Abstract
Glycogen storage disease type III (GSD III), a rare autosomal recessive disease characterized by hepatomegaly, fasting hypoglycemia, growth retardation, progressive myopathy and cardiomyopathy, is caused by deficiency of the glycogen debranching enzyme (AGL). Direct sequencing of human AGL cDNA and genomic DNA has enabled analysis of the underlying genetic defects responsible for GSD III. To date, the frequent mutations in different areas and populations have been described in Italy, Japan, Faroe Islands and Mediterranean area, whereas little has been performed in Chinese population. Here we report a sequencing-based mutation analysis in 43 Chinese patients with GSD III from 41 families. We identified 51 different mutations, including 15 splice-site (29.4%), 11 small deletions (21.6%), 12 nonsense (23.5%), 7 missense (13.7%), 5 duplication (9.8%) and 1 complex deletion/insertion (2.0%), 31 of which are novel mutations. The most common mutation is c.1735+1G>T (11.5%). The association of AGL missense and small in-frame deletion mutations with normal creatine kinase level was observed. Our study extends the spectrum of AGL mutations and suggests a genotype–phenotype correlation in GSD III.
Similar content being viewed by others
Introduction
Glycogen storage disease type III (GSD III; MIM #232400) is a rare genetic disorder of glycogen metabolism inherited as an autosomal recessive trait. It is caused by a deficiency of the glycogen debranching enzyme (AGL), which has two independent catalytic activities as follows: 4-alpha-glucanotransferase (EC 2.4.1.25) and amylo-1,6-glucosidase (EC 3.2.1.33). GSD III can be classified into four types (a, b, c and d) according to the different affected organs and loss of different catalytic activities of the glycogen debranching enzyme.1 GSD IIIa is the most common type that accounts ~80% of all GSD III and differentiated with other types by both liver and muscle involved. All types of GSD III are caused by various mutations of the AGL gene (MIM #610860). Human AGL gene is located on chromosome 1p21 and consists of 35 exons spanning ~85 kb of genomic DNA. The AGL gene encodes about 7 kb mRNA and produces at least six isoforms by alternative splicing.2 The major mRNA isoform present in both muscle and liver encodes a protein consisting of 1532 amino-acid residues,2, 3 and is involved in the development of GSD IIIa. Clinical manifestations of GSD III include hepatomegaly, fasting hypoglycemia, growth retardation, and, in many patients, progressive myopathy and cardiomyopathy.4 Frequently, hepatomegaly tends to resolve spontaneously. In patients with GSD IIIa, cardiomyopathy may become predominant in adults.
There are about 170 different disease-causing AGL mutations recorded in the Human Gene Mutation Database (http://www.hgmd.org) and other references. The majorities are nonsense, deletion, insertion and splicing mutations.5 The prevalent mutations of AGL in GSD III vary among the ethnic groups. The most frequent mutation among the Italian patients is a splice-site mutation (c.2681+1G>A) and it accounts for 20.5% of 42 patients;6 the prevalent mutation in Faroe Islands is a nonsense mutation (p.R408X) caused by the founder effect;7 the frequent mutations in the United States are p.R864X (10.3%), c.3964delT (6.7%), c.4260-12 A>G (5.5%) and p.R1228X (5.2%), which together account for 28% of 29 patients.5 To date, 37 AGL mutations have been reported in Chinese patients or patients of Chinese descent, including 12 nonsense, 11 small deletions, 9 splicing, 4 insertions and 1 missense mutation.8, 9, 10, 11, 12, 13
In order to get a better understanding of the molecular basis of the AGL gene in Chinese GSD III patients, we studied 43 patients from 41 families by direct sequencing. A spectrum of 51 AGL mutations, including 31 novel mutations, was identified. A total of 41 genotypes were characterized. A genotype–phenotype correlation was observed.
Materials and methods
Patients
A total of 43 Chinese GSD III patients, including 28 males and 13 females, aged between 1 and 21 years, from 41 unrelated families were collected from the Clinic of Pediatric Genetics at the Peking Union Medical College Hospital. All of them were Han Chinese. Patients 28 and 29 were monozygous twin daughters of a consanguineous couple, and patients 24 and 33 were siblings in a family. The clinical information was not available in patients 42 and 43. All patients were presented with hepatomegaly and fasting hypoglycemia. Biochemical studies showed increased aspartate aminotransferase (AST) and variable creatine kinase (CK). Biochemical assays (AST, CK, triglyceride (TG)) were performed by automatic biochemical analyzer (Beckman Coulter AU5800, Brea, CA, USA). Glucose response to epinephrine stimulation was flat after overnight fasting, but a significant elevation (blood glucose increased ⩾2.5 mmol l−1) post prandial. None of our patients was included in the study by Zhuang et al.10 and Wang et al.12 The study was approved by the Peking Union Medical College Hospital Institutional Review Board, and peripheral blood samples were collected from the patients and their family members with informed consent.
Clinical information
Except for patients 42 and 43, whose clinical data were not available, the other 41 were long-term follow-up patients in our clinic and were managed by corn starch supplement only. The age of disease onset, sex, physical examination, biochemistry data from recent clinical visit were shown in Supplementary Table S1. The duration of patient follow-up in our clinic varied from 1 year to 12 years. The age at symptom onset was between 3 months and 3 years old, and the age at last visiting was from 1 year to 27 years old. AST was increased in all patients. The TG was within normal range in nine cases. Of 32 cases with increased TG, 26 had the level lower than 4 mmol l−1.
Mutation analysis
Genomic DNA of patients and their family members was isolated from peripheral blood leukocytes using a TAKARA Universal Genomic DNA Extraction kit (Takara, Otsu, Japan). Total RNA was extracted from fresh whole blood by QIAamp RNA Blood Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. Messenger RNAs were reverse-transcribed by PrimeScript One Step RT-PCR Kit (Takara). The full coding exons, their relevant exon–intron boundaries, the 5′- and 3′-flanking regions of the AGL gene and the full cDNA were amplified using primers designed according to the published sequence of AGL isoform 1 (GenBank accession no. NM_000642). Primers and conditions are available upon request. After purification, the polymerase chain reaction (PCR) products were subjected to automatic DNA sequencing and the result sequences were compared with the normal control. All mutations found were confirmed by sequencing of a second PCR amplicon. As reference, the A of the ATG translation initiation codon of coding sequence of AGL is referred to as nucleotide +1. Mutation description follows the standard nomenclature (http://www.hgvs.org/mutnomen/). To confirm the pathogenicity of the novel missense mutations, restriction fragment length polymorphism experiments were conducted with 50 normal Han Chinese controls. PCR primers and restriction endonucleases using restriction fragment length polymorphism detection for five novel missense mutations of the AGL gene are showed in Supplementary Table S2. Restriction digests were analyzed on 8% polyacrylamide gel or 2% agarose gel.
Bioinformatic protein analysis
ClustalW2 (http://www.ebi.ac.uk/Tools/clustalw2/index.html) from EMBL-EBI and BLAT (http://genome.ucsc.edu/cgi-bin/hgBlat?hgsid=208883171) from UCSC were used to examine evolutionary conservation of the amino-acid residues in these variants by multiple sequence alignment.
PolyPhen-2 program (http://genetics.bwh.harvard.edu/pph2/dokuwiki/start) was used to evaluate possible biologic effects of the amino-acid substitution on the structure and function of the AGL protein.
Statistical analysis
The patients under 10 years old were classified into two groups according to the serum CK value. Group 1 stands for serum CK value >198 IU l−1, and group 2 stands for CK value ⩽198 IU l−1 (normal CK level: 0
Results
Mutational spectrum and frequency of AGL in GSD III patients
By direct Sanger sequencing, we identified 51 mutations in AGL (Figure 1) in patients from the 41 study families. For two patients (12 and 25), only one mutation could be identified, making the mutation detection rate up to 97.56% (80/82) in our present study. Those mutations are comprised of 15 splice-site (29.4%), 11 small deletions (21.6%), 12 nonsense (23.5%), 7 missense (13.7%), 5 duplication (9.8%) and 1 complex deletion/insertion (2.0%) mutations. Two splice-site mutations, c.1735+1G>T and c.665-1G>A, were detected in 9 (11.1%) and 4 (4.9%) independent alleles, respectively. The rest 49 mutations were each identified in less than four alleles (Supplementary Table S1). Of the 51 mutations, 31 were novel. The majority (25/31, 80.6%) of the novel mutations identified here were predicted to result in premature termination codons (PTCs). These 25 mutations comprised 10 frameshift, nine splicing and six nonsense. All these 51 mutations are not present in the NCBI SNP database Build 142, the 1000 genomes browser (http://www.ncbi.nlm.nih.gov/variation/tools/1000genomes/), and NIH Heart, Lung and Blood Institute exome sequencing project exome variant server (http://evs.gs.washington.edu/EVS/).
AGL mutational spectrum in Chinese GSD III patients. Schematic representation of the AGL gene comprising 35 exons. Exons are not drown in scale. The mutations identified in this study are depicted in blue, and known mutations previously reported by the literature (Chinese population) are in black. Mutation marked in red stands for two different nucleotide changes result in same nonsense mutation. Blue boxes stand for exons considered encoding the putative transferase catalytic residues. Green boxes stand for exons considered encoding the putative glucosidase catalytic residues and orange ones stand for exons considered encoding putative glycogen-binding domain.
RT-PCR analysis of splice-site mutations
Four splice-site mutations, including three novel ones and our previously reported c.2546+1G>A, were further analyzed by RT-PCR assays on available RNA samples to verify the transcriptional effect (Qiu et al.,13 Figure 2). Three of them brought to a skipping of one exon and resulted in translational frameshifts. The c.2546+1G>A, c.3588+1G>A and c.4347+1G>A mutations led to a skipping of exon 20 (113 bp), exon 27 (226 bp) and exon 33 (88 bp), respectively. And one acceptor splice-site mutation, c.1424-1G>C, resulted in an in-frame deletion encompassing exons 13 and 14 (312 bp).
Consequences of four splice-site mutations on AGL pre-mRNA splicing. Genomic DNA and cDNA sequencing demonstrating the heterozygous c.1424-1G>C, c.2546+1G>A, c.3588+1G>A, c.4347+1G>A splicing mutations in AGL and showing the aberrant splicing with variable skipping of adjacent exons in cDNA templates.
Bioinformatics analysis of missense mutations
Five missense variants, c.499G>C (p.G167R), c.713 T>G (p.L238R), c.742 T>A (p.W248R), c.1513C>T (p.L505F), c.3299G>A (p.G1100E), were considered to be likely pathogenic after bioinformatics analysis. PolyPhen-2 classified all five mutations as ‘probably damaging’ with scores from 0.996 to 1. These scores indicate the mutations are predicted to affect protein function or structure with a high degree of confidence. Determination of the evolutionary conservation of p.G167R, p.L238R, p.W248R, p.L505F and p.G1100E by multiple amino-acid sequence alignment of human (Homo Sapiens) AGL with the protein sequences derived from mouse (Mus musculus), rat (Rattus norvegicus), chimpanzee(Pan troglodytes), rabbit (Oryctolagus cuniculus), horse (Equus callabus), dog (Canis lupus familiaris), cow (Bos Taurus), chicken (Gallus gallus) and zebra fish (Danio rerio) showed that all residues 167, 238, 248, 505 and 1100 in AGL were well conserved across deferent species (Figure 3).
Multiple amino-acid sequence alignment of AGL among human and other species in the region surrounding mutated residues. Sequences of human (Homo sapiens NP_000019), chimpanzee (Pan troglodytes XP_524777), cow (Bos taurus XP_595566), horse (Equus callabus NP_001103778), rabbit (Oryctolagus cuniculus NP_001075716), dog (Canis familiaris NP_001041561), mouse (Mus musculus NP_001074795), rat (Rattus norvegicus NP_001102034), chicken (Gallus gallus XP_422317) and zebra fish (Danio rerio XP_696194). A full color version of this figure is available at the Journal of Human Genetics journal online.
Genotypes of Chinese GSD III patients
The characterization of the genotypes about the 41 families is shown in Supplementary Table S1. Compound heterozygous mutations were found in 33 families, homozygous mutations in six families. Patients 4 and 40 shared identical mutations, c.1298_1299delCA (p.P433LfsX2) and c.1937_1939delTTG (p.V647del). Patient 9 and the affected brothers (patients 24 and 33) shared another compound heterozygous mutations, c.665-1G>A and c.1571G>A (p.R524H). Mutation analysis in the 62 parental samples available for sequencing showed that all the mutations were inherited from their carrier parents.
Genotype–phenotype correlation
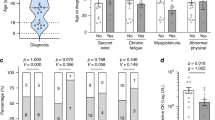
In our present study, we observed a possible association between the age and the serum CK level. All six patients equal to or older than 10 years old (⩾10y) had a higher serum CK value than normal. However, 24 (68.6%) of the 35 patients under 10 years of age (<10y), had elevated CK. We then examined the genotype and the serum CK level in the patient group of <10y. Nine of the 11 patients, who carried missense mutations or small in-frame deletion, showed a normal CK level, whereas 22 of the 24 patients with PTC mutations had an elevated CK, suggesting the presence of a genotype–phenotype correlation in this patient group.
Discussion
Mutational spectrum
In our present study, a total of 51 AGL mutations on 81 independent mutant alleles had been identified in 43 Chinese patients with GSD IIIa. Review of the literature indicated that 37 mutations had been reported in Chinese population. Except for the c.1735+1G>T, we found the recurrence of 13 previously reported mutations in Chinese, including c.665-1G>A on four alleles, c.100C>T (p.R34X) and c.4234delC (p.G1413AfsX2) on three alleles, c.958+1G>T, c.2929C>T (p.R977X) and c.4284T>G (p.Y1428X) on two alleles each, c.664+1G>A, c.1177C>T (p.Q393X), c.2520_2524dupGTCTC (p.P842RfsX28), c.2546+1G>A, c.2717_2721delAGATC (p.Q906PfsX5), c.2906_2907delAT (p.Y969CfsX3) and c.4260-12A>G on one allele. Our data expanded the spectrum of AGL mutations in Chinese patients from 37 to 72. These mutations scattered throughout the entire AGL gene (shown in Figure 1) and no mutational hot spot region was found.
The frequent mutations detected in our GSD III cohort were c.1735+1G>T (11.1%) and c.665-1G>A (4.9%), which together account for 16.6% of 81 independent alleles. Our result was in agreement with Wang et al.11 and Wang et al.12 that c.1735+1G>T is the most common mutation in Chinese GSD III patients. The c.1735+1G>T mutation was first reported in a Japanese patient.14 This donor splice-site mutation could cause a skipping of exon 14 (1612–1735th nucleotides of the AGL cDNA), which resulted in a translational frameshift. This mutation was also reported with a highest frequency of 11.8% in Korean and Japanese patients.15, 16 On the basis of the higher frequency of this mutation found in Japanese, Korean and Chinese patients, we concluded that c.1735+1G>T might be the most prevalent mutation in Asians.
The nucleotide substitutions at the highly conserved GT–AG splice sites were relatively common in our study cohort. The functional consequence of such mutations is difficult to predict without analyzing mRNA/cDNA. Of the 15 splice-site mutations found in our study, 9 were novel. We investigated the effect of four mutations with available samples using RT-PCR. Skipping of a single exon was confirmed for mutations c.2546+1G>A, c.3588+1G>A and c.4347+1G>A, whereas skipping deletion of two exons was detected for the c.1424-1G>C mutation. It is noteworthy that a novel mutation, c.1735+1G>A, located at the same nucleotide position as the frequently detected c.1735+1G>T, was also identified. As c.1735+1G>T had been proved to cause exon 14 skipping, we predicted that c.1735+1G>A might have similar effect on the RNA product.
All the five novel missense mutations, p.G167R, p.L238R, p.W248R, p.L505F and p.G1100E, changed the highly conserved amino acids (Figure 3). As shown in Figure 1, p.G167R, p.L505R and p.G1100E, were located in exons 6, 13 and 26 of AGL, respectively. Exon 6 and exon 13 were considered encoding the putative transferase catalytic residues, and exon 26 was considered encoding the putative glucosidase catalytic residues of the AGL enzyme. Therefore, these mutations were likely to influence the activities of AGL.5 The p.W248R mutation was identified on three mutant alleles. One patient (patient 1) was homozygous and the other (patient 39) was compound heterozygous with this mutation. The p.L238R mutation was found to be compound heterozygous in patient 34 with another mutation c.4481+2T>A. No other variants in AGL were found in the above-mentioned six patients except common single nucleotide polymorphisms (⩾1%). In addition, none of these five missense mutations was detected in 50 normal controls (100 chromosomes) nor was present in public databases, suggesting that they are pathogenic mutations.
The novel three base-pair deletion, c.3878_3880delCTG (p.A1293del), was identified in one compound heterozygous patient (patient 6). This small in-frame deletion in exon 30 affected an amino that is not highly conserved among human and other species (data not shown). The pathogenic effect of this deletion was uncertain, even though the patient had a typical presentation at the age of 2.
Genotype–phenotype correlation
GSD III has a poor correlation between the genotype and phenotype.17 However, the homozygous p.Y1510X nonsense mutation in the 3′-coding region of AGL had been suggested to be associated with a severe phenotype in GSD III;18 and the homozygous c.4260-12A>G splice-site mutation was reported in multiple patients and was associated with mild clinical symptoms.19
The involvement of skeletal muscle may differentiate GSD IIIa from GSD IIIb. The increased serum CK is a suggestive and nonspecific marker of muscle breakdown, although the correlation between the level of CK and the severity of muscle involvement was not known yet. In our present study, we found significant association between PTC mutations and elevated serum CK value (Table 1). And most of the missense and the small in-frame deletion mutations in AGL were found in GSD III patients with normal serum CK levels. This may imply relatively less or late muscle involvement.
In summary, we have identified 31 novel mutations and extended the mutation spectrum of AGL in Chinese patients with GSD III. In addition, our results suggested a correlation between the missense and small in-frame deletion mutations and a normal serum CK level in GSD III.
References
Ding, J. H., de Barsy, T., Brown, B. I., Coleman, R. A. & Chen, Y. T. Immunoblot analyses of glycogen debranching enzyme in different subtypes of glycogen storage disease type III. J. Pediatr. 116, 95–100 (1990).
Yang-Feng, T. L., Zheng, K., Yu, J., Yang, B. Z., Chen, Y. T. & Kao, F. T. Assignment of the human glycogen debrancher gene to chromosome 1p21. Genomics 13, 931–934 (1992).
Bao, Y., Yang, B. Z. & Chen, Y. T. Structural organization of the multifunctional human glycogen debrancher gene. Am. J. Hum. Genet. 53, 662A (1993).
Chen, Y.T., Burchell, A. in The Metabolic and Molecular Bases of Inherited Disease (eds Scriver, C. R., Beaudet, A. L., Sly, W. S. & Valle, D.) 935–965 (McGraw-Hill, New York, 1995).
Shen, J. J. & Chen, Y. T. Molecular characterization of glycogen storage disease type III. Curr. Mol. Med. 2, 167–175 (2002).
Lucchiari, S., Donati, M. A., Parini, R., Melis, D., Gatti, R., Bresolin, N. et al. Molecular characterisation of GSD III subjects and identification of six novel mutations in AGL. Hum. Mutat. 20, 480 (2002).
Santer, R., Kinner, M., Steuerwald, U., Kjaergaard, S., Skovby, F., Simonsen, H. et al. Molecular genetic basis and prevalence of glycogen storage disease type IIIA in the Faroe Islands. Eur. J. Hum. Genet. 9, 388–391 (2001).
Horinishi, A., Okubo, M., Tang, N. L., Hui, J., To, K. F., Mabuchi, T. et al. Mutational and haplotype analysis of AGL in patients with glycogen storage disease type III. J. Hum. Genet. 47, 55–59 (2002).
Lam, C. W., Lee, A. T., Lam, Y. Y., Wong, T. W., Mak, T. W., Fung, W. C. et al. DNA-based subtyping of glycogen storage disease type III: mutation and haplotype analysis of the AGL gene in Chinese. Mol. Genet. Metab. 83, 271–275 (2004).
Zhuang, T. F., Qiu, Z. Q., Wei, M. & Huang, S. Z. Mutation analysis of glycogen debrancher enzyme gene in five Chinese patients with glycogen storage disease type III. Chin. J. Pediatr. 43, 85–88 (2005).
Wang, X., Qiu, W. J., Ye, J., Han, L. S., Zhang, H. W., Jiang, L. R. et al. Molecular genetic analysis of 10 Chinese patients with glycogen storage disease type III. Zhonghua Er Ke Za Zhi. 47, 416–420 (2009).
Wang, W., Wei, M., Song, H. M., Qiu, Z. Q., Zhang, W. M., Wu, X. Y. et al. Diagnosis of glycogen storage disease type IIIa by detecting glycogen debranching enzyme activity, glycogen content and structure in muscle. Chin. J. Pediatr. 47, 608–612 (2009).
Qiu, Z. Q., Wang, W., Wei, M., Qiu, J. J., Zhang, H. B. & Wu, X. Y. Mutation analysis of AGL gene in 7 Chinese patients with glycogen storage disease type III. Zhonghua Er Ke Za Zhi. 31, 471–474 (2011).
Okubo, M., Aoyama, Y. & Murase, T. A novel donor splice site mutation in the glycogen debranching enzyme gene is associated with glycogen storage disease type III. Biochem. Biophys. Res. Commun. 224, 493–499 (1996).
Kim, K. O., Lee, H. J., Choi, J. W., Eun, J. R. & Choi, J. H. An adult case of glycogen storage disease type IIIa. Korean J. Hepatol. 14, 219–225 (2008).
Oh, S. H., Park, H. D., Ki, C. S., Choe, Y. H. & Lee, S. Y. Biochemical and molecular investigation of two Korean patients with glycogen storage disease type III. Clin. Chem. Lab. Med. 46, 1245–1249 (2008).
Lucchiari, S., Fogh, I., Prelle, A., Parini, R., Bresolin, N., Melis, D. et al. Clinical and genetic variability of glycogen storage disease type IIIa: seven novel AGL gene mutations in the Mediterranean area. Am. J. Med. Genet. 109, 183–190 (2002).
Shen, J. J., Bao, Y. & Chen, Y. T. A nonsense mutation due to a single base insertion in the 3-prime-coding region of glycogen debranching enzyme gene associated with a severe phenotype in a patient with glycogen storage disease type IIIa. Hum. Mutat. 9, 37–40 (1997).
Shaiu, W. L., Kishnani, P. S., Shen, J., Liu, H. M. & Chen, Y. T. Genotype-phenotype correlation in two frequent mutations and mutation update in type III glycogen storage disease. Mol. Genet. Metab. 69, 16–23 (2000).
Acknowledgements
We thank all patients and their family members who participated in this study. This work was funded by the National 863 Program of China (2007AA02Z440), the National Science & Technology Pillar Program during the Twelfth Five-year Plan Period (2012BAI09B00), the National Natural Science Foundation of China (81241032 and 30871355), the Public Science and Technology Research Funds Projects of Health (201302001), and the Presidential Foundation of Chinese Academy of Medical Sciences & Peking Union Medical College.
Author contributions
ZQ, MS and XZ conceived and designed the study. CL carried out the experiments and analyzed the gene mutation results. WM, ZQ and WW performed the study of the clinical part. CL, ZQ, MS and XZ wrote the manuscript. All authors revised and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Lu, C., Qiu, Z., Sun, M. et al. Spectrum of AGL mutations in Chinese patients with glycogen storage disease type III: identification of 31 novel mutations. J Hum Genet 61, 641–645 (2016). https://doi.org/10.1038/jhg.2016.24
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2016.24
This article is cited by
-
The biallelic novel pathogenic variants in AGL gene in a chinese patient with glycogen storage disease type III
BMC Pediatrics (2022)
-
A novel homozygous splicing mutation of the AGL gene in a Chinese patient with severe myopathy involvement of glycogen storage disease type IIIa
Neurological Sciences (2021)
-
Hepatic Manifestations in Glycogen Storage Disease Type III
Current Pathobiology Reports (2018)