Abstract
Objective:
To identify the specific microRNA (miRNA) biomarkers of preeclampsia (PE), the miRNA profiles analysis were performed.
Study Design:
The blood samples were obtained from five PE patients and five normal healthy pregnant women. The small RNA profiles were analyzed to identify miRNA expression levels and find out miRNAs that may associate with PE. The quantitative reverse transcriptase–PCR (qRT-PCR) assay was used to validate differentially expressed peripheral leucocyte miRNAs in a new cohort.
Result:
The data analysis showed that 10 peripheral leucocyte miRNAs were significantly differently expressed in severe PE patients. Four differently expressed miRNAs were successfully validated using qRT-PCR method.
Conclusion:
We successfully constructed a model with high accuracy to predict PE. A combination of four peripheral leucocyte miRNAs has great potential to serve as diagnostic biomarkers of PE.
Similar content being viewed by others
Introduction
Preeclampsia (PE), a disease characterized by high blood pressure and proteinuria, is one of the causes of maternal and fetal morbidity and mortality and occurs in around 8% of pregnancies.1 Part of the PE may progress quickly and cause eclampsia, placental abruption, HELLP syndrome, neonatal asphyxia, disseminated intravascular coagulation, stroke, heart failure or death despite intensive treatment. Previous studies have shown that placental hypoxia enhanced platelet aggregation and endothelial dysfunction in vascular aspect and immunological dysfunctions were responsible for the occurring of PE.2 Even though several factors have been investigated, few effective results were revealed.3, 4, 5, 6 However, the pathogenesis of PE has not been elucidated, although it has been suggested that PE encompasses a combination of genetic, immune and environment aspects. Neither its useful biomarkers nor its risk factor has been found to reach an agreement.
MicroRNAs (miRNAs) are small non-coding RNAs of 18 to 25 nt long that affect the stability and translational efficiency of target mRNAs.7, 8 Abnormal expression levels of miRNAs have been associated with various diseases, including PE.9, 10 The specific expressed miRNA might be used as the biomarker of diseases.
Circulating miRNAs have proven as powerful biomarkers for human disease.11, 12 Many researchers have found a handful of miRNAs expressed as aberrant in PE samples.13, 14 Anton et al.15 reported that miR-210 may be a novel predictive serum biomarker for PE. On the other hand, Luque et al.16 recently declared maternal serum miRNA assessment at the first trimester of pregnancy showed no predictive value for early PE.
The miRNA expression in leukocytes has been confirmed a stable state17 that can be several diseases’ predicted markers.18, 19 To better address the role of miRNAs in PE, we extracted leukocytes from peripheral blood of PE patients and performed the miRNA sequencing of PE patients and normal healthy pregnant women using the second-generation sequencing technology. We found that a collection of miRNAs was expressed aberrantly in PE samples compared with the healthy pregnant women. Different expressed miRNA were validated by quantitative reverse transcriptase–PCR (qRT-PCR) in a large sample set. The receiver operating characteristic (ROC) analysis illustrated that a combination of four miRNA expression profiles might be used as biological markers for PE.
Methods
Ethics statement and samples
This study has approved by the Clinical Research Ethics Committee of Shanxi Medical College. PE was defined according to the criteria by American College of Obstetricians and Gynecologists practice bulletin on diagnosing and managing PE and eclampsia.20 A total of 575 cases were sampled from the Second Hospital of Shanxi Medical University between 2011 and 2014. We filtered the cases of drug treatment, severe anemia, kidney failure, severe hypoalbuminemia, disagreement for experiment and failure in RNA extraction, and 57 cases were excluded from the experiment. Written informed consents were obtained from all participates. Five PE patients and five normal pregnant women were included in the first discovery stage (Supplementary Tables S1 and S2). Forty-seven samples were included in the second validation stage (Supplementary Table S3), which were separated into four groups: the control (N) group (13 normal cases of pregnant women); the PE group (13 cases of PE women aged between 21 and 29 years); PE with advanced age (PA) group (13 cases of PE women aged >30 years); and PE women with complications (PC) group (8 cases of PE women with chronic hypertension and gestational diabetes). Considering the ethics and convenience of pregnancy test, the peripheral blood were collected from all participant.
RNA extraction and sequencing
The procedure of RNA extraction was as follows: (a) an equal volume phosphate-buffered saline (1 ×) was added to fresh whole blood with anticoagulant and mixed; (b) the mixture slowly transferred to another centrifuge tube, which contained the lymphocyte separated liquids and mixture was allowed to settle over lymphocyte separated liquid (so as to not mix the two liquids and keep the interface clear), centrifuged at 3000 × g for 30 min at 4 °C; (c) leucocyte was separated with a pipette carefully, washed with phosphate-buffered saline (1 ×), collected using centrifuge and completely removed and discarded the supernate; (d) TRIzol reagent was added with 20 volumes of lymphocyte, washed the lymphocyte until the cell block was broken, kept the entire solution clear but not the viscous state; and (e) the solution was poured into dry ice or stored at −80 °C. RNA quantity and integrity were evaluated using Agilent 2100 BioAnalyzer (Agilent Technologies, Santa Clara, California, USA). Small RNA libraries were constructed using the method described in previous study.21, 22 Briefly, for each library, 50 μg of the total RNA was size-fragmented on a 15% tris-borate-EDTA (TBE) urea polyacrylamide gel (Invitrogen, Waltham, Massachusetts, USA) and 15 to 30 base pair (bp) fraction was excised, using 10 bp ladder (Invitrogen) as marker. RNA was eluted from the polyacrylamide gel slice in 600 μl of 0.3 M NaCl overnight at 4 °C. The resulting gel slurry was passed through a Spin-X cellulose acetate filter column (Corning, Corning, New York, USA) and precipitated in two 300-μl aliquots by the addition of 750 μl of ethanol and 3 μl of glycogen (5 mg ml−1; Invitrogen). After washing with 75% ethanol, the pellets were allowed to air dry at 25 °C and dissolved in diethylpyrocarbonate (DEPC) water. The RNA was dephosphorylated by alkaline phosphatase and recovered by ethanol precipitation. The small RNA was ligated with 5′ adapter (5′-GUUCAGAGUUCUACAGUCCGACGAUC-3′) using T4 RNA ligase (Promega, Madison, Wisconsin, USA) in the presence of RNase Out (Invitrogen) overnight at 20 °C. The ligation reaction was stopped by the addition of 10 μl of 2 × Gel Loading Buffer II (Ambion, Waltham, Massachusetts, USA). The ligated RNA was size-fractionated on a 15% TBE urea polyacrylamide gel (Invitrogen), and a 40 to 70 bp fraction was excised. The RNA was eluted from the gel and precipitated as described above followed by resuspension in DEPC-treated water. The precipitated RNA was subsequently ligated to the 3′ RNA adapter (5′-pUCGUAUGCCGUCUUCUGC UUGidT-3′; p, phosphate; idT, inverted deoxythymidine) using T4 RNA ligase (Promega) in the presence of RNase Out (Invitrogen) overnight at 25 °C. The ligation reaction was stopped by the addition of 10 μl of 2 × Gel Loading Buffer II (Ambion). Ligated RNA was size-fractionated on a 10% TBE urea polyacrylamide gel (Invitrogen), and the 60 to 100 bp fraction was excised. The RNA was eluted from the polyacrylamide gel and precipitated from the gel as described above and resuspended in 5.0 μl of DEPC water. The RNA was converted to single-stranded cDNA using Superscript II reverse transcriptase (Invitrogen) and Illumina’s small RNA RT-Primer (5′-CAAGCA GAAGACGGCATACGA-3′) following the manufacturer’s instructions. The resulting cDNA was PCR-amplified with Hotstart Phusion DNA Polymerase (NEB, Ipswich, Massachusetts, USA) in 15 cycles using Illumina’s small RNA primer set (5′-CAAGCAGAAGACGGCATA CGA-3′; 5′-AATGATACGGCGACCACCGA-3′).
PCR products were purified on a 12% TBE urea polyacrylamide gel (Invitrogen), and the 100-bp fraction was excised. The DNA was eluted into 100 μl of 1 × NEBuffer 2 at room temperature for 2 h. The resulting gel slurry was passed through a Spin-X filter (Corning) and precipitated by the addition of 325 μl of ethanol, 10 μl of 3 M sodium acetate and 3 μl of glycogen (5 mg ml−1; Invitrogen). After washing with 75% ethanol, the pellet was allowed to air dry at 25 °C and dissolved in 10 ml of resuspension buffer (10 mM Tris-HCl, pH 8.5) by incubation at 4 °C for 10 min. The purified PCR products were quantified on the Agilent DNA 1000 chip and diluted to 10 nM for sequencing on the Illumina Hiseq 2000 platform (San Diego, California, USA) according to the manufacturer’s protocol.
Sequencing data analysis
After removing low-quality reads and adapter reads, the remaining reads with at least 18 nucleotides (nt) were aligned against human reference genome (version hg19) using SOAP (version 2.0, Microsoft Corporation, Redmond, Washington, USA).22 Perfect match sequence tags were then annotated as coding exons, repeats, rRNA, tRNA, snRNA and snoRNA by aligning sequence tags to the databases of UCSC RefGene, Repbase, NCBI Refseq and NCBI Genbank (http://www.ncbi.nlm.nih.gov/). Expression levels of known miRNAs were calculated after mapping the tags to the precursors/mature miRNA sequence in miRBase (version 18).23 NOISeq24 was used in the detection of differentially expressed miRNAs between the PE and healthy group under the criteria: probability >0.70 and log2 ratio >1.5. The target genes for differentially expressed miRNAs were predicted by DIANA-microT.25 To obtain dependable prediction, the DIANA-microT threshold was set to 0.9. Gene Ontology (GO) functional analysis26 and KEGG pathway27 analysis were carried out to identify significantly enriched GO terms and pathways associated with the target genes using miRPath,28 with a cutoff of corrected P-value<0.05.
Real-time PCR validation
miRNA-451a, miRNA-15a-3P, miRNA-31-3P and miRNA-122-5P were selected to conduct real-time PCR validation in a four-group data set as mentioned above. The known miRNA has-U6 was used as the endogenous control during the process. RT reactions were carried out by miRcute miRNA First-Strand cDNA Synthesis Kit (TIANGEN, Beijing, China) according to the manufacturer’s protocol. Amplification were conducted under the following condition: 94 °C for 20 s, then 94 °C for 20 s, and finally 60 °C for 34 s with 40 cycles. All reactions were run in triplicate. The relative expression level was calculated using the comparative CT method and normalized by the expression level of control miRNA has-U6.
The differential expression analysis of miRNAs between difference groups were analyzed by the method of analysis of variance tests using the software SPSS (version 18.0, SPSS Inc., Chicago, Illinois, USA). The significance P-value cutoff was set as 0.05.
Disease prediction model construction and ROC analysis
Logistic regression analysis of the four miRNA combination regression was carried out using glm package of R (http://www.r-project.org/). Relative expression level profiles of four miRNAs from qRT-PCR experiments were used as independent variable, and the PE status was used as dependent variable. Leave-one-out cross-validation accuracy was calculated using cv.binary from DAAG package of R (http://www.stats.uwo.ca/DAAG/). ROC analysis was carried out using pROC package of R.29
Results
Sequencing data description and small RNA annotations
Ten small RNA libraries were constructed and sequenced using second-generation sequencing technology. Primary sequencing data of each sample contained 12.98 M raw reads on an average. After filtering out about 3% low-quality and adaptor reads, 12.76 M final reads per sample were reserved. The length of most reads was located in 20 to 24 nt. In our study, >70% small RNAs were identified as miRNA.
Differentially expressed miRNAs
Eight known miRNAs were significantly differentially expressed. Among these eight miRNAs, four were significantly upregulated in PE with up to 10-fold changed level, and the expression of the other four miRNAs were downregulated in patients compared with the normal group (Table 1).
Based on the prediction of MircoT, 125 genes were identified as target genes for these eight miRNAs (Supplementary Figure S1). GO terms of sensory perception of chemical stimulus (GO: 0007606, P=0.044) and G-protein coupled receptor activity (GO:0004930, P=0.005) were significantly enriched in these target genes of differentially expressed of miRNAs. The overrepresented pathways of these target genes included mucin type O-Glycan biosynthesis (ko 00512, P=4.85 × 10−4) and endocytosis (ko 04144, P=7.83 × 10−4) (Table 2).
qRT-PCR validations
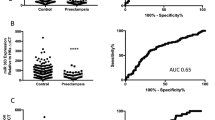
Of these eight differential expressed miRNAs we found in the sequencing stage, hsa-miR-31 and hsa-miR-15 were reported to be deregulated in other PE studies either by sequencing method or PCR validations,30, 31 miR-451 were differentially regulated in primary trophoblasts exposed to hypoxia32 and miR-122 were reported to be associated with gene GIT1 (G-protein-coupled receptor kinase interactor 1), a gene which is related to the endothelial cell repair.33 Therefore, we selected has miR-31, miR15, miR 451 and miR122 to perform qRT-PCR validation in other 47 subjects. Analysis of variance test showed that there were significant different means of the expression levels of each miRNA between groups. T-test showed that the expression levels of miR-15a-3p and miR-31-3p were significantly downregulated in the three patients groups compared with the healthy group. The expression levels of miRNA-451a and miRNA-122-5p were upregulated in PE group and downregulated in the PA and PC groups (Figure 1 and Table 3). We also found that expression of miR-122-5p was downregulated in the PA and PC groups compared with the PE group. Meanwhile, the expression of miR-122-5p in the PC group was significantly increased compared with the PA group.
PE prediction model construction and ROC analysis
As four miRNAs could significantly separate PE patients from healthy women, these miRNAs can be used as biomarkers for PE. We constructed a logistic regression model to evaluate these four miRNAs and compare the model with single marker using ROC analysis. The results showed that the combination of four miRNAs achieved the highest area under the ROC curve (AUC=0.99, Figure 2) and was better than other four single markers. To prevent over fitting of the logistic regression model, the leave-one-out cross validation were performed, and we got an estimated accuracy of 0.94.
Receiver operating characteristic (ROC) curves to assess the utility of microRNAs (miRNAs) to differentiate preeclampsia and healthy women. ROC curves for miR-122 (yellow, area under the curve (AUC)=0.59), miR-451 (black, AUC=0.90), miR-15a (green, AUC=0.96), miR-31 (blue, AUC=0.95) and all four miRNAs (red, AUC=0.99).
Discussion
The activity of neutrophil may have a significant impact on the pathophysiology of vascular endothelial cells for PE.34, 35 The supernatants of placental villus cultured from PE patients can significantly increase the expression of CD62L and CD1lb.36 All these studies indicated that the biomarkers might be located in leukocyte. Thus our study mainly focused on finding circulating miRNAs from leukocyte that can distinguish PE from healthy pregnant women.
GIT1, a targeted gene of hsa-miR-122-5p, controls the activity of endothelial nitric-oxide synthase in endothelial cells, and down regulation of GIT1 can induce damages to endothelial cells.33 Previous study37 showed that the changes initiated by endothelial cell injury results in the clinical syndrome of PE. In our study, the upregulation of miR-122-5p in PE may induce the vascular endothelial damage, immune dysfunction and abnormal liver function.
The overactivity of miR-451 might also be responsible for the damage of the patients’ circulatory system and nervous system.38 Previous studies39, 40 showed that miR-451 has a crucial role in promoting erythroid maturation under conditions of stress. Overexpression of miR-451 could significantly inhibit growth and induce cell apoptosis.41, 42 The downregulated miR-451 has been reported in other studies.43, 44 In our study, the upregulated miR-451 might be responsible for placenta hypoxia in the PE group. Meanwhile, the abnormal expression of miR-451a in PA and PC may be complicated. Further study are needed to explore the mechanism of miR-451a in PA and PC.
miR-15a negatively regulate BCL2 at a posttranscriptional level.45 In our study, downregulation of miR-15a-3p might lead to an increased expression of BCL2 in leukocytes and induce normal leukocyte apoptosis by increasing oxidative stress and eventually lead to vascular endothelial injury.
miR-31 is blood vascular endothelial cell-specific posttranscriptional regulatory mechanism that has negative function in the development and function of lymphatic vascular system.46 In our study, downregulation of miR-31-3p may enhance immune response and induce endothelial damage and pathogenesis of PE.
Screening by maternal serum pregnancy-associated plasma protein A, free β-hCG, activin A and inhibin A concentrations at 22+0 to 24+6 weeks’ gestation detect 75% and 92% of patients who subsequently develop pre-eclampsia for false positive rates of 5% and 10%, respectively.47 A multivariate predictive model combining 14 metabolites as metabolomic biomarkers in PE obtained an AUC of 0.94.48 To distinguish PE patients from healthy women, we constructed the prediction model of PE using four miRNAs. Three of them achieved a very high AUC range from 0.9 to 0.96. In our study, the prediction model achieved 0.99 AUC and estimated accuracy of 0.94 when using the combination of these four miRNAs to predict the status of samples. Therefore, the constructed model is fit for predication of PE using these four miRNAs.
In summary, a set of miRNAs were found that significantly differentially expressed in PE samples based on the results of data analysis of next-generation sequencing and qRT-PCR validation. We successfully constructed a logistic regression model to distinguish PE patients from healthy women with extremely high accuracy. We provide information regarding the valuable miRNAs for diagnosing the systematic subtypes of PE and advance our understanding of the pathogenesis of PE.
References
Redman CW, Sargent IL . Latest advances in understanding preeclampsia. Science 2005; 308 (5728): 1592–1594.
Redman CW, Sargent IL . Immunology of pre-eclampsia. Am J Reprod Immunol 2010; 63 (6): 534–543.
Carty DM, Delles C, Dominiczak AF . Novel biomarkers for predicting preeclampsia. Trends Cardiovasc Med 2008; 18 (5): 186–194.
Romero R, Nien JK, Espinoza J, Todem D, Fu W, Chung H et al. A longitudinal study of angiogenic (placental growth factor) and anti-angiogenic (soluble endoglin and soluble vascular endothelial growth factor receptor-1) factors in normal pregnancy and patients destined to develop preeclampsia and deliver a small for gestational age neonate. J Matern Fetal Neonatal Med 2008; 21 (1): 9–23.
Grill S, Rusterholz C, Zanetti-Dallenbach R, Tercanli S, Holzgreve W, Hahn S et al. Potential markers of preeclampsia—a review. Reprod Biol Endocrinol 2009; 7: 70.
Hawfield A, Freedman BI . Pre-eclampsia: the pivotal role of the placenta in its pathophysiology and markers for early detection. Ther Adv Cardiovasc Dis 2009; 3 (1): 65–73.
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA 2008; 105 (30): 10513–10518.
Wang K, Zhang S, Marzolf B, Troisch P, Brightman A, Hu Z et al. Circulating microRNAs, potential biomarkers for drug-induced liver injury. Proc Natl Acad Sci USA 2009; 106 (11): 4402–4407.
Choudhury M, Friedman JE . Epigenetics and microRNAs in preeclampsia. Clin Exp Hypertens 2012; 34 (5): 334–341.
Chen DB, Wang W . Human placental microRNAs and preeclampsia. Biol Reprod 2013; 88 (5): 130.
Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 2008; 18 (10): 997–1006.
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci 2008; 105 (30): 10513–10518.
Wu L, Zhou H, Lin H, Qi J, Zhu C, Gao Z et al. Circulating microRNAs are elevated in plasma from severe preeclamptic pregnancies. Reproduction 2012; 143 (3): 389–397.
Stubert J, Koczan D, Richter DU, Dieterich M, Ziems B, Thiesen HJ et al. miRNA expression profiles determined in maternal sera of patients with HELLP syndrome. Hypertens Pregnancy 2014; 33 (2): 215–235.
Anton L, Olarerin-George AO, Schwartz N, Srinivas S, Bastek J, Hogenesch JB et al. miR-210 inhibits trophoblast invasion and is a serum biomarker for preeclampsia. Am J Pathol 2013; 183 (5): 1437–1445.
Luque A, Farwati A, Crovetto F, Crispi F, Figueras F, Gratacos E et al. Usefulness of circulating microRNAs for the prediction of early preeclampsia at first-trimester of pregnancy. Sci Rep 2014; 4: 4882.
Ward JR, Heath PR, Catto JW, Whyte MK, Milo M, Renshaw SA . Regulation of neutrophil senescence by microRNAs. PLoS One 2011; 6 (1): e15810.
Larsen MT, Hother C, Häger M, Pedersen CC, Theilgaard-Mönch K, Borregaard N et al. MicroRNA profiling in human neutrophils during bone marrow granulopoiesis and in vivo exudation. PLoS One 2013; 8 (3): e58454.
Chang CW, Wu HC, Terry MB, Santella RM . microRNA expression in prospectively collected blood as a potential biomarker of breast cancer risk in the BCFR. Anticancer Res 2015; 35 (7): 3969–3977.
Schroeder BM . ACOG practice bulletin on diagnosing and managing preeclampsia and eclampsia. American College of Obstetricians and Gynecologists. Am Fam Physician 2002; 66 (2): 330–331.
Morin RD, O'Connor MD, Griffith M, Kuchenbauer F, Delaney A, Prabhu AL et al. Applicationof massively parallel sequencing to microRNA profiling and discovery in human embryonic stem cells. Genome Res 2008; 18 (4): 610–621.
Xu G, Wu J, Zhou L, Chen B, Sun Z, Zhao F et al. Characterization of the small RNA transcriptomes of androgen dependent and independent prostate cancer cell line by deep sequencing. PLoS One 2010; 5 (11): e15519.
Li R, Li Y, Kristiansen K, Wang J . SOAP: short oligonucleotide alignment program. Bioinformatics 2008; 24 (5): 713–714.
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ . miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 2006; 34 (Database issue): D140–D144.
Tarazona S, Garcia-Alcalde F, Dopazo J, Ferrer A, Conesa A . Differential expression in RNA-seq: a matter of depth. Genome Res 2011; 21 (12): 2213–2223.
Paraskevopoulou MD, Georgakilas G, Kostoulas N, Vlachos IS, Vergoulis T, Reczko M et al. DIANA-microT web server v5.0: service integration into miRNA functional analysis workflows. Nucleic Acids Res 2013; 41 (Web Server issue): W169–W173.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 2000; 25 (1): 25–29.
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res 2008; 36 (Database issue): D480–D484.
Vlachos IS, Kostoulas N, Vergoulis T, Georgakilas G, Reczko M, Maragkakis M et al. DIANA miRPath v.2.0: investigating the combinatorial effect of microRNAs in pathways. Nucleic Acids Res 2012; 40 (Web Server issue): W498–W504.
Robin X, Turck N, Hainard A, Tiberti N, Lisacek F, Sanchez JC et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 2011; 12: 77.
Zhao G, Zhou X, Chen S, Miao H, Fan H, Wang Z et al. Differential expression of microRNAs in decidua-derived mesenchymal stem cells from patients with pre-eclampsia. J Biomed Sci 2014; 21: 81.
Ishibashi O, Ohkuchi A, Ali MM, Kurashina R, Luo SS, Ishikawa T et al. Hydroxysteroid(17-beta) dehydrogenase 1 is dysregulated by miR-210 and miR-518c that are aberrantly expressed in preeclamptic placentas: a novel marker for predicting preeclampsia. Hypertension 2012; 59 (2): 265–273.
Mouillet JF, Chu T, Nelson DM, Mishima T, Sadovsky Y . MiR-205 silences MED1 in hypoxic primary human trophoblasts. FASEB J 2010; 24 (6): 2030–2039.
Liu S, Premont RT, Rockey DC . G-protein-coupled receptor kinase interactor-1 (GIT1) is a new endothelial nitric-oxide synthase (eNOS) interactor with functional effects on vascular homeostasis. J Biol Chem 2012; 287 (15): 12309–12320.
Belo L, Santos-Silva A, Caslake M, Cooney J, Pereira-Leite L, Quintanilha A et al. Neutrophil activation and C-reactive protein concentration in preeclampsia. Hypertens Pregnancy 2003; 22 (2): 129–141.
Cadden KA, Walsh SW . Neutrophils, but not lymphocytes or monocytes, infiltrate maternal systemic vasculature in women withpreeclampsia. Hypertens Pregnancy 2008; 27 (4): 396–405.
Wang Y, Gu Y, Philibert L, Lucas MJ . Neutrophil activation induced by placental factors in normal and pre-eclamptic pregnancies in vitro. Placenta 2001; 22 (6): 560–565.
Roberts JM, Taylor RN, Musci TJ, Rodgers GM, Hubel CA, McLaughlin MK . Preeclampsia: an endothelial cell disorder. Am J Obstet Gynecol 1989; 161 (5): 1200–1204.
Godlewski J, Nowicki MO, Bronisz A, Nuovo G, Palatini J, De Lay M et al. MicroRNA-451 regulates LKB1/AMPK signaling and allows adaptation to metabolic stress in glioma cells. Mol Cell 2010; 37 (5): 620–632.
Rasmussen KD, Simmini S, Abreu-Goodger C, Bartonicek N, Di Giacomo M, Bilbao-Cortes D et al. The miR-144/451 locus is required for erythroid homeostasis. J Exp Med 2010; 207 (7): 1351–1358.
Pase L, Layton JE, Kloosterman WP, Carradice D, Waterhouse PM, Lieschke GJ . miR-451 regulates zebrafish erythroid maturation in vivo via its target gata2. Blood 2009; 113 (8): 1794–1804.
Wu LN, Wei XW, Fan Y, Miao JN, Wang LL, Zhang Y et al. Altered expression of 14-3-3ζ protein in spinal cords of rat fetuses with spina bifida aperta. PLoS One 2013; 8 (8): e70457.
Bian HB, Pan X, Yang JS, Wang ZX, De W . Upregulation of microRNA-451 increases cisplatin sensitivity of non-small cell lung cancer cell line (A549). J Exp Clin Cancer Res 2011; 30 (1): 20.
Murata K, Yoshitomi H, Furu M, Ishikawa M, Shibuya H, Ito H et al. MicroRNA-451 down-regulates neutrophil chemotaxis via p38 MAPK. Arthritis Rheumatol 2014; 66 (3): 549–59.
Zhang F, Huang W, Sheng M, Liu T . MiR-451 inhibits cell growth and invasion by targeting CXCL16 and is associated with prognosis of osteosarcoma patients. Tumour Biol 2015; 36 (3): 2041–2048.
Cimmino A, Calin GA, Fabbri M, Iorio MV, Ferracin M, Shimizu M et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA 2005; 102 (39): 13944–13949.
Pedrioli DM, Karpanen T, Dabouras V, Jurisic G, van de Hoek G, Shin JW et al. miR-31 functions as a negative regulator of lymphatic vascular lineage-specific differentiation in vitro and vascular development in vivo. Mol Cell Biol 2010; 30 (14): 3620–3634.
Spencer K, Yu CK, Savvidou M, Papageorghiou AT, Nicolaides KH . Prediction of pre-eclampsia by uterine artery Doppler ultrasonography and maternal serum pregnancy-associated plasma protein-A, free beta-human chorionic gonadotropin, activin A and inhibin A at 22+0 to 24+6 weeks' gestation. Ultrasound Obstet Gynecol 2006; 27 (6): 658–663.
Kenny LC, Broadhurst DI, Dunn W, Brown M, North RA, McCowan L et al. Robust early pregnancy prediction of later preeclampsia using metabolomic biomarkers. Hypertension 2010; 56 (4): 741–749.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Journal of Perinatology website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Wang, Y., Yang, X., Yang, Y. et al. High-throughput deep screening and identification of four peripheral leucocyte microRNAs as novel potential combination biomarkers for preeclampsia. J Perinatol 36, 263–267 (2016). https://doi.org/10.1038/jp.2015.192
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jp.2015.192