Abstract
Tafazzin, encoded by the TAZ gene, is a mitochondrial membrane-associated protein that remodels cardiolipin (CL), an important mitochondrial phospholipid. TAZ mutations are associated with Barth syndrome (BTHS). BTHS is an X-linked multisystemic disorder affecting usually male patients. Through sequence analysis of TAZ, we found one novel mutation c.39_60del p.(Pro14Alafs*19) by whole-exome sequencing and a reported missense mutation c.280C>T p.(Arg94Cys) by Sanger sequencing in two male patients (Pt1 and Pt2). Patient with c.280C>T mutation had dilated cardiomyopathy, while another patient with c.39_60del mutation had no feature of cardiomyopathy. A reported m.1555A>G homoplasmic variant was also identified in the patient having mutation c.39_60del by whole mitochondrial DNA sequencing method. This variant was not considered to be the main cause of mitochondrial dysfunction based on a cytoplasmic hybrid (cybrid) assay. Tafazzin expression was absent in both patient-derived fibroblast cells. Complementation of TAZ expression in fibroblasts from the patient with the novel mutation c.39_60del restored mitochondrial respiratory complex assembly. High-performance liquid chromatography–tandem mass spectrometry-based metabolic analysis revealed the decline of CL and the accumulation of monolysocardiolipin, indicating the loss of tafazzin activity. Owing to phenotypic variability, it is difficult to diagnose BTHS based on clinical features only. We conclude that genetic analysis should be performed to avoid underdiagnosis of this potentially life-threatening inborn error of metabolism.
Similar content being viewed by others
Introduction
Tafazzin, encoded by the nuclear gene TAZ, is primarily found in the inner mitochondrial membrane homology to acyl transferases.1 TAZ is located on chromosome Xq28 and comprises 11 exons and 2 alternative translation initiation sites.2 Among the four major transcripts: TAZ-FL (full length), TAZ-Δ5 (lacking exon 5), TAZ-Δ7 (lacking exon 7), and TAZ-Δ5;7 (lacking exons 5 and 7), TAZ-Δ5 is the most common.3 Mutations in TAZ may be found in each of its exons and introns. The mutations include single-nucleotide substitutions, insertions, partial gene deletions, large deletions of several exons, partial gene duplications, splice site mutations4 and deep intronic mutation with a new 5′ splice site.5 Ferri et al.6 identified a novel synonymous substitution, which is located in exon 4, and analysis of mRNA revealed aberrant skipping of 24 nucleotides in the mature tafazzin mRNA, leading to the deletion of eight amino acids in the tafazzin protein. Therefore, gene expression can be damaged by apparently harmless silent variants in the TAZ gene.6
Mutations in TAZ are associated with Barth syndrome (BTHS), an X-linked multisystem disorder exclusively affecting male patients, characterized by cardiomyopathy, skeletal myopathy, growth retardation, neutropenia, characteristic dysmorphism, male fetal death resulting in miscarriage and stillbirth.7 The mitochondria of BTHS patients have abnormal ultrastructure and have several respiratory chain complex deficiencies in muscle and fibroblast cells.8
There is lack of established genotype–phenotype correlations in BTHS9; patients with the same mutation may manifest extremely diverse phenotypes, even within the same family,10 varying age at clinical presentation11 and no correlation with cardiac phenotype or disease severity.12 Patients who inherited de novo mutation owing to maternal gonadal mosaicism, not to somatic mutation, has been reported.13 Clinical phenotypes of BTHS were confirmed with deletions of exons 1–5 in the TAZ gene in a female infant and mosaicism for monosomy X and a ring X chromosome with deletion of the Xq28 region were identified by cytogenetic analysis.14 A somatic and gonadal mosaicism has been described in a female carrier.15 Based on high variability in the clinical presentations of BTHS, 9% of BTHS patients never exhibit cardiomyopathy16 and about 10% do not exhibit neutropenia.17 Bowron et al.10 described two asymptomatic adults, one of whom never experienced any features of BTHS.10 Sequence analysis of TAZ gene mutation revealed deletions of exons 6–11 in two patients on the basis of exons; even though originally they possess different genetic rearrangements of TAZ, both patients had dilated cardiomyopathy and died owing to heart failure.18
Cardiolipin (CL) is a phospholipid dimer with four fatty acids. It represents 10–15% of total phospholipids in mitochondria and predominantly resides in the inner mitochondrial membrane and to a lesser extent in the outer mitochondrial membrane.19 CL is synthesized from its precursor phosphatidylglycerol, a common substrate in triacylglycerol and glycerolipid metabolism. After synthesis of primary CL, its final mature composition is achieved through remodeling of its acyl chains. The mature form of CL is essential for proper mitochondrial functioning.20 In patients with a TAZ mutation, acyl chain remodeling is defective, which leads to significant changes in CL acyl chain composition and loss of molecular symmetry.21 The mature tissue-specific acyl pattern of CL is critical for normal mitochondrial physiology. In mitochondria, CL helps to maintain mitochondrial ultrastructure and stabilizes the assembly of respiratory chain complexes.22 Only in the presence of CL, the respiratory complexes are able to form stable oligomeric supercomplexes.23
BTHS is characterized by reduced levels of CL and reduced linoleic acid incorporation into CL owing to defective acyl chain remodeling,1 as well as accumulation of monolysocardiolipin (MLCL, CL lacking one acyl chain), and a highly abnormal MLCL/CL ratio.24
In this study, we present the clinical, biochemical and molecular characteristics of two patients from two unrelated families diagnosed with BTHS. We focus specifically on the novel TAZ mutation, which is not associated with characteristic signs and symptoms of BTHS such as cardiomyopathy, neutropenia or 3-methylglutaconic aciduria to shed light on the atypical clinical manifestations and to avoid underdiagnosis of BTHS.
Materials and methods
Patient information
Skin fibroblasts and blood spots from the two patients were examined after obtaining informed written consent from their parents, in accordance with the ethical committee recommendations of Saitama Medical University.
Patient summaries are shown in Table 1. Patient 1 (Pt1) is a 6-year-old boy born to a Japanese family after full-term pregnancy. His weight at birth was 2.6 kg and Apgar score: 3–8–9 (1′–3′–5′). At birth, he exhibited respiratory distress and was hospitalized in the Neonatal Intensive Care Unit for approximately 1 month. Automated auditory brainstem response and brain magnetic resonance imaging showed no pathology when he left the hospital. Developmental delay was noted at 2 years of age: height (−4.0 s.d.), weight (−2.8 s.d.), and head circumference (−1.8 s.d.). Gradually he developed hypotonia, facial dysmorphism, depressed nasal root, prominent forehead and speech difficulty. His echocardiography revealed normal left ventricular (wall) motion and atrial septal defect (II), and he had no history of taking any drugs for cardiac problems. He underwent surgical closure of atrial septal defect. Repeated echocardiography confirmed the absence of cardiomyopathy. He had a history of slightly elevated lactate and pyruvate levels but no history of neutropenia. His mother had been treated with antipsychotic drugs during pregnancy for schizophrenia. Investigation of mitochondrial function from skin fibroblasts revealed reduced mitochondrial complex I (CI) and complex IV (CIV) activities. We confirmed the diagnosis by identification of TAZ mutation c.39_60del p.(Pro14Alafs*19) and MLCL/CL analysis. The ratio of MLCL/CL was 9.30 on dried blood spot (reference range 0–0.3). By whole mitochondrial DNA (mtDNA) sequencing, we identified deafness-associated m.1555A>G variant in mitochondrial 12S rRNA (MT-RNR1) in Pt1. mtDNA m.1555A>G variant was identified in his mother. This variant was homoplasmic in Pt1 and in his mother. We had validated this variant by direct sequencing and PCR–RFLP (restriction fragment length polymorphisms) analysis. Neither Pt1 nor his mother exhibited the phenotype of deafness and had no history of aminoglycoside antibiotic exposure.
Patient 2 (Pt2) is a 9-year-old boy born to a Japanese family after full-term pregnancy. His birth weight was 2.5 kg and Apgar: 8–9 (1′–5′). Shortly after birth, he developed respiratory distress. BTHS was suspected when dilated cardiomyopathy, neutropenia and hypotonia were present and was confirmed by identification of the TAZ mutation c.280C>T p.(Arg94Cys) in exon 3. His mother was not a carrier for TAZ gene mutation. Biochemical analysis also revealed lactic acidosis, and using skin fibroblasts, mitochondrial CI deficiency was detected. The ratio of MLCL/CL on dried blood spot was 6.10 (reference range 0–0.3).
Whole-exome sequencing and Sanger sequence validation
Whole-exome sequencing was performed as previously reported.25 Sanger sequencing was performed on cDNA extracted from patients’ fibroblasts using ABI 3130XL and BigDye v3.1 Terminators (Applied Biosystems, Foster City, CA, USA) per the manufacturer’s protocols. Sequencing primers were as follows: (chrX: 153640216–153640449) TAZ_F_5′-GCTCCCCAGTGACGAGAGA-3′ and TAZ_R_5′-CTCGTACAGCACCTCCCTGTT-3′.
Whole mtDNA sequencing
To avoid contamination of mitochondrial-origin nuclear genome sequences,26 a long-range mtDNA PCR amplification followed by sequencing was performed in this study. DNA was isolated from the patient’s skin fibroblast cells. Long-range PCR was performed using amplicon 1 (rCRS 619–8988) and amplicon 2 (rCRS 8749–895) primers: 5′-GACGGGCTCACATCACCCCATAA-3′ and 5′-GCGTACGGCCAGGGCTATTGGT-3′ for amplicon 1, and 5′-GCCACAACTAACCTCCTCGGGCTCCT-3′ and 5′-GGTGGCTGGCACGAAATTGACC-3′ for amplicon 2. According to the manufacturer’s protocol, PCR fragment libraries were prepared from patient mtDNA using the Nextera XT DNA Sample Prep Kit (Illumina, San Diego, CA, USA). Sequencing was performed as previously described.25
PCR–RFLP and direct sequencing for m.1555A>G variant
Total genomic DNA was extracted from blood according to standard techniques. To detect the m.1555A>G variant, we performed PCR–RFLP analysis. PCR was performed with the following primers: (rCRS 1410–1585) 1555_PCR_RFLP_F_5′-GGGTCGAAGGTGGATTTAGCAGTAAAC-3′; 1555_PCR_RFLP_R_5′-TTTCCAGTACACTTACCATGTTACGACTgG-3′. PCR products were digested by HaeIII restriction enzyme (Toyobo, Osaka, Japan) and were subjected to electrophoresis in 8% polyacrylamide gel. Fragments containing the wild-type allele produced restriction fragments of 121 and 55 bp after digestion. As reverse primer contains a mismatch G instead of T at 1557, the m.1555A>G mutation creates a novel HaeIII restriction site, resulting in three fragments 91, 55 and 30 bp in length. Sanger sequencing was performed using ABI 3130XL and BigDye v3.1 Terminators (Applied Biosystems) per the manufacturer’s protocols. Sequencing primers were as follows: (rCRS 1410–1690) 1555_seq_F_5′-GGTCGAAGGTGGATTTAGCA-3′ and 1555_seq_R_5′-GGGTTTGGGGCTAGGTTTAG-3′.
Cell culture
Patient-derived fibroblasts and control fibroblast cells were cultured at 37 °C and 5% CO2 in Dulbecco’s modified Eagle’s medium (4.5 or 1.0 g l−1 glucose; Nacalai Tasque Inc., Kyoto, Japan) supplemented with 10–20% fetal bovine serum and 1% penicillin–streptomycin. Normal neonatal human dermal fibroblasts (Takara Bio, Shiga, Japan) and normal fetal human dermal fibroblasts (fHDFs; Toyobo) were used as control fibroblast cells.
RNA extraction and cDNA synthesis
Total RNA was purified from fibroblast cells using the SV Total RNA Isolation System (Promega, Madison, WI, USA). Total RNA was also isolated from HEK293FT cells transfected with expression plasmids of wild-type or mutated TAZ cDNA using the same isolation protocol. cDNA was synthesized from total RNA using ReverTra Ace (Toyobo).
Quantitation of TAZ mRNA by quantitative reverse transcriptase PCR (qRT-PCR)
qRT-PCR was performed to quantify TAZ mRNA. qRT-PCR of cDNA isolated from human cells was performed using Power SYBR Green PCR Master Mix (Life Technologies, Warrington, UK), and Mx3000P (Agilent Technologies, Santa Clara, CA, USA). The relative mRNA concentration was normalized to the average expression of GAPDH. Primers for qRT-PCR were as follows: (NM_000116: CDS 68–218) TAZ_rt-F_5′-TCGTCATGGGCTTGGTG-3′ and TAZ_rt-R_5′-ATGCAGGACTGGTGATTGGA-3′ for TAZ; (NM_001289745: CDS 866–983) GAPDH_F_5′-ACACCCACTCCTCCACCTTT-3′ and GAPDH_R_5′-ATGAGGTCCACCACCCTGT-3′ for GAPDH.
Complementation test
Patient and control fibroblasts were infected with a lentiviral mammalian expression vector system expressing mitochondria-targeted TurboRFP (mtTurboRFP) and TAZ cDNA (NM_000116).
Cells were cultured at 37 °C and 5% CO2 in Dulbecco’s modified Eagle’s medium (4.5 or 1.0 g l−1 glucose; Nacalai Tasque) supplemented with 10%–20% fetal bovine serum and 1% penicillin–streptomycin. Normal fHDFs (Toyobo) were used as control fibroblast cells. Open reading frame (ORF) of candidate gene was PCR amplified from cDNA. Primer sequences used to clone TAZ (NM_000116: CDS 1–876) cDNA were as follows: CSCA-XhoI-Kosak-TAZ-F_5′-AGTGGCGGCCGctcgagCCACCatgCCTCTGCACGTGAAGTG-3′; V5-XhoI-TAZ_R_5′-TAGGCTTACCctcgagTCTCCCAGGCTGGAGGTG-3′. ORF and pTurboRFP-mito (TurboRFP fused to a mitochondrial targeting sequence derived from the subunit VIII of human cytochrome C oxidase; Evrogen, Moscow, Russia) were cloned into the CS-CA-MCS lentiviral vector with a C-terminal V5 tag, CAG promoter for mammalian cell expression and blasticidin resistance using the In-Fusion HD Cloning Kit (Clontech Laboratories, Shiga, Japan). Following this, 2 × 106 HEK293FT cells were seeded in six-cm plates and co-transfected with ViraPower Packaging vectors (pLP1, pLP2, pLP/VSVG; Invitrogen, Carlsbad, CA, USA) and a pCS-CA-ORF(candidate gene)-blast vector. Transfection was performed using Lipofectamine 2000 (Invitrogen).
After 24 h of transfection, medium was replaced with fresh medium. Supernatant containing the viral particles was collected 48 h after transfection and filtered through a 0.45 μm filter. Patients’ skin fibroblasts were infected with the viral supernatant and 5 μg ml−1 polybrene (Sigma, St Louis, MO, USA) for 24–48 h. After 5–7 days, selection was initiated with 1–2 μg ml−1 blasticidin. After 1–2 months of selection, mitochondria were harvested from the cells for sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) or blue native (BN)-PAGE.
Respiratory chain enzyme assays
To prepare enriched mitochondria, cell pellets were resuspended in ice-cold MegaFb Buffer (250 mM sucrose, 2 mM HEPES, 0.1 mM EGTA, pH 7.4) and homogenized with 20 strokes at 4200 r.p.m. The homogenates were centrifuged for 10 min at 600 g. Supernatants were centrifuged for an additional 10 min at 14 400 g. Pellets were resuspended in 900 μl MegaFb buffer, and 450 μl aliquots were resuspended in hypotonic buffer (25 mM potassium phosphate, pH 7.2, 5 mM MgCl2) for complex I and IV, citrate synthase and protein concentration assays and centrifuged for 10 min at 14 400 g. Pellets were resuspended in Hypotonic Buffer and subjected to three freeze–thaw cycles. These samples were stored at −80 °C prior to enzyme assays. Respiratory chain enzyme activities were measured using cary300 (Agilent Technologies).25 CI and CIV activities were expressed as percentages of citrate synthase activity.
SDS-PAGE, BN-PAGE and western blotting
To isolate mitochondria, cell pellets were suspended in mitochondria isolation buffer A (220 mM mannitol, 20 mM HEPES, 70 mM sucrose, 1 mM EDTA, pH 7.4, 2 mg ml−1 bovine serum albumin, 1 × protease inhibitor cocktail) and homogenized with 20 strokes on ice at 4200 r.p.m. Homogenates were separated into cytosolic and nuclear fractions after centrifugation at 700 g for 5 min at 4 °C. The supernatants were centrifuged at 10 000 g for 10 min at 4 °C. Mitochondrial pellets were rinsed twice with mitochondria isolation buffer B (220 mM mannitol, 20 mCC HEPES, 70 mM sucrose, 1 mM EDTA, pH 7.4, 1 × protease inhibitor cocktail). Mitochondrial protein levels were determined using a bicinchoninic acid assay (Pierce BCA Protein Assay Kit; Thermo Scientific, Rockford, IL, USA).
For SDS-PAGE analyses, total cell lysates were solubilized in M-PER Mammalian Protein Extraction Reagent (Thermo Fisher Scientific, Waltham, MA, USA) and denatured for 30 min at 37 °C. Enriched mitochondria were denatured for 5 min at 95 °C. Prepared samples were separated by electrophoresis on 10% SDS-PAGE gels, depending on the size of the detected protein. For BN-PAGE analyses, the NativePAGE Novex Bis-Tris Gel System (Life Technologies Corporation, Carlsbad, CA, USA) was used according to the manufacturer’s protocol. Mitochondrial fractions were solubilized in NativePAGE sample buffer containing 1% Triton X-100 or 1% digitonin and separated on 4–16% NativePAGE gels. Anti-NDUFA9 (CI), anti-70 kDa Fp Subunit (CII), anti-core 1 (CIII), anti-subunit 1 (CIV) and anti-V5 antibodies were purchased from Life Technologies. Anti-HSP60 was purchased from Abcam (Cambridge, UK); two different anti-tafazzin antibodies were purchased from GeneTex (Irvine, CA, USA) and Abcam; and anti-β-actin was purchased from Sigma.
Cybrid assay
To generate transmitochondrial cytoplasmic hybrids (cybrids), an mtDNA-less (rho0) derivative of HeLa cells were fused with patient’s fibroblasts (Pt1) and normal fibroblasts (fHDFs) as previously described.27
Morphological study
Control and patient cells were seeded on a 35-mm glass bottom dish. Mitochondria were stained with 50 nM MitoTracker Orange CMTMRos (Molecular probes, Eugene, OR, USA) for 30 min in Dulbecco’s modified Eagle’s medium containing 10% fetal bovine serum. Cells were visualized with a Leica TCS SP8 (Leica Microsystems, Wetzlar, Germany) confocal microscope.
Measurement of CL and MLCL
CL and MLCL from blood spots were analyzed as described by Kulik et al.28 The measurement of MLCL/CL was performed at the Academic Medical Center in Amsterdam as previously reported by them.28
Results
Whole-exome sequencing and whole mtDNA sequencing of Pt1 identified a novel hemizygous frameshift mutation c.39_60del p.(Pro14Alafs*19) in exon 1 of TAZ and the deafness-associated m.1555A>G variant in mitochondrial 12S rRNA (MT-RNR1), respectively.25 The homoplasmicity of m.1555A>G variant in Pt1 was confirmed by PCR–RFLP analysis (Supplementary Figure S1c). Inheritance of m.1555A>G variant from mother was confirmed by direct sequencing (Supplementary Figure S1b).
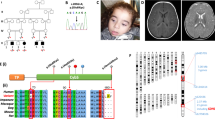
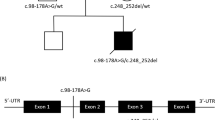
The presence of the TAZ mutation and its inheritance were validated and confirmed by trio-based Sanger sequencing. The patient’s mother carried the c.39_60del mutation in TAZ (Figure 1a). Tafazzin has a conserved N-terminal transmembrane domain and C-terminal cytosolic domain. The affected residue p.(Pro14Alafs*19) is located in the N-terminal transmembrane domain of tafazzin. The mutation was not present in the Human Tafazzin (TAZ) Gene Mutation and Variation Database (revision 28 March 2015; http://www.barthsyndrome.org).
Identification of a novel mutation in TAZ. (a) The pedigree of Pt1 showed inheritance of a hemizygous mutation {NM_000116: c.39_60del p.(Pro14Alafs*19)} from the mother. (b) Trio-based sequencing chromatogram illustrated the TAZ mutation in Pt1. (c) Schematic diagram representing the relative position of two mutations in TAZ (NM_000116).
In Pt2, a c.280C>T mutation resulting in p.(Arg94Cys) substitution was identified in exon 3 of TAZ by Sanger sequencing. This mutation has been reported previously.2, 29
To observe the effect of mutations on TAZ expression, we assessed the level of tafazzin protein and TAZ mRNA expression. We observed an apparent loss of tafazzin protein in fibroblasts from Pt1 and Pt2 (Figure 2a). The molecular weight of tafazzin varied between total cell lysates and the mitochondrial fraction (MT) of fibroblasts. In MT, tafazzin was approximately 44 kDa, whereas tafazzin in total cell lysates was 28 kDa. This may be due to the association of tafazzin with multiple protein complexes. Previously, it was demonstrated that tafazzin is incorporated into multiple protein complexes and small tafazzin protein complexes remain stable in the absence of CL. CL and other phospholipids are required for assembly of larger tafazzin complexes.30 In contrast with protein expression, TAZ transcript levels were upregulated in Pt1 fibroblast cells (Figure 2b). To further investigate the effect of these mutations, mutant or wild-type TAZ cDNA expression plasmids were introduced into HEK293FT cells. Although TAZ cDNA derived from Pt1 fibroblast cells were overexpressed in HEK293FT cells, we found loss of tafazzin protein and upregulated TAZ mRNA (Figures 2d and e). These results strongly suggest that p.(Pro14Alafs*19) induced the upregulation of TAZ mRNA expression to compensate for a lack of tafazzin protein.
Tafazzin protein syntheses, TAZ mRNA expression and morphological features of mitochondria in patient-derived fibroblasts. (a) SDS-PAGE/western blotting showed a decrease in endogenous tafazzin protein in total cell lysate (TCL) and mitochondrial extract (MT) of Pt1 and Pt2 fibroblast. β-Actin and HSP60 were used as loading controls. C1, control 1 (fHDF); C2, control 2 (NHDF). (b and c) TAZ mRNA expression levels were determined by qRT-PCR. Relative expression was measured to GAPDH mRNA in control and patient-derived fibroblasts. Each value referred to the mean of three independent experiments. (d) Mitochondrial extracts of V5-tagged mtTurboRFP, wild-type TAZ cDNA and mutated TAZ (Pro14Alafs*19 and Arg94Cys) cDNA overexpressing HEK293FT cells were isolated and V5 fusion proteins were detected by SDS-PAGE/western blotting analysis with an anti-V5 antibody. HSP60 was used as a loading control. Blasticidin-S deaminase (BSD) in TCL of overexpressed cells was detected by SDS-PAGE/western blotting with anti-BSD antibody. BSD encoded by the CS-CA-MCS plasmids was assessed as a transfection control. (e) qRT-PCR-based determination of TAZ mRNA expression in transfected HEK293FT cell lines. (f) Mitochondria of C2, and Pt1 and Pt2 fibroblast cells were stained with MitoTracker Orange and visualized by confocal microscopy. Scale bar, 50 μm.
Previously, it was reported that loss of CL leads to decreased mitochondrial fusion and fragmented mitochondria.31 TAZ mutation causes reduction of CL levels thereby affecting mitochondrial morphology. To observe the effect of TAZ mutation on mitochondrial morphology, we visualized mitochondrial morphology of patient fibroblast cells by confocal microscopy. The fragmented mitochondria were increased in both patients’ cells (Figure 2f).
To determine the pathogenicity of the m.1555A>G mutation in Pt1, we generated transmitochondrial cybrid lines harboring mutated m.1555A>G or wild-type TAZ by fusing rho0-HeLa cells with enucleated fibroblasts. We analyzed mitochondrial respiratory function in these cybrid cells. For BN-PAGE analysis, cells were solubilized using the strong detergent Triton X-100 and the mild detergent digitonin. BN-PAGE analysis using 1% Triton X-100 showed that Pt1 fibroblasts have CI and CIV assembly defects (Figure 3a). In addition, BN-PAGE using digitonin-solubilized cells revealed destabilization of CI+CIII2+CIV and CI+CIII2 supercomplexes in Pt1 fibroblast cells (Figure 3b). However, the cybrid cells from Pt1 showed normal respiratory chain assembly and stable supercomplexes (Figures 3a and b). In patient fibroblast, CI and CIV enzyme activities were decreased in comparison to control fibroblast (Figure 3c). Furthermore, the cybrids harboring mutation m.1555A>G exhibited a recovery of CI and CIV enzyme activities (Figure 3d) compared with cybrids derived from control fibroblasts. SDS-PAGE analysis of cybrid cells showed that endogenous tafazzin protein can ameliorate the deficiency caused by mutant TAZ (Figure 3e). From these findings, we conclude that the m.1555A>G mutation was not pathogenic for Pt1 and that wild-type TAZ derived from HeLa nuclear DNA recovers mitochondrial functions.
Mitochondrial complex assembly status and measurement of mitochondrial complex enzyme activities in cybrid cell lines and complementation assay. (a) BN-PAGE/western blotting analysis of mitochondrial respiratory chain complexes with anti-complex I–IV antibodies in control, patient fibroblast and cybrid cells. Three control and three patient cybrid clones were obtained and analyzed. Cybrids were established from rho0-HeLa cell and control or Pt1 fibroblasts. (b) Supercomplex stabilization revealed by BN-PAGE/western blotting using 1% digitonin-solubilized control, patient fibroblasts and cybrid cells. (c and d) Biochemical assessment of mitochondrial complex activities in (c) control and patient fibroblasts; activities were normalized relative to C1, and (d) control and patient cybrid cells; activities were normalized relative to control-1 cybrid cells. (e) Expression of endogenous tafazzin protein in cybrid cells. (f) The mitochondria were isolated from control or patient fibroblasts following the lentiviral-mediated expression of mtTurboRFP-V5 or TAZ-V5 cDNA and were analyzed by BN-PAGE/western blotting. Complementation with TAZ-V5 restored the assembly levels of both complexes I and IV. (g) mtTurboRFP-V5 and TAZ-V5 proteins in the isolated mitochondria were detected by SDS-PAGE/western blotting. HSP60 was used as a loading control.
We further performed a cellular complementation experiment to determine whether wild-type TAZ can rescue the mitochondrial defects caused by TAZ deficiency in Pt1. Pt1 fibroblasts and control fibroblasts were transfected with a lentiviral vector expressing the mtTurboRFP construct (control) and wild-type TAZ cDNA (TAZ-Δ5). To investigate whether wild-type TAZ could restore the defect in respiratory complex assembly caused by TAZ mutation, we performed BN-PAGE analysis. We found that CI and CIV assemblies were rescued by wild-type TAZ expression (Figure 3f). SDS-PAGE/western blotting analysis confirmed the expression of tafazzin in transfected cells (Figure 3g).
High-performance liquid chromatography–tandem mass spectrometry analyses of dried blood spots were performed to measure the MLCL/CL ratio in Pt1 and Pt2 and were 9.30 and 6.10, respectively (Table 1). Reference value of the measurement was 0–0.3.
Discussion
In the present study, we report phenotypic variability in patients with TAZ mutations confirmed by whole-exome sequencing, Sanger sequencing and whole mtDNA sequence analyses. Immunoblotting analysis revealed that tafazzin protein was undetectable in patient-derived fibroblasts. Exogenous expression of c.39_60del p.(Pro14Alafs*19) recombinant mutant protein in HEK293FT cells showed that this mutation affected tafazzin protein expression. Using cellular complementation, we verified that the mitochondrial dysfunction in Pt1 was caused by TAZ deficiency.
Patient harboring TAZ mutation c.39_60del p.(Pro14Alafs*19) had never been reported to manifest cardiomyopathy, whereas patient with c.280C>T p.(Arg94Cys) mutation had dilated cardiomyopathy, which is one of the common features of BTHS. This result, which is similar to the findings of a previous report,32 showed that 10% of patients with BTHS did not develop cardiomyopathy. Patients with BTHS may not exhibit cardiac problems or may be asymptomatic for life.10 This suggests the presence of important, unidentified modifying factors that affect the phenotype or severity of cardiomyopathy. Such modifications may include environmental influences or other modifying factors.33 CL deficiency is tissue specific, and disease severity and phenotypic variability does not correlate with the degree of CL deficiency. Future research should therefore be focused on identifying additional modifying factors.34
Patients with BTHS may not have profound CL deficiency, and indeed elevated MLCL represents the abnormality in CL remodeling owing to TAZ mutation. Therefore, the MLCL/CL ratio above the reference range is a better diagnostic marker rather than CL assay alone. Bowron et al.10 found patients with normal CL level previously; revising these patients owing to family history and clinical features, they found MLCL/CL ratios were significantly high, which underscore the importance of the measurement of this ratio.10 Although Pt1 had no history of cardiomyopathy, MLCL/CL ratio was significantly high in this patient. Therefore, BTHS was confirmed biochemically by this lipid assay in both of our patients.
Myopathy, which is a common feature of BTHS,32 was also presented by our two patients. Mazurová et al.29 also observed muscle hypotonia in four of their patients with BTHS. They developed myopathy shortly after birth and motor milestones were also delayed. Two patients experienced fatigue. Elevated activity of creatine kinase was not found.29 Growth failure is another common feature in BTHS patients. Pt1 had growth retardation and speech difficulty. At birth, our patients had normal weight; however, about 30% of newborns with BTHS had a birth weight below the third percentile and patients with growth retardation revealed low level of arginine.16 Arginine supplementation may be used as an auxiliary therapy for improving the growth rate of BTHS patients.16 There may be a partial defect in the Krebs cycle, which metabolizes arginine to alpha-ketoglutaric acid and protein synthesis affected by arginine depletion.35
We observed loss of tafazzin protein expression in our patients, with increased TAZ mRNA expression in one patient and reduced mRNA expression in the other. mRNA levels do not necessarily reflect protein levels and the increased mRNA expression in BTHS fibroblasts may have been a compensatory measure in response to loss of functional enzyme. The reduction of mRNA is likely caused by reduced mRNA stability or increased degradation.20 There are several TAZ mutations that affect the proportion of splice variants and induce unusual TAZ mRNA expression.36 The c.39_60del mutation may therefore upregulate the expression of unusual TAZ variants.
In Pt1 fibroblasts, we observed a decrease in CI and CIV assembly, destabilization of supercomplexes and decrease of CI and CIV enzymatic activity. Respiratory complex analysis revealed destabilization of respiratory supercomplexes and decreased CI, CIV and CV in BTHS cells.37 They hypothesized that CL is required for association/stabilization of the complexes into supercomplexes and to modulate the abundance of individual respiratory chain complexes. Mutated tafazzin may incorrectly remodel CL and affect the assembly and stability of the electron transport chain.38
Both our patients exhibited fragmented mitochondrial morphology. This may be due to decreased CL levels caused by TAZ mutation. In a yeast study, it was observed that CL and phosphatidylethanolamine deficiency affects mitochondrial fusion and induces mitochondrial fragmentation. They found that loss of CL led to reduced steady-state levels of short and long isoforms of the mitochondrial fusion protein Mgm1p (human homolog Opa1) and fragmented mitochondria were observed owing to fusion defects.31 Morphological variations in the mitochondria of lymphoblasts from BTHS patients were observed, including enlarged size, fragmentation, adhesion of cristae membranes and deformed intercristae space, which was also reported in a mouse model of BTHS.39, 40
Cybrid study is an effective tool to discriminate pathogenicity of either mtDNA or nuclear DNA origin. Observation of transmitochondrial cybrids with a uniform nuclear background of HeLa cell demonstrated no clear difference in the stability of respiratory chain complexes, supercomplexes nor enzyme activities between Pt1 and control. This revealed that m.1555A>G mutation was not pathogenic to this patient. Though m.1555A>G mutation is considered as the most common cause of maternally inherited deafness in 12S rRNA gene,41 Pt1 had never presented any sign of deafness.
Early diagnosis of BTHS is a prerequisite for early and effective therapy and for preventing crisis owing to sepsis. Moreover, prompt molecular characterization of BTHS patients is vital for prenatal diagnosis, preventing future BTHS cases in the same family and to increase the quality of life of an affected individual providing supportive management. Genetic analysis of TAZ mutation and measurement of MLCL/CL ratio are necessary to confirm clinical and other biochemical diagnosis even with an incomplete phenotype, in absence of cardiac disease or other typical signs of BTHS and to avoid underdiagnosis of this potentially life-threatening inborn error of metabolism.
References
Vreken, P., Valianpour, F., Nijtmans, L. G., Grivell, L. a, Plecko, B., Wanders, R. J. et al. Defective remodeling of cardiolipin and phosphatidylglycerol in Barth syndrome. Biochem. Biophys. Res. Commun. 279, 378–382 (2000).
Johnston, J., Kelley, R. I., Feigenbaum, A, Cox, G. F., Iyer, G. S., Funanage, V. L. et al. Mutation characterization and genotype-phenotype correlation in Barth syndrome. Am. J. Hum. Genet. 61, 1053–1058 (1997).
Gonzalez, I. L. Barth syndrome:TAZ gene mutations, mRNAs, and evolution. Am. J. Med. Genet. Part A 134A, 409–414 (2005).
Aprikyan, A. A. & Khuchua, Z. Advances in the understanding of Barth syndrome. Br. J. Haematol. 161, 330–338 (2013).
Sakamoto, O., Ohura, T., Katsushima, Y., Fujiwara, I., Ogawa, E., Miyabayashi, S. et al. A novel intronic mutation of the TAZ (G4.5) gene in a patient with Barth syndrome: creation of a 5′ splice donor site with variant GC consensus and elongation of the upstream exon. Hum. Genet. 109, 559–563 (2001).
Ferri, L., Dionisi-Vici, C., Taurisano, R., Vaz, F. M., Guerrini, R. & Morrone, A. When silence is noise: infantile-onset Barth syndrome caused by a synonymous substitution affecting TAZ gene transcription. Clin. Genet. 90, 461–465 (2016).
Jefferies, J. L. Barth syndrome. Am. J. Med. Genet. Part C Semin. Med. Genet. 163, 198–205 (2013).
Barth, P. G., Valianpour, F., Bowen, V. M., Lam, J., Duran, M., Vaz, F. M. et al. X-linked cardioskeletal myopathy and neutropenia (Barth syndrome): an update. Am. J. Med. Genet. 126A, 349–354 (2004).
Roberts, A. E., Nixon, C., Steward, C. G., Gauvreau, K., Maisenbacher, M., Fletcher, M. et al. The Barth Syndrome Registry: distinguishing disease characteristics and growth data from a longitudinal study. Am. J. Med. Genet. A 158A, 2726–2732 (2012).
Bowron, A., Honeychurch, J., Williams, M., Tsai-Goodman, B., Clayton, N., Jones, L. et al. Barth syndrome without tetralinoleoyl cardiolipin deficiency: a possible ameliorated phenotype. J. Inherit. Metab. Dis. 38, 279–286 (2015).
Ronvelia, D., Greenwood, J., Platt, J., Hakim, S. & Zaragoza, M. V. Intrafamilial variability for novel TAZ gene mutation: Barth syndrome with dilated cardiomyopathy and heart failure in an infant and left ventricular noncompaction in his great-uncle. Mol. Genet. Metab. 107, 428–432 (2012).
Moric-Janiszewska, E. & Markiewicz-Łoskot, G. Genetic heterogeneity of left-ventricular noncompaction cardiomyopathy. Clin. Cardiol. 31, 201–204 (2008).
Chang, B., Momoi, N., Shan, L., Mitomo, M., Aoyagi, Y., Endo, K. et al. Gonadal mosaicism of a TAZ (G4.5) mutation in a Japanese family with Barth syndrome and left ventricular noncompaction. Mol. Genet. Metab. 100, 198–203 (2010).
Cosson, L., Toutain, A., Simard, G., Kulik, W., Matyas, G., Guichet, A. et al. Barth syndrome in a female patient. Mol. Genet. Metab. 106, 115–120 (2012).
Ferri, L., Donati, M. a, Funghini, S., Cavicchi, C., Pensato, V., Gellera, C. et al. Intra-individual plasticity of the TAZ gene leading to different heritable mutations in siblings with Barth syndrome. Eur. J. Hum. Genet. 23, 1708–1712 (2015).
Rigaud, C., Lebre, A.-S., Touraine, R., Beaupain, B., Ottolenghi, C., Chabli, A. et al. Natural history of Barth syndrome: a national cohort study of 22 patients. Orphanet J. Rare Dis. 8, 70 (2013).
Clarke, S. L. N., Bowron, A., Gonzalez, I. L., Groves, S. J., Newbury-Ecob, R., Clayton, N. et al. Barth syndrome. Orphanet J. Rare Dis. 8, 23 (2013).
Ferri, L., Donati, M. A., Funghini, S., Malvagia, S., Catarzi, S., Lugli, L. et al. New clinical and molecular insights on Barth syndrome. Orphanet J. Rare Dis. 8, 27 (2013).
Horvath, S. E. & Daum, G. Lipids of mitochondria. Prog. Lipid Res. 52, 590–614 (2013).
Houtkooper, R. H., Turkenburg, M., Poll-The, B. T., Karall, D., Pérez-Cerdá, C., Morrone, A. et al. The enigmatic role of tafazzin in cardiolipin metabolism. Biochim. Biophys. Acta Biomembr. 1788, 2003–2014 (2009).
Schlame, M., Ren, M., Xu, Y., Greenberg, M. L. & Haller, I. Molecular symmetry in mitochondrial cardiolipins. Chem. Phys. Lipids 138, 38–49 (2005).
Houtkooper, R. H. & Vaz, F. M. Cardiolipin, the heart of mitochondrial metabolism. Cell Mol. Life Sci. 65, 2493–2506 (2008).
Pfeiffer, K., Gohil, V., Stuart, R. a., Hunte, C., Brandt, U., Greenberg, M. L. et al. Cardiolipin stabilizes respiratory chain supercomplexes. J. Biol. Chem. 278, 52873–52880 (2003).
Vaz, F. M., Houtkooper, R. H., Valianpour, F., Barth, P. G. & Wanders, R. J. A. Only one splice variant of the human TAZ gene encodes a functional protein with a role in cardiolipin metabolism. J. Biol. Chem. 278, 43089–43094 (2003).
Kohda, M., Tokuzawa, Y., Kishita, Y., Nyuzuki, H., Moriyama, Y., Mizuno, Y. et al. A comprehensive genomic analysis reveals the genetic landscape of mitochondrial respiratory chain complex deficiencies. PLoS Genet. 12, e1005679 (2016).
Hazkani-Covo, E., Zeller, R. M. & Martin, W. Molecular poltergeists: mitochondrial DNA copies (numts) in sequenced nuclear genomes. PLoS Genet. 6, e1000834 (2010).
Hayashi, J., Ohta, S., Kikuchi, A., Takemitsu, M., Goto, Y. & Nonaka, I. Introduction of disease-related mitochondrial DNA deletions into HeLa cells lacking mitochondrial DNA results in mitochondrial dysfunction. Proc. Natl Acad. Sci. USA 88, 10614–10618 (1991).
Kulik, W., Van Lenthe, H., Stet, F. S., Houtkooper, R. H., Kemp, H., Stone, J. E. et al. Bloodspot assay using HPLC-tandem mass spectrometry for detection of barth syndrome. Clin. Chem. 54, 371–378 (2008).
Mazurová, S., Tesařová, M., Magner, M., Houšťková, H., Hansíková, H., Augustínová, J. et al. Novel mutations in the TAZ gene in patients with Barth syndrome. Prague Med. Rep. 114, 139–153 (2013).
Xu, Y., Malhotra, A., Claypool, S. M., Ren, M. & Schlame, M. Tafazzins from Drosophila and mammalian cells assemble in large protein complexes with a short half-life. Mitochondrion 21, 27–32 (2015).
Joshi, A. S., Thompson, M. N., Fei, N., Ttemann, M. H. & Greenberg, M. L. Cardiolipin and mitochondrial phosphatidylethanolamine have overlapping functions in mitochondrial fusion in Saccharomyces cerevisiae. J. Biol. Chem. 287, 17589–17597 (2012).
Spencer, C. T., Bryant, R. M., Day, J., Gonzalez, I. L., Colan, S. D., Thompson, W. R. et al. Cardiac and clinical phenotype in Barth syndrome. Pediatrics 118, e337–e346 (2006).
Ren, M., Phoon, C. K. L. & Schlame, M. Metabolism and function of mitochondrial cardiolipin. Prog. Lipid Res. 55, 1–16 (2014).
Schlame, M., Kelley, R. I., Feigenbaum, A., Towbin, J. a., Heerdt, P. M., Schieble, T. et al. Phospholipid abnormalities in children with Barth syndrome. J. Am. Coll. Cardiol. 42, 1994–1999 (2003).
Coman, D., Yaplito-Lee, J. & Boneh, A. New indications and controversies in arginine therapy. Clin. Nutr. 27, 489–496 (2008).
Kirwin, S. M., Manolakos, A., Barnett, S. S. & Gonzalez, I. L. Tafazzin splice variants and mutations in Barth syndrome. Mol. Genet. Metab. 111, 26–32 (2014).
Gonzalvez, F., D’Aurelio, M., Boutant, M., Moustapha, A., Puech, J. P., Landes, T. et al. Barth syndrome: cellular compensation of mitochondrial dysfunction and apoptosis inhibition due to changes in cardiolipin remodeling linked to tafazzin (TAZ) gene mutation. Biochim. Biophys. Acta Mol. Basis Dis. 1832, 1194–1206 (2013).
Schlame, M., Rua, D. & Greenberg, M. L. The biosynthesis and functional role of cardiolipin. Prog. Lipid Res. 39, 257–288 (2000).
Acehan, D., Xu, Y., Stokes, D. L. & Schlame, M. Comparison of lymphoblast mitochondria from normal subjects and patients with Barth syndrome using electron microscopic tomography. Lab. Invest. 87, 40–48 (2007).
Acehan, D., Vaz, F., Houtkooper, R. H., James, J., Moore, V., Tokunaga, C. et al. Cardiac and skeletal muscle defects in a mouse model of human Barth syndrome. J. Biol. Chem. 286, 899–908 (2011).
Prezant, T. R., Agapian, J. V., Bohlman, M. C., Bu, X., Oztas, S., Qiu, W. Q. et al. Mitochondrial ribosomal RNA mutation associated with both antibiotic-induced and non-syndromic deafness. Nat. Genet. 4, 289–294 (1993).
Acknowledgements
This work was supported by a grant of the Innovative Cell Biology by Innovative Technology (Cell Innovation Program) from the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan to YO; the Support Project and a grant of Strategic Research Center in Private Universities from the MEXT, Japan to Saitama Medical University Research Center for Genomic Medicine. This work was also supported by Grants-in-Aid of the Research on Intractable Diseases (Mitochondrial Disorder) from the Ministry of Health, Labor and Welfare of Japan to KM. YO received a Special research grant from Takeda Science Foundation. NNB is a recipient of the Saitama Medical University Research Fellowship. We acknowledge the Laboratory Genetic Metabolic Disease in the Academic Medical Center in Amsterdam for measuring the MLCL/CL ratio. We thank Atsuko Imai for providing valuable comments. We are grateful to Ryoko Nakamura, Department of Pediatric Neurology, Fukuoka Children’s Hospital, Fukuoka, Japan; who is an attending physician of one of our patients.
Author contributions
NNB, YK and YO designed the study. NNB, YK and KI acquired data. KN, J-IH, YT, MK, HN, YY-S, KM and AO gave critical comments. YO, NNB and YK wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Borna, N., Kishita, Y., Ishikawa, K. et al. A novel mutation in TAZ causes mitochondrial respiratory chain disorder without cardiomyopathy. J Hum Genet 62, 539–547 (2017). https://doi.org/10.1038/jhg.2016.165
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2016.165
This article is cited by
-
A high mutation load of m.14597A>G in MT-ND6 causes Leigh syndrome
Scientific Reports (2021)