Abstract
Aberrant expression of cytosolic sulfotransferase 2B1b (SULT2B1b) has been reported in several human malignancies. However, the expression pattern and clinical significance of SULT2B1b in colorectal carcinoma (CRC) remains unknown. Real-time quantitative PCR, western blot, and immunohistochemistry analyses were used to determine SULT2B1b expression in CRC clinical samples and CRC-derived cell lines. Kaplan–Meier and Cox proportional regression analyses were used to evaluate the association between SULT2B1b expression and patient survival in two independent cohorts of 485 patients with CRC. Gain- and loss-of-function approaches were employed to investigate the role of SULT2B1b in regulation of CRC cell growth and invasion. We found that SULT2B1b expression was frequently upregulated in CRC clinical samples and CRC-derived cell lines and was significantly correlated with lymph node metastasis and TNM stage in both the training and validation cohorts. Patients with higher intratumoral SULT2B1b expression had a significantly shorter disease-specific survival (DSS) and disease-free survival (DFS) than those with lower expression. Importantly, increased expression of SULT2B1b significantly predicted poor DSS and DFS and was an independent unfavorable prognostic indicator for stage II patients in both cohorts. Functional studies revealed that overexpression of SULT2B1b promoted CRC cell growth and invasion in vitro. Conversely, knockdown of SULT2B1b inhibited these processes. In conclusion, our findings suggest that SULT2B1b expression correlates with disease progression and metastasis and may serve as a novel prognostic biomarker and potential therapeutic target for patients with CRC.
Similar content being viewed by others
Main
Colorectal carcinoma (CRC) is the third most commonly diagnosed cancer in men and the second in women worldwide, accounting for more than 1.2 million new cases and 600 000 deaths per year.1, 2 Although the survival of patients with CRC has improved in western societies over the past decades, mortality rates have continued to increase in countries or areas including China.3, 4 Surgery remains the mainstay of curative treatment. Nevertheless, due to postsurgical recurrence and fatal distant metastasis, the prognosis for CRC patients has shown only limited improvement despite recent advancements in the treatment approaches. Therefore, an urgent search is needed for novel prognostic biomarkers and therapeutic targets for CRC.
Cytosolic sulfotransferase 2B1b (SULT2B1b), a member of the SULT2 family, is expressed in multiple normal tissues including prostate, skin, breast, intestine, lung, and placenta.5, 6 Previous studies have demonstrated that SULT2B1b plays an important role in cholesterol and oxysterol sulfate synthesis.7, 8, 9 It is highly selective for the addition of sulfate groups to 3β-hydroxysteroids such as cholesterol, oxysterols, DHEA, and is required for sulfating 25-hydroxychoelsterol into 5-Cholesten-3β-25-diol-3-sulfate (25HC3S), which is a new regulatory oxysterol.10, 11, 12 Recently, the growth-promoting effect of SULT2B1b has been reported. Zhang et al13 showed that overexpression of SULT2B1b promotes hepatocyte proliferation by inactivating oxysterol/LXR signaling, indicating its involvement in liver proliferation.
Up to now, aberrant expression of SULT2B1b has been documented in several human malignancies including breast, endometrial, prostate, and hepatocellular carcinomas.14, 15, 16, 17, 18, 19 It has been shown that expression of SULT2B1b is frequently upregulated in breast cancer, endometrial cancer, and liver cancer, but downregulated in prostate cancer. Functional studies revealed that SULT2B1b has the ability to promote proliferation of hepatocellular carcinoma cells both in vitro and in vivo,19 whereas impairs the proliferative potential of prostate cancer cells.18 These characteristics indicate that SULT2B1b have different expression patterns and functions in different types of human cancers.
A recent cDNA microarray-based analysis conducted by our group revealed that SULT2B1b is transcriptionally upregulated in CRC tissues. However, its expression pattern and clinical relevance in CRC has not been investigated to date. In the present study, we examined both the mRNA and protein expression levels of SULT2B1b in CRC clinical samples and cell lines and analyzed the correlation of SULT2B1b expression with clinicopathologic features and with patient survival in a training cohort and further validated our findings in an independent external cohort. Moreover, we assessed whether SULT2B1b influenced CRC cell proliferation, migration, and invasion in vitro. Our data suggest that SULT2B1b might represent a novel prognostic indicator and potential therapeutic target in CRC.
MATERIALS AND METHODS
Patients and Follow-Up
We obtained pathologically confirmed formalin-fixed paraffin-embedded tissue specimens of 485 stages I–III CRC patients with typical adenocarcinoma histology. Of these, 202 received curative surgery in 150th Hospital of PLA (Luoyang, China) between June 2006 and April 2008 and 283 received curative surgery in Changzheng Hospital, Second Military Medical University (Shanghai, China) between January 2008 and December 2009. Distribution of the continuous variables of the two study cohorts was listed in Supplementary Table S1 and S2. Detailed clinicopathologic features of CRC patients were listed in Table 1. The follow-up period was defined as the interval from the date of surgery to the date of death or last follow-up. The final date of follow-up was 28 March 2014 for patients from 150th Hospital of PLA (the Luoyang cohort) and 21 February 2014 for patients from Changzheng Hospital (the Shanghai cohort). Disease-specific survival (DSS) was defined as the interval from the date of surgery to the date that patient died of CRC. Disease-free survival (DFS) was defined as the interval from the date of surgery to the date of disease recurrence; if recurrence was not diagnosed, patients were censored on the date of death or last follow-up. Patients were excluded from the study cohorts with the following exclusion criteria: previously received any anticancer therapy; impaired heart, lung, liver, or kidney function; previous malignant disease. TNM staging was classified according to the American Joint Committee on Cancer staging manual (seventh edition).
Fresh-frozen CRC samples obtained from 90 stages I–III primary CRC patients who received curative surgery in 150th Hospital of PLA from April 2013 to September 2013 were used for quantitative PCR (qPCR) and western blot analysis. Written informed consent was obtained from each patient and this study was approved by the Ethical Committee of 150th Hospital of PLA and Changzheng Hospital.
Real-Time qPCR Analysis
Real-time qPCR analysis was performed as described previously.20 Briefly, total RNAs were isolated from frozen specimens using TRIzol Reagent (Ambion, 80706, USA). Reverse transcription was performed using RevertAidTM First Strand cDNA Synthesis Kit (Thermo Scientific, K1622, Lithuania) according to the manufacturer’s instructions. QPCR was performed on ABI Prism 7500 Sequence Detection System with SYBR Premix Ex Taq II (Takara, RR820A, Japan) using the 2−ΔΔCT method. Gene expression results were normalized by internal control β-actin. The primers used in this study are as follows: SULT2B1b (NM_177973.1) forward, 5′-GCTTGTGGGACACCTATGAAGA-3′; reverse, 5′-TGTTCTCCGCCAAGCTGATG-3′; β-actin forward, 5′-AATCGTGCGTGACATTAAGGAG-3′; reverse, 5′-ACTGTGTTGGCGTACAGGTCTT-3′. Each sample was tested in triplicate.
Western Blot Analysis
Western blot analysis was performed as described previously.21 Briefly, tumor specimens were prepared in lysis buffer (Tris-HCl (20 mM), pH 7.4, NaCl (150 mM), glycerol (10%), Nonidet P-40 (0.2%), EDTA (1 mM), EGTA (1 mM), PMSF (1 mM), NaF (10 mM), aprotinin (5 mg/ml), leupeptin (20 mM), and sodium orthovanadate (1 mM)) and centrifuged at 12 000 g for 30 min. Protein concentrations were measured using the BCA assay. Immunoblotting was performed using a primary antibody specific for SULT2B1b (Abnova, H00006820-B01P, Taiwan) and immunocomplexes were incubated with an anti-mouse horseradish-peroxidase-conjugated secondary antibody, and then detected using an enhanced chemiluminescence kit (Santa Cruz Biotechnology, sc-2048, Japan). β-actin was used as a loading control (Santa Cruz Biotechnology, sc-47778).
Tissue Microarray and Immunohistochemistry
Tissue microarrays containing the specimens from 150th Hospital of PLA were constructed (Shanghai Biochip Company, Shanghai, China). Immunohistochemistry was performed as described previously.22 Briefly, sections were deparaffinized and rehydrated. The endogenous peroxidase activity was blocked with 3% H2O2 for 10 min. Antigens were retrieved with citrate buffer (10 mM, pH 6.0) for 15 min at 100 °C in a microwave oven. After blocking, the sections were incubated with a primary anti-SULT2B1b antibody (R&D Systems, AF6174, USA) with 1:100 dilution at 4 °C overnight in a moist chamber followed by incubaton with an anti-sheep peroxidase-conjugated secondary antibody (Abcam, ab97130, USA) at room temperature for 30 min. Finally, the visualization signal was developed with diaminobenzidine and the slides were counterstained with hematoxylin.
Stained sections were evaluated in a blinded manner without prior knowledge of the clinical data using the German immunoreactive score (IRS) as described previously.23, 24 Briefly, staining intensity was graded as ‘0’ (negative), ‘1’ (weak), ‘2’ (moderate), and ‘3’ (strong); staining extent was graded as ‘0’ (<5%), ‘1’ (5–25%), ‘2’ (25–50%), ‘3’ (50–75%), or ‘4’ (>75%). Values of the staining intensity and the staining extent were multiplied as a final IRS of SULT2B1b expression, which ranged from 0 to 12. Intratumoral SULT2B1b expression was defined as follows: low expression with the IRS<6 and high expression with the IRS≥6. Discrepancies in the IRS were resolved by discussing together with other pathologists to reach a consensus. Tissue samples of patients from the Luoyang cohort were used as a training set. Prognostic value of the expression of SULT2B1b was subsequently validated in the patients from the Shanghai cohort as an external validation set.
Cell Lines, Cell Culture, and Lentivirus Infection
Normal human colon mucosal epithelial cell line NCM460 and CRC cell lines LS174T, DLD1, HCT15, CaCO-2, HT29, SW480, RKO, SW620, HCT116, and LOVO were purchased from Cell Bank of Type Culture Collection of Chinese Academy of Sciences (Shanghai, China). All cell lines were maintained at 37 °C in a humidified incubator containing 5% CO2 in Dulbecco’s modified Eagle’s medium or RPMI-1640 supplemented with 10% heat-inactivated fetal bovine serum and passed every 2–3 days to maintain logarithmic growth.
Lentiviral vectors expressing full-length human SULT2B1b protein (LV-SULT2B1b) or GFP (LV–GFP) and lentiviral vectors containing siRNA targeting human SULT2B1b (LV-siSULT2B1b) or scrambled control siRNA (LV-siSCR) were prepared by the Genechem Company (Shanghai, China). The following siRNA specific for SULT2B1b was used: 5′-CAGAUCUUCACCAAGGCCUUCUUCA-3′ (Invitrogen #SULT2B1HSS186159). LV–GFP and LV-siSCR were used as negative controls. For lentivirus infection, cells were infected with the indicated virus at a multiplicity of infection (MOI) of 10 in the presence of polybrene (8 mg/ml) for 8 h. Twenty-four hours later, the supernatant was replaced with fresh medium. After infection, stable colonies were selected in medium containing 3 μg/ml puromycin for 2–3 weeks. Expression of SULT2B1b in the infected cells was validated by Western blot assay.
Cell Proliferation Assay
The cell proliferation assay was performed using the Cell Counting Kit-8 solution (Dojindo Laboratories, Japan) according to the manufacturer’s instruction. Briefly, cells were seeded at a density of 4 × 103/well in 96-well plates and treated with 10 μl/well of the Cell Counting Kit-8 solution and cell viability was measured at the indicated times. The optical density of the well was measured at 450 nm using a microplate reader.
Colony-Formation Assay
Cells were trypsinized to generate a single-cell suspension, and 500 cells/well were seeded into 6-well plates. Dishes were returned to the incubator for 14 days, and the colonies were fixed with methanol for 1 h at room temperature and then stained with 0.5% crystal violet for additional 1 h.
Cell Migration and Invasion Assay
Migration and invasion assays were performed as described previously.25 Briefly, Cells were trypsinized, centrifuged, and resuspended in serum-free medium followed by plating into the upper chamber at a density of 2 × 105/well. Complete medium (700 μl) was added to the lower chamber as a chemoattractant. After incubation for 16–18 h for the migration assay, or after incubation for 20–24 h for the invasion assay, cells were fixed in methanol and stained with 0.1% crystal violet. Cells on the upper surface of the chamber were removed by wiping with a cotton swab and migration and invasion were determined by counting the cells that migrated to the lower side of the chamber using a microscope at × 100 magnification. Six random microscopic fields were counted per chamber in each group, and these experiments were repeated at least three times.
Statistical Analysis
Mann–Whitney U-test was used to compare SULT2B1b levels between groups. Pearson χ2-test or Fisher’s exact test was used to analyze the relationship between SULT2B1b expression and clinicopathologic features. Kaplan–Meier analysis with log-rank test was used to estimate DSS and DFS between subgroups. Multivariate Cox regression analysis was performed to determine the effect of the clinicopathologic variables and SULT2B1b expression on survivals. All statistical tests were two-sided and P-value<0.05 was considered to be statistically significant. Statistical analyses were performed using SPSS PASW Statistics 18.0 software (SPSS, Chicago, USA).
RESULTS
SULT2B1b is Frequently Overexpressed in CRC Tissues and Cell Lines
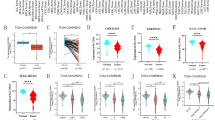
We first determined the expression levels of SULT2B1b mRNA in 55 paired primary CRC tissues and corresponding adjacent nontumor tissues using real-time qPCR methods. As shown in Figure 1a, SULT2B1b mRNA expression was significantly upregulated in tumor tissues compared with adjacent nontumor tissues (41/55, P<0.001). In addition, Western blot analysis from an independent set of 35 paired CRC and adjacent nontumor specimens confirmed that SULT2B1b protein levels were significantly higher in tumor tissues than in adjacent nontumor tissues (Figure 1b, 29/35, P<0.001).
SULT2B1b is frequently upregulated in CRC tissues and cell lines. (a) SULT2B1b mRNA expression in 55 paired human primary CRC tissues and corresponding adjacent nontumor tissues were determined by real-time qPCR methods. Gene expression results were normalized by internal control β-actin. (T, tumor tissues; N, adjacent nontumor tissues) (b) Protein levels of SULT2B1b in an independent set of 35 paired CRC and adjacent nontumor specimens were determined by western blot assay. β-actin was used as a loading control. (c and d) Expression levels of SULT2B1b mRNA (c) and protein (d) in NCM460 and CRC-derived cell lines. CRC, colorectal carcinoma; qPCR, quantitative PCR; SULT2B1b, sulfotransferase 2B1b.
Furthermore, we examined the levels of SULT2B1b mRNA and protein in 10 human CRC-derived cell lines (LS174T, DLD1, HCT15, CaCO-2, HT29, SW480, RKO, SW620, HCT116, and LOVO) and the normal colon epithelial cell line NCM460. Consistent with the results for the tissues, SULT2B1b was significantly upregulated in the most CRC cell lines examined when compared with the NCM460 cells at both the mRNA and protein levels (Figures 1c and d). Thus, these data demonstrate that SULT2B1b expression is frequently upregulated in CRC tissues and cell lines.
Expression of SULT2B1b in CRC Specimens is Associated with Aggressive Clinicopathologic Features
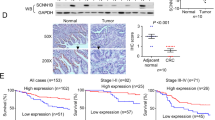
To investigate the clinical significance of SULT2B1b in the development and progression of CRC, SULT2B1b protein expression levels were determined immunohistochemically in two independent cohorts of patients with CRC who had undergone surgical resection. A group of 202 patients from the 150th Hospital of PLA were enrolled as the training group (Luoyang cohort). In addition, a second group of 283 patients from the Changzheng Hospital were enrolled as the validation group (Shanghai cohort). The representative staining of the SULT2B1b protein (negative, weak, moderate, strong) in CRC tissues was shown in Figure 2a, and positive staining of SULT2B1b was observed mainly in the cytoplasm. In the cancerous specimens tested from the Luoyang cohort, 19.3% (39/202) of cases presented strong immunostaining, 45% (91/202) showed moderate staining, 23.3% (47/202) showed weak staining and only 12.4% (25/202) showed negative staining of SULT2B1b protein. In striking contrast, only 19.8% (40/202) of the adjacent normal mucosa tissues showed weak to moderate SULT2B1b immunoreactivity (Figure 2b, P<0.001). Likewise, immunohistochemical data from the Shanghai cohort yielded a similar result (Figure 2c, P<0.001).
Upregulated SULT2B1b expression predicts poor prognosis in patients with CRC. (a) Representative immunohistochemical expression patterns of SULT2B1b in cancerous and adjacent normal mucosa specimens were shown. (Magnification: upper panel, × 100; lower panel, × 400) (b and c) Percentage of cases with different staining intensity of SULT2B1b in the tumor or adjacent normal tissues in the Luoyang cohort (b) and Shanghai cohort (c). **P<0.001. (d) Kaplan–Meier curves for disease-specific survival and disease-free survival of all patients (upper panel) or stage II patients (lower panel) in the Luoyang cohort according to SULT2B1b expression status. The P-value was determined using the log-rank test. (e) Kaplan–Meier curves for disease-specific survival and disease-free survival of all patients (upper panel) or stage II patients (lower panel) in the Shanghai cohort according to SULT2B1b expression status. The P-value was determined using the log-rank test. SULT2B1b, sulfotransferase 2B1b.
To evaluate the association between SULT2B1b expression levels and clinicopathologic characteristics, the patients in the Luoyang cohort were divided into high and low SULT2B1b expression subgroups with the median IRS value as the cut-off. As shown in Table 2, the upregulation of SULT2B1b in CRC tissues was significantly correlated with several aggressive clinicopathologic features, including poor differentiation grade (P=0.049), advanced local invasion (P=0.001), increased lymph node metastasis (P=0.048), advanced TNM stage (P=0.004), early disease recurrence (P<0.001), and patient death (P<0.001), while no significant correlations were observed between SULT2B1b expression and age, sex, tumor location, or tumor size.
We then applied the same cut-off to dichotomize the study patients in the Shanghai cohort. Consistently, high expression of SULT2B1b was significantly associated with increased lymph node metastasis (P=0.028), advanced TNM stage (P=0.022), early recurrence (P<0.001), and death (P<0.001). Hence, these results support the notion that SULT2B1b may be involved in the progression of CRC.
Upregulated SULT2B1b Expression Predicts Poor Prognosis in Patients with CRC
We next assessed the association between SULT2B1b expression and clinical prognosis of CRC patients. Kaplan–Meier survival analyses revealed that patients with higher intratumoral SULT2B1b expression had significantly poorer DSS (P<0.001) and DFS (P<0.001) rates than those with lower SULT2B1b expression in the Luoyang cohort (Figure 2d). The cumulative 5-year DSS and DFS rates were 79.1% and 66.3% in the low-SULT2B1b-expression patients, whereas it was only 44.0% and 37.9% in the high-SULT2B1b-expression ones, respectively. Similarly, patients with high SULT2B1b expression exhibited significantly shorter DSS (P<0.001) and DFS (P<0.001) than those with low SULT2B1b expression in the Shanghai cohort (Figure 2e). Importantly, increased expression of SULT2B1b significantly predicted poor DSS (P<0.001) and DFS (P<0.001) for stage II patients in the Luoyang cohort (Figure 2d). Similarly, high levels of SULT2B1b protein also significantly predicted unfavorable DSS (P<0.001) and DFS (P<0.001) for stage II patients in the Shanghai cohort (Figure 2e). In both cohorts, patients with stage III tumors had a significantly worse prognosis compared with those with stage I or II tumors (Supplementary Figure S1). These data were consistent with the established adverse prognostic effect of tumor stage26 and confirmed that our cohorts were representative and that the survival analyses were valid.
The multivariate Cox regression analyses further demonstrated that SULT2B1b expression level (P<0.001), together with TNM stage (P<0.001) and patient age (P=0.015), was an independent risk factor for DSS, while SULT2B1b expression level (P=0.002) and TNM stage (P<0.001) were independent risk factors for DFS in the Luoyang cohort (Table 3). Similarly, high SULT2B1b expression and advanced TNM stage were independently associated with poor DSS and DFS in the Shanghai cohort (Table 4). More importantly, further analyses revealed that increased expression of SULT2B1b also was an independent predictor of poor prognosis for stage II CRC patients in both cohorts (Table 5). Collectively, these findings suggest that the expression level of SULT2B1b could be used as an independent factor for predicting the prognosis of CRC.
Exogenous Expression of SULT2B1b Promotes CRC Cell Growth and Invasion
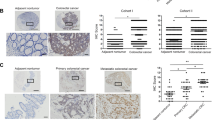
To examine the role of SULT2B1b in CRC progression, we established stable expression of SULT2B1b in two human CRC cell lines, SW480 and RKO, which have low levels of endogenous SULT2B1b, using a lentiviral system (Figure 3a). The Cell Counting Kit-8 assay showed that forced expression of SULT2B1b induced a significant increase of the proliferation rate (Figure 3b). The enhanced proliferative ability of SULT2B1b- overexpressing cells was further validated by the colony-formation assay (Figure 3c). In addition, The transwell migration and matrigel invasion assay demonstrated that upregulation of SULT2B1b markedly enhanced the migratory and invasive abilities of SW480 and RKO cells (Figures 3d and e). These data indicate that SULT2B1b contributes to the progression of CRC in vitro.
Overexpression of SULT2B1b promotes CRC cell proliferation, motility, and invasion. (a) SW480 or RKO cells were infected with lentiviral-delivering GFP (LV–GFP) or SULT2B1b (LV-SULT2B1b) and protein levels of SULT2B1b were evaluated by western blot assay. (b) Cell viability of SW480 or RKO cells infected with LV–GFP or LV-SULT2B1b was determined by the Cell Counting Kit 8 assay. Plots are represented as mean±s.e.m. of data from three independent experiments. *P<0.05. (c–e) Effects of SULT2B1b overexpression on the proliferation, migration, and invasion of SW480 or RKO cells were assessed by the colony-forming assay (c), transwell migration assay (d), and matrigel invasion assay (e), respectively. Representative results are shown in the left panel. Plots in the right panel are represented as mean±s.e.m. of data from three independent experiments. *P<0.05.
SULT2B1b Silencing Suppresses the Proliferative and Invasive Potential of CRC Cells
To further determine whether SULT2B1b is required for the growth and motility of CRC cells, we then used lentivirus-mediated siRNA to generate SULT2B1b-knockdown DLD1 and HCT15 cell models (Figure 4a). As expected, a significant reduction in cell growth was observed in SULT2B1b-depleted DLD1 and HCT15 cells (Figure 4b). In addition, knockdown of SULT2B1b strongly impaired the ability to form colonies in each cell line (Figure 4c). Moreover, SULT2B1b depletion significantly suppressed the migration and invasion capabilities of DLD1 and HCT15 cells (Figures 4d and e). Taken together, these results suggest that SULT2B1b is essential for the proliferative and invasive properties of CRC cells.
SULT2B1b silencing inhibits the proliferation, motility, and invasion of CRC cells. (a) DLD1 or HCT15 cells were infected with lentiviral-delivering scrambled control siRNA (LV-siSCR) or SULT2B1b siRNA (LV-siSULT2B1b) and protein levels of SULT2B1b were evaluated by western blot assay. (b) Cell viability of DLD1 or HCT15 cells infected with LV-siSCR or LV-siSULT2B1b was determined by the Cell Counting Kit 8 assay. Plots are represented as mean±s.e.m. of data from three independent experiments. *P<0.05. (c–e) Effects of SULT2B1b depletion on the proliferation, migration, and invasion of DLD1 or HCT15 cells were assessed by the colony-forming assay (c), transwell migration assay (d), and matrigel invasion assay (e), respectively. Representative results are shown in the left panel. Plots in the right panel are represented as mean±s.e.m. of data from three independent experiments. *P<0.05.
DISCUSSION
Over the past decades, great efforts have been made to elucidate the molecular mechanisms underlying the tumorigenesis and progression of CRC to identify novel prognostic biomarkers and therapeutic targets for this malignancy.3, 27, 28 To date, there is evidence linking SULT2B1b gene to tumor growth of hepatocellular and prostate carcinomas.18, 19 However, the reported effects of SULT2B1b on tumor biology are conflicting. Yang et al19 showed that overexpression of SULT2B1b promoted the proliferation of human hepatocarcinoma cells, whereas Seo et al18 demonstrated that SULT2B1b expression was reduced in prostate cancer and knockdown of SULT2B1b resulted in increased proliferation rate of prostate cancer cells. These conflicting results indicate that the role of SULT2B1b in different cancers is tissue dependent and varies with the type of malignancy. Recently, several SNPs in SULT2B1 gene have been reported to be associated with risk of progression in prostate and esophageal carcinomas,29, 30 implying a possible relationship between SULT2B1b and clinical prognosis of cancer patients. Nevertheless, the expression pattern and clinical relevance of SULT2B1b has not been assessed in CRC.
In the current study, we found that expression of SULT2B1b was significantly upregulated in CRC tissues as compared with adjacent noncancerous counterparts at both the mRNA and protein levels. Subsequent immunohistochemical analysis of CRC specimens from the training and validation cohorts demonstrated that 87.6 and 90.2% of the cancerous tissues tested were positive for SULT2B1b protein expression, whereas only 19.8 and 19.4% of the adjacent normal mucosa tissues were SULT2B1b positive. These findings definitely confirmed the significant upregulation of SULT2B1b protein in CRC, suggesting that SULT2B1b might be a new candidate diagnostic tool or marker for CRC. Similar to what we observed in the present study, earlier investigations showed that expression of SULT2B1b was significantly increased in breast and liver cancer tissues relative to their corresponding nontumor counterparts.6, 14, 19, 31 Of note, although SULT2B1a and SULT2B1b are splice variants of the same gene differing only at their amino-terminal ends, we did not observe a substantial increase in the expression level of SULT2B1a mRNA in cancerous tissues (Supplementary Figure S2), indicating that the two isoforms have different expression patterns in CRC. Nevertheless, the molecular basis for SULT2B1b overexpression in CRC is currently unknown and requires further investigation.
Our immunohistochemical results revealed that positive staining of SULT2B1b was mainly located in the cytoplasm of carcinoma cells. Previously, He et al5 reported that SULT2B1b was expressed only in the cytosol of human prostate epithelial cells, prostate benign prostatic hyperplasia, prostate adenocarcinoma, and LNCaP prostate adenocarcinoma cells. However, Dumas et al31 demonstrated that SULT2B1b was localized in both cytosol and nuclei of cancerous and associated-normal breast tissue. In addition, they showed that there was a different expression pattern of cytoplasmic: nuclear localization of SULT2B1b in breast cancer in Caucasians and African Americans.31 While, in the present study, we did not observe apparent nuclear localization of SULT2B1b in CRC tissues or adjacent normal tissues. The discrepant expression pattern of SULT2B1b in different malignancies indicates that its subcellular localization is tissue specific; however, the underlying mechanism for this discrepancy remains to be elucidated.
Previous studies have showed that SULT2B1b is largely involved in the control of cell growth and survival. Its expression is increased in liver regeneration after partial hepatectomy32 and it has the ability to promote hepatocyte proliferation and protect cells against 7-ketocholesterol-induced loss of cell viability.8, 13 Conversely, SULT2B1b depletion by siRNA induces cell-cycle arrest and triggers apoptosis.19 Interestingly, in this study, correlation analyses with clinicopathologic features from the two independent patient cohorts unanimously revealed a significant association between SULT2B1b expression and lymph node metastasis, TNM stage, and disease recurrence. These results suggest that SULT2B1b may have an important role in tumor progression and dissemination. Therefore, we further examined the effects of SULT2B1b overexpression as well as knockdown on the aggressive behaviors of CRC in vitro. Our results demonstrated that SULT2B1b played a positive role in regulating the proliferation, migration, and invasion of CRC cells, which may explain why higher SULT2B1b expression was significantly correlated with more lymph node metastasis, more advanced TNM stage, and earlier recurrence. In addition, results from this study suggest that pharmacological suppression of SULT2B1b may represent a promising approach for CRC treatment.
Another interesting finding of the present study lies on the survival analysis results. Increased expression of SULT2B1b was demonstrated to be associated with shortened survival for CRC patients not only in the training cohort but also in the validation cohort. In the multivariate Cox regression analyses, SULT2B1b protein emerged as a significant independent predictor of survival along with TNM stage in both of the two cohorts. Our current results indicated that TNM stage also is an important prognostic factor in CRC, which is consistent with the well recognized unfavorable prognostic effect of tumor stage.26 In general, patients with early-stage CRC (stages I-II) have a better prognosis than those with advanced-stage CRC (stages III-IV). Nevertheless, a subgroup of stage II CRC patients has an increased risk of early recurrence and death. Therefore, identification of this high-risk subgroup would be of particular clinical importance in the selection of patients for appropriate treatment. To further evaluate the prognostic value of SULT2B1b and its potential implication in the therapeutic decision-making, we performed survival analyses based on TNM stage. Importantly, SULT2B1b efficiently discriminated stage II patients with distinct prognosis, and moreover it was still an independent unfavorable prognostic indicator for stage II patients in both cohorts. Thus, our results suggest that SULT2B1b expression status may serve as a valuable prognostic marker to stratify stage II CRC patients into different risk subgroups and help clinicians to design individualized therapies in time.
This study had some limitations. Although our results demonstrated the prognostic value of SULT2B1b expression in two cohorts of patients with CRC and its effects on CRC cells, they did not explore the consequences of SULT2B1b overexpression or knockdown in animal models of CRC. In addition, the underlying mechanism by which SULT2B1b contributes to the development and progression of CRC has yet to be elucidated. Therefore, further in vivo experiments and mechanistic studies are warranted to confirm our findings and to provide a better understanding of the molecular events involved in the SULT2B1b-mediated cancer development and progression.
In conclusion, we report here, for the first time, that SULT2B1b is frequently upregulated in CRC tissues and CRC-derived cell lines. Increased expression of SULT2B1b protein was significantly correlated with disease progression and poor postoperative prognosis of CRC patients. SULT2B1b may have a major role in CRC proliferation and metastasis and could be a promising prognostic predictor for both DSS and DFS in CRC patients. Our data was validated in two independent cohorts. Combination of SULT2B1b with other prognostic biomarkers may enhance its performance in prognosis prediction. In addition to its prognostic value, our findings pave the road for further investigation of SULT2B1b as a potential therapeutic target for the treatment of CRC.
References
Jemal A, Bray F, Center MM et al. Global cancer statistics. CA Cancer J Clin 2011;61:69–90.
Ferlay J, Shin HR, Bray F et al. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer 2010;127:2893–2917.
Brenner H, Kloor M, Pox CP . Colorectal cancer. Lancet 2014;383:1490–1502.
Guo P, Huang ZL, Yu P et al. Trends in cancer mortality in China: an update. Ann Oncol 2012;23:2755–2762.
He D, Meloche CA, Dumas NA et al. Different subcellular localization of sulphotransferase 2B1b in human placenta and prostate. Biochem J 2004;379:533–540.
Falany CN, He D, Dumas N et al. Human cytosolic sulfotransferase 2B1: isoform expression, tissue specificity and subcellular localization. J Steroid Biochem Mol Biol 2006;102:214–221.
Strott CA . Sulfonation and molecular action. Endocr Rev 2002;23:703–732.
Fuda H, Javitt NB, Mitamura K et al. Oxysterols are substrates for cholesterol sulfotransferase. J Lipid Res 2007;48:1343–1352.
Falany CN, Rohn-Glowacki KJ . SULT2B1: unique properties and characteristics of a hydroxysteroid sulfotransferase family. Drug Metab Rev 2013;45:388–400.
Li X, Pandak WM, Erickson SK et al. Biosynthesis of the regulatory oxysterol, 5-cholesten-3beta,25-diol 3-sulfate, in hepatocytes. J Lipid Res 2007;48:2587–2596.
Bai Q, Zhang X, Xu L et al. Oxysterol sulfation by cytosolic sulfotransferase suppresses liver X receptor/sterol regulatory element binding protein-1c signaling pathway and reduces serum and hepatic lipids in mouse models of nonalcoholic fatty liver disease. Metabolism 2012;61:836–845.
Ren S, Ning Y . Sulfation of 25-hydroxycholesterol regulates lipid metabolism, inflammatory responses, and cell proliferation. Am J Physiol Endocrinol Metab 2014;306:E123–E130.
Zhang X, Bai Q, Xu L et al. Cytosolic sulfotransferase 2B1b promotes hepatocyte proliferation gene expression in vivo and in vitro. Am J Physiol Gastrointest Liver Physiol 2012;303:G344–G355.
Bieche I, Girault I, Urbain E et al. Relationship between intratumoral expression of genes coding for xenobiotic-metabolizing enzymes and benefit from adjuvant tamoxifen in estrogen receptor alpha-positive postmenopausal breast carcinoma. Breast Cancer Res 2004;6:R252–R263.
Tozlu S, Girault I, Vacher S et al. Identification of novel genes that co-cluster with estrogen receptor alpha in breast tumor biopsy specimens, using a large-scale real-time reverse transcription-PCR approach. Endocr Relat Cancer 2006;13:1109–1120.
Hevir N, Sinkovec J, Rizner TL . Disturbed expression of phase I and phase II estrogen-metabolizing enzymes in endometrial cancer: lower levels of CYP1B1 and increased expression of S-COMT. Mol Cell Endocrinol 2011;331:158–167.
He D, Falany CN . Inhibition of SULT2B1b expression alters effects of 3beta-hydroxysteroids on cell proliferation and steroid hormone receptor expression in human LNCaP prostate cancer cells. Prostate 2007;67:1318–1329.
Seo YK, Mirkheshti N, Song CS et al. SULT2B1b sulfotransferase: induction by vitamin D receptor and reduced expression in prostate cancer. Mol Endocrinol 2013;27:925–939.
Yang X, Xu Y, Guo F et al. Hydroxysteroid sulfotransferase SULT2B1b promotes hepatocellular carcinoma cells proliferation in vitro and in vivo. PLoS One 2013;8:e60853.
Hu L, Chen L, Yang G et al. HBx sensitizes cells to oxidative stress-induced apoptosis by accelerating the loss of Mcl-1 protein via caspase-3 cascade. Mol Cancer 2011;10:43.
Hu L, Chen L, Li L et al. Hepatitis B virus X protein enhances cisplatin-induced hepatotoxicity via a mechanism involving degradation of Mcl-1. J Virol 2011;85:3214–3228.
Li L, Chen L, Hu L et al. Nuclear factor high-mobility group box1 mediating the activation of Toll-like receptor 4 signaling in hepatocytes in the early stage of nonalcoholic fatty liver disease in mice. Hepatology 2011;54:1620–1630.
Tang L, Tan YX, Jiang BG et al. The prognostic significance and therapeutic potential of hedgehog signaling in intrahepatic cholangiocellular carcinoma. Clin Cancer Res 2013;19:2014–2024.
Li MH, Dong LW, Li SX et al. Expression of cytoskeleton-associated protein 4 is related to lymphatic metastasis and indicates prognosis of intrahepatic cholangiocarcinoma patients after surgery resection. Cancer Lett 2013;337:248–253.
Wang RY, Chen L, Chen HY et al. MUC15 inhibits dimerization of EGFR and PI3K-AKT signaling and is associated with aggressive hepatocellular carcinomas in patients. Gastroenterology 2013;145:1436–1448, e1431-e1412.
Webber C, Gospodarowicz M, Sobin LH et al. Improving the TNM classification: findings from a 10-year continuous literature review. Int J Cancer 2014;135:371–378.
Malesci A, Laghi L . Novel prognostic biomarkers in colorectal cancer. Dig Dis 2012;30:296–303.
Ludwig JA, Weinstein JN . Biomarkers in cancer staging, prognosis and treatment selection. Nat Rev Cancer 2005;5:845–856.
Levesque E, Laverdiere I, Audet-Walsh E et al. Steroidogenic germline polymorphism predictors of prostate cancer progression in the estradiol pathway. Clin Cancer Res 2014;20:2971–2983.
Hyland PL, Freedman ND, Hu N et al. Genetic variants in sex hormone metabolic pathway genes and risk of esophageal squamous cell carcinoma. Carcinogenesis 2013;34:1062–1068.
Dumas NA, He D, Frost AR et al. Sulfotransferase 2B1b in human breast: differences in subcellular localization in African American and Caucasian women. J Steroid Biochem Mol Biol 2008;111:171–177.
Lo Sasso G, Celli N, Caboni M et al. Down-regulation of the LXR transcriptome provides the requisite cholesterol levels to proliferating hepatocytes. Hepatology 2010;51:1334–1344.
Acknowledgements
This work was supported by funds from the National Natural Science Foundation of China (81301811).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website
Increased cytosolic sulfotransferase 2B1b (SULT2B1b) expression correlates with aggressive clinical behaviors and poor prognosis of patients with colorectal carcinoma (CRC), and contributes to cancer cell growth and invasion. This paper suggests that SULT2B1b has an important role in CRC progression and could be a novel prognostic marker and potential therapeutic target for CRC patients.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Hu, L., Yang, GZ., Zhang, Y. et al. Overexpression of SULT2B1b is an independent prognostic indicator and promotes cell growth and invasion in colorectal carcinoma. Lab Invest 95, 1005–1018 (2015). https://doi.org/10.1038/labinvest.2015.84
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2015.84
This article is cited by
-
Association study of genetic variants in estrogen metabolic pathway genes and colorectal cancer risk and survival
Archives of Toxicology (2018)
-
Metabolomics guided pathway analysis reveals link between cancer metastasis, cholesterol sulfate, and phospholipids
Cancer & Metabolism (2017)
-
The Anti-Inflammatory Effect and Intestinal Barrier Protection of HU210 Differentially Depend on TLR4 Signaling in Dextran Sulfate Sodium-Induced Murine Colitis
Digestive Diseases and Sciences (2017)