validation, design issues and standard error application. Genome Biol 2001; 2: RESEARCH0032.
Li C, Wong WH . Model-based analysis of oligonucleotide arrays: expression index computation and outlier detection. Proc Natl Acad Sci USA 2001; 98: 31–36.
Nguyen C, Liang G, Nguyen TT, Tsao-Wei D, Groshen S, Lübbert M et al. Susceptibility of nonpromoter CpG islands to de novo methylation in normal and neoplastic cells. J Natl Cancer Inst 2001; 93: 1465–1472.
Ehrich M, Nelson MR, Stanssens P, Zabeau M, Liloglou T, Xinarianos G et al. Quantitative high-throughput analysis of DNA methylation patterns by base-specific cleavage and mass spectrometry. Proc Natl Acad Sci USA 2005; 102: 15785–15790.
Momparler RL, Samson J, Momparler LF, Rivard GE . Cell cycle effects and cellular pharmacology of 5-aza-2′-deoxycytidine. Cancer Chemother Pharmacol 1984; 13: 191–194.
Momparler RL . Epigenetic therapy of cancer with 5-aza-2′-deoxycytidine (decitabine). Semin Oncol 2005; 32: 443–451.
Sampath D, Cortes J, Estrov Z, Du M, Shi Z, Andreeff M et al. Pharmacodynamics of cytarabine alone and in combination with 7-hydroxystaurosporine (UCN-01) in AML blasts in vitro and during a clinical trial. Blood 2006; 107: 2517–2524.
Raj K, John A, Ho A, Chronis C, Khan S, Samuel J et al. CDKN2B methylation status and isolated chromosome 7 abnormalities predict responses to treatment with 5-azacytidine. Leukemia 2007; 21: 1937–1944.
Hackanson B, Bennett KL, Brena RM, Jiang J, Claus R, Chen SS et al. Epigenetic modification of CCAAT/enhancer binding protein alpha expression in acute myeloid leukemia. Cancer Res 2008; 68: 3142–3151.
Lemaire M, Momparler LF, Bernstein ML, Marquez VE, Momparler RL . Enhancement of antineoplastic action of 5-aza-2′-deoxycytidine by zebularine on L1210 leukemia. Anticancer Drugs 2005; 16: 301–308.
Sato N, Matsubayashi H, Abe T, Fukushima N, Goggins M . Epigenetic down-regulation of CDKN1C/p57KIP2 in pancreatic ductal neoplasms identified by gene expression profiling. Clin Cancer Res 2005; 11: 4681–4688.
Nakamura T, Largaespada DA, Lee MP, Johnson LA, Ohyashiki K, Toyama K et al. Fusion of the nucleoporin gene NUP98 to HOXA9 by the chromosome translocation t(7;11)(p15;p15) in human myeloid leukaemia. Nat Genet 1996; 12: 154–158.
Armstrong SA, Staunton JE, Silverman LB, Pieters R, den Boer ML, Minden MD et al. MLL translocations specify a distinct gene expression profile that distinguishes a unique leukemia. Nat Genet 2002; 30: 41–47.
Lasa A, Carnicer MJ, Aventin A, Estivill C, Brunet S, Sierra J et al. MEIS 1 expression is downregulated through promoter hypermethylation in AML1-ETO acute myeloid leukemias. Leukemia 2004; 18: 1231–1237.
Accordi B, Pillozzi S, Dell’Orto MC, Cazzaniga G, Arcangeli A, Kronnie GT et al. Hepatocyte growth factor receptor c-MET is associated with FAS and when activated enhances drug-induced apoptosis in pediatric B acute lymphoblastic leukemia with TEL-AML1 translocation. J Biol Chem 2007; 282: 29384–29393.
Nigten J, Breems-de Ridder MC, Erpelinck-Verschueren CA, Nikoloski G, van der Reijden BA, van Wageningen S et al. ID1 and ID2 are retinoic acid responsive genes and induce a G0/G1 accumulation in acute promyelocytic leukemia cells. Leukemia 2005; 19: 799–805.
Sakajiri S, O’kelly J, Yin D, Miller CW, Hofmann WK, Oshimi K et al. Dlk1 in normal and abnormal hematopoiesis. Leukemia 2005; 19: 1404–1410.
Chen K, Tu Y, Zhang Y, Blair HC, Zhang L, Wu C . PINCH-1 regulates the ERK–Bim pathway and contributes to apoptosis resistance in cancer cells. J Biol Chem 2008; 283: 2508–2517.
Suzuki H, Gabrielson E, Chen W, Anbazhagan R, van Engeland M, Weijenberg MP et al. A genomic screen for genes upregulated by demethylation and histone deacetylase inhibition in human colorectal cancer. Nat Genet 2002; 31: 141–149.
Schmelz K, Sattler N, Wagner M, Lübbert M, Dörken B, Tamm I . Induction of gene expression by 5-Aza-2′-deoxycytidine in acute myeloid leukemia (AML) and myelodysplastic syndrome (MDS) but not epithelial cells by DNA-methylation-dependent and -independent mechanisms. Leukemia 2005; 19: 103–111.
Lee T, Karon M, Momparler RL . Kinetic studies on phosphorylation of 5-azacytidine with the purified uridine-cytidine kinase from calf thymus. Cancer Res 1974; 34: 2482–2488.
Ben Kasus T, Ben Zvi Z, Marquez VE, Kelley JA, Agbaria R . Metabolic activation of zebularine, a novel DNA methylation inhibitor, in human bladder carcinoma cells. Biochem Pharmacol 2005; 70: 121–133.
Cheng JC, Weisenberger DJ, Gonzales FA, Liang G, Xu GL, Hu YG et al. Continuous zebularine treatment effectively sustains demethylation in human bladder cancer cells. Mol Cell Biol 2004; 24: 1270–1278.
Liang G, Gonzales FA, Jones PA, Orntoft TF, Thykjaer T . Analysis of gene induction in human fibroblasts and bladder cancer cells exposed to the methylation inhibitor 5-aza-2′-deoxycytidine. Cancer Res 2002; 62: 961–966.
Nguyen CT, Weisenberger DJ, Velicescu M, Gonzales FA, Lin JC, Liang G et al. Histone H3-lysine 9 methylation is associated with aberrant gene silencing in cancer cells and is rapidly reversed by 5-aza-2′-deoxycytidine. Cancer Res 2002; 62: 6456–6461.
Stresemann C, Brueckner B, Musch T, Stopper H, Lyko F . Functional diversity of DNA methyltransferase inhibitors in human cancer cell lines. Cancer Res 2006; 66: 2794–2800.
Gius D, Cui H, Bradbury CM, Cook J, Smart DK, Zhao S et al. Distinct effects on gene expression of chemical and genetic manipulation of the cancer epigenome revealed by a multimodality approach. Cancer Cell 2004; 6: 361–371.
Karpf AR, Lasek AW, Ririe TO, Hanks AN, Grossman D, Jones DA . Limited gene activation in tumor and normal epithelial cells treated with the DNA methyltransferase inhibitor 5-aza-2′-deoxycytidine. Mol Pharmacol 2004; 65: 18–27.
Cheng JC, Yoo CB, Weisenberger DJ, Chuang J, Wozniak C, Liang G et al. Preferential response of cancer cells to zebularine. Cancer Cell 2004; 6: 151–158.
Rüter B, Wijermans P, Claus R, Kunzmann R, Lübbert M . Preferential cytogenetic response to continuous intravenous low-dose decitabine (DAC) administration in myelodysplastic syndrome with monosomy 7. Blood 2007; 110: 1080–1082.
Burnett AK, Milligan D, Prentice AG, Goldstone AH, McMullin MF, Hills RK et al. A comparison of low-dose cytarabine and hydroxyurea with or without all-trans retinoic acid for acute myeloid leukemia and high-risk myelodysplastic syndrome in patients not considered fit for intensive treatment. Cancer 2007; 109: 1114–1124.
Borthakur G, Ahdab SE, Ravandi F, Faderl S, Ferrajoli A, Newman B et al. Activity of decitabine in patients with myelodysplastic syndrome previously treated with azacitidine. Leuk Lymphoma 2008; 49: 690–695.
Acknowledgements
Grant support: Kind Philipp Foundation T237/15893/2006 (to CF); José Carreras Leukemia Foundation DJCLS R06/42f (to ML); Research Commission of the Faculty of Medicine of the University of Freiburg (to CF and ML); Deutsche Forschungsgemeinschaft (to RC). We thank Dr A Heinzmann for providing the Affymetrix facility and J Heinze for excellent technical assistance with capillary sequencing. We sincerely thank Dr Peter A Jones, Dr Richard Momparler and Dr James Downing for critically reading the paper and providing helpful comments.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Leukemia website (http://www.nature.com/leu)
Rights and permissions
About this article
Cite this article
Flotho, C., Claus, R., Batz, C. et al. The DNA methyltransferase inhibitors azacitidine, decitabine and zebularine exert differential effects on cancer gene expression in acute myeloid leukemia cells. Leukemia 23, 1019–1028 (2009). https://doi.org/10.1038/leu.2008.397
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2008.397
Keywords
This article is cited by
-
Prophylactic versus Preemptive modified donor lymphocyte infusion for high-risk acute leukemia after allogeneic hematopoietic stem cell transplantation: a multicenter retrospective study
Bone Marrow Transplantation (2024)
-
The methylome of Biomphalaria glabrata and other mollusks: enduring modification of epigenetic landscape and phenotypic traits by a new DNA methylation inhibitor
Epigenetics & Chromatin (2021)
-
Micro-RNA-125a mediates the effects of hypomethylating agents in chronic myelomonocytic leukemia
Clinical Epigenetics (2021)
-
Identification of RASSF1A promoter hypermethylation as a biomarker for hepatocellular carcinoma
Cancer Cell International (2020)
-
Engineering nanoparticles to reprogram radiotherapy and immunotherapy: recent advances and future challenges
Journal of Nanobiotechnology (2020)