Abstract
A genome-wide association study (GWAS) for cold medicine-related Stevens–Johnson syndrome (CM-SJS) with severe ocular complications (SOC) was performed in a Japanese population. A recently developed ethnicity-specific array with genome-wide imputation that was based on the whole-genome sequences of 1070 unrelated Japanese individuals was used. Validation analysis with additional samples from Japanese individuals and replication analysis using samples from Korean individuals identified two new susceptibility loci on chromosomes 15 and 16. This study might suggest the usefulness of GWAS using the ethnicity-specific array and genome-wide imputation based on large-scale whole-genome sequences. Our findings contribute to the understanding of genetic predisposition to CM-SJS with SOC.
Similar content being viewed by others
Introduction
Stevens–Johnson syndrome (SJS) is an acute inflammatory vesiculobullous reaction of the skin and mucosa. It tends to involve the ocular surface, oral cavity and genitals. In patients with extensive skin detachment and a poor prognosis, the condition is called toxic epidermal necrolysis (TEN). Although these reactions are rare (annual incidence 1–6 cases per 106 persons),1, 2, 3 the mortality rate is 3% for SJS and 27% for TEN.4 Survivors often suffer severe sequelae such as vision loss due to severe ocular complications (SOC).5
About 40% of SJS/TEN patients diagnosed by dermatologists develop SOC.2 Among SJS and TEN patients, especially those with SJS/TEN with SOC, cold medicines (CM), including multi-ingredient cold medications and non-steroidal anti-inflammatory drugs, were identified as causative.2, 6, 7, 8, 9 We reported that in the Japanese, CM-related SJS/TEN (CM-SJS/TEN) with SOC was strongly associated with HLA-A*02:06 and significantly associated with HLA-B*44:03.9 In our first genome-wide association study (GWAS), we analyzed SJS/TEN with SOC patients using the Affymetrix GeneChip Mapping 500 K array set (Affymetrix, Santa Clara, CA, USA). We found an association between prostaglandin E receptor 3 (PTGER3) and SJS/TEN with SOC.6, 10 We subsequently documented that this association was stronger in CM-SJS/TEN patients with SOC than in all SJS/TEN patients with SOC.11 Moreover, we identified an interaction with additive effects between HLA-A*02:06 and the high-risk genotypes PTGER3 rs1327464 GA or AA.11 A second GWAS using the Affymetrix AXIOM Genome-Wide ASI 1 array (Affymetrix) identified the association between IKAROS family zinc-finger 1 (IKZF1) and CM-SJS/TEN with SOC in the Japanese.7 These findings were indicative of a genetic predisposition for CM-SJS/TEN with SOC.
A single-nucleotide polymorphism (SNP) array (Japonica Array, Toshiba, Japan) specifically designed for the genome-wide study of the Japanese population was developed by Kawai et al.12 Its haplotype collection was constructed from whole-genome sequences of 1070 Japanese individuals.13 We posited that whole-genome imputation with the Japonica array would increase the statistical power for detecting associated variants of CM-SJS/TEN with SOC than was possible with the AXIOM Genome-wide ASI 1 array we reported previously.7
In the current study, we performed GWAS using the Japonica array and imputation with the 1KJPN panel (Tohoku Medical Megabank Organization constructed the reference panel) to look for additional genetic predispositions for CM-SJS/TEN with SOC in the Japanese population.
Materials and methods
Patients
This study was approved by the institutional review board of Kyoto Prefectural University of Medicine (KPUM), the University of Tokyo and other collaborating research institutes (Seoul National University College of Medicine, Yonsei University College of Medicine, Chonnam National University Medical School and College of Medicine, The Catholic University of Korea in Korea).
We used the same samples from 117 Japanese individuals with CM-SJS/TEN with SOC and 691 healthy Japanese volunteers as were used in our earlier GWAS with the Affymetrix AXIOM Genome-Wide ASI 1 array.7
SOC patients were defined as those with ocular sequelae such as severe dry eye, trichiasis, symblepharon and conjunctival invasion into the cornea in the chronic stage,5 and as patients who manifested severe conjunctivitis with pseudomembranes and epithelial defects on the ocular surface (cornea and/or conjunctiva) in the acute stage.14 We have classified the patients who had taken CMs such as non-steroidal anti-inflammatory drugs and multi-ingredient cold medications for a few–several days before the disease onset for common-cold symptoms as CM-SJS/TEN. The specific drugs they used were not named by all patients (Supplementary Table 1). In Japan, doctors in hospital usually prescribed CM such as non-steroidal anti-inflammatory drugs and acetaminophen with some antibiotics. However, we previously reported that the use of CM such as non-steroidal anti-inflammatory drugs and cold-remedies was significantly associated with SJS/TEN with SOC, whereas the use of antibiotics was not associated.2 Detailed information on the SJS/TEN patients and the controls we analyzed are shown in Supplementary Methods in the Online Repository.
GWAS, imputation with 1KJPN and validation analysis
In the GWAS, we genotyped 808 samples from the same SJS/TEN patients (n=117) and the healthy Japanese controls (n=691) used in our earlier study.7 We used the Japonica SNP array12 according to the manufacturer’s instructions. All genotyped samples passed the recommended sample quality control metric for the AXIOM arrays (dish QC>0.82); we excluded three control samples with an overall call rate <97%. We recalled the remaining 805 samples with Genotyping Console 4.2.0.26 software (Affymetrix).
Because we had already performed identity-by-descent estimation and principal component analysis for all 808 samples genotyped with the AXIOM Genome-Wide ASI 1 array;7 all samples used in the current study were known to be from unrelated individuals and the effect of population stratification was negligible. A quantile–quantile plot of the distribution of test statistics for the comparison of genotype frequencies in our patients and controls showed that the inflation factor λ was 1.024 for all tested SNPs and decreased to 1.021 when SNPs in the human leukocyte antigen (HLA) region (chr 6: 29 645 038–33 360 787) were excluded (Supplementary Figure 1).
For imputation, the clustering plots were classified by the Ps classification function in the SNPolisher package (version 1.5.2; Affymetrix). SNPs that were assigned 'recommended' by the Ps classification were retained. Excluded were SNPs with a call rate <99.0%, a Hardy–Weinberg equilibrium (HWE) test result of P<0.0001 or a minor allele frequency (MAF) <0.5%. Prephasing was conducted first with these SNPs using SHAPEIT (v.2.r644);15 the options were —burn 10, —prune 10 and —main 25. Genotype imputation was performed on the phased genotypes with IMPUTE2 (ver. 2.3.1)16 using a phased reference panel of 1070 healthy Japanese individuals (1KJPN panel).13 For IMPUTE2, the applied options were -Ne 2000, -k hap 1000, -k 120, -burnin 15, and -iter 50.
After genome-wide imputation genotyped with the Japonica array, we applied a SNP call rate⩾95%, MAF⩾1%, and HWE P⩾0.001 for SNP quality control for data cleaning of the samples. The 6,714,496 SNPs and insertions and deletions (indels) on autosomal chromosomes that passed the quality control filters were used for GWAS.
Confirmation of the genotypes at 6 candidate SNPs with P<10−7 in the GWAS and the replication study using Korean samples was with the TaqMan SNP genotyping assay (Applied Biosystems, Foster City, CA). We genotyped extended sample sets of Japanese SJS patients (n=138) and controls (n=883) to validate the GWAS results. Samples from Korean patients (n=31) and controls (n=110) were genotyped to replicate the GWAS results (Supplementary Methods in the Online Repository).
Statistical analysis
For the validation of GWAS and the replication study, we applied the χ2 or the Fisher's exact test on a two-by-two contingency table of the allele frequency, the dominant model and the recessive model. The odds ratio (OR) and the confidence interval (CI) were calculated using the major alleles as references. Meta-analysis was with the Mantel–Haenszel method. Heterogeneity among studies was examined using the Breslow–Day test.
Results
GWAS of CM-SJS/TEN with SOC in Japanese individuals using the Japonica array and imputation with 1KJPN
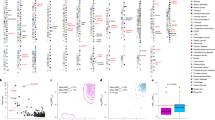
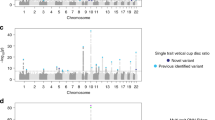
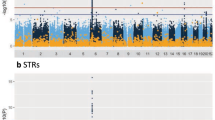
Figure 1 presents a genome-wide view of the SNP association data for 117 Japanese CM-SJS/TEN patients with SOC and 688 controls (3 of the 691 control samples were excluded because their overall call rate was <97%). We used the Japonica array and performed genome-wide imputation based on 1KJPN; the associations were based on allele frequencies (MAF >0.01, HWE P>0.001, 6 714 496 SNPs). Imputation with 1KJPN clearly increased the number of candidate SNPs because SNP association data acquired with the Japonica array without imputation with 1KJPN included only 612 877 sites (MAF>0.01, HWE P>0.001) (Supplementary Figure 2).
The HLA-A region exhibited the strongest association with susceptibility to CM-SJS/TEN with SOC (P=2.2 × 10−19, OR=4.6). This is consistent with findings reported in our earlier studies; they had shown a strong association between SJS/TEN with SOC and HLA-A*02:06.8, 9, 17, 18, 19
Outside of the HLA region, there were 82 polymorphisms manifesting associations with P<10−7 in the allele frequency model. Of these, four exhibited MAF>0.05 and they were located on chromosome 16. There was a strong linkage disequilibrium (LD) between rs6500265 and rs9933632 and between rs9888871 and rs76213482. The other 78 polymorphisms manifested 0.02>MAF>0.01 and were located in a single LD block on chromosome 15 (Supplementary Table 2). For validation, we selected one to three SNPs from each LD block; three SNPs (rs16957893, rs12324361, rs113301740) were from the LD block on chromosome 15 and one (rs9888871) or two SNPs (rs6500265, rs9933632) were from the LD blocks on chromosome 16. Validation analysis of these six SNPs by applying the TaqMan assay to samples from the 117 Japanese CM-SJS/TEN with SOC patients and the 691 controls showed genome-wide significant associations for three of these six SNPs (Table 1).
Extended analysis with additional Japanese and Korean samples
A different additional Japanese sample set of 213 samples (21 Japanese CM-SJS/TEN-SOC patients and 192 Japanese healthy controls) was used in a subsequent extended analysis to further evaluate these six SNPs. We found that despite the relatively small sample size, one of the six SNPs exhibited a significant association after applying Bonferroni correction (Table 1).
It was difficult to evaluate the three SNPs with MAF <0.01 because of the low sample number (n=21). The combined results of the GWAS validation analysis and the analysis of additional sample sets showed that four of the six SNPs exhibited significant genome-wide associations (Table 1).
We also genotyped the 6 SNPs in 141 samples from a Korean population (31 CM-SJS/TEN with SOC patients and 110 controls). Because their number was small, we found no significant association between Korean CM-SJS/TEN-SOC and the six SNPs after applying Bonferroni correction (Supplementary Table 3). However, the OR of the six SNPs exhibited the same direction of association as the OR of the Japanese population. Furthermore, meta-analysis using the Japanese and Korean samples revealed a significant genome-wide association with rs16957893 ((C vs G), OR=5.61, P=1.72 × 10−8), with rs12324361 ((C vs T), OR=5.58, P=2.63 × 10−8) and with rs113301740 ((A vs C), OR=5.58, P=2.63 × 10−8) (Supplementary Table 4).
Discussion
GWAS using the Japonica array, an ethnicity-specific array and imputation with a Japanese 1070 whole-genome-sequenced panel (1KJPN) revealed that outside of the HLA region there were 82 polymorphisms manifesting associations with P<10−7 in the allele frequency model. We found that four SNPs with P<10−7 in two LD blocks on chromosome 16 were present in an intergenic region between ZNF423 and CNEP1R1; their functions are unclear. Of the 78 variants with P<10−7 in a single LD block on chromosome 15, 47 (60.3%) were located in introns of the REC114 gene; 16 (20.5%) in introns of the NPTN gene, 5 (6.4%) in an exon (no amino-acid change) or an intron of LOC105370891, 4 (5.1%) in an intergenic region between LOC105370891 and CD276 and the remaining 6 (7.7%) in introns of the CD276 gene. There was no variant with amino-acid changes among the above genes.
REC114, a meiotic recombination protein, may have a role in meiotic DNA double-strand breaks.20 The function of LOC105370891 has not been studied in detail. Neuroplastin encodes a type I transmembrane protein belonging to the immunoglobulin superfamily; it may be involved in cell–cell interactions or cell–substrate interactions.21 CD276 is a member of the immunoglobulin superfamily and is thought to participate in the regulation of the T-cell-mediated immune response.22 However, the molecular mechanisms involved in the associations with CM-SJS/TEN-SOC remain to be identified.
Validation analysis of selected six SNPs by applying the TaqMan assay to samples from the 138 (117 of GWAS and additional 21) Japanese CM-SJS/TEN with SOC patients and the 883 (691 of GWAS and additional 192) controls showed genome-wide significant associations for four of these six SNPs. However, when we performed logistic regression analysis adjusted for sex in Japanese 138 cases and 883 controls because of the difference of sex, P-values of the four SNPs observed slightly increased (adjusted P-values of rs16957893, rs6500265, rs9888871 and rs9933632 was 4.66 × 10−6, 1.32 × 10−7, 6.62 × 10−8 and 7.17 × 10−8, respectively). Moreover, when we performed logistic regression analysis adjusted for sex using genome-wide imputed GWAS data, we detected two additional regions (chromosome 3 and 20) with P-values <10−6. This finding is consistent with our previous report.7 Thus, further investigation with more case and control samples which are sex-matched might be needed.
While typing with the Japonica array detected 612 877 sites (MAF>0.01, HWE P>0.001), in combination with imputation we detected 6 101 619 more; that is, a total of 6 714 496 polymorphisms. Thus, the combination identified ~10 times more polymorphisms than did typing with the Japonica array alone. Moreover, among 82 variants with P<10−7 in the allele frequency detected by GWAS using the Japonica array and imputation with 1KJPN, only three (3.7%) were identified when typing was with the Japonica array alone. The combination of the Japonica array and imputation with the 1KJPN panel identified new SNPs.
Elsewhere we reported that in Japanese individuals, HLA-A*02:06 was strongly associated with CM-SJS/TEN with SOC.9 We also documented that HLA-A*02:06 with TLR3 polymorphisms exerted more than additive effects23 and that HLA-A*02:06 with PTGER3 polymorphisms exhibited additive effects in CM-SJS/TEN with SOC.11 Although our sample numbers were small, we found that HLA-A*02:06 with Rec114 rs16957893 CG might also exert more than additive effects in CM-SJS/TEN with SOC (OR=110, P=4.45 × 10−8) (Supplementary Table 5). The combination of variants' effects between HLA-A*02:06 and Rec114 rs16957893 CG might be independent of the interactions between HLA-A*02:06 and TLR3 rs3775296 T/T because CM-SJS/TEN with SOC in Japanese patients with both HLA-A*02:06 and Rec114 rs16957893 CG did not harbor TLR3 rs3775296 T/T (data not shown).
In Japanese individuals, combined genotyping for the associated variants thus far identified; that is, HLA-A*02:06, TLR3 rs3775296 T/T and Rec114 rs16957893 CG, may help to predict the risk for CM-SJS/TEN with SOC. Based on our previous and current observations, we suggest that besides microbial infections and CMs, the combination of multiple gene polymorphisms and their interactions contributes strongly to the onset of CM-SJS/TEN with SOC.
In the current GWAS, we used the Japonica array and imputation with the 1KJPN panel to investigate CM-SJS/TEN-SOC in Japanese individuals. We also performed replication studies using Korean samples to identify host genetic factors that predispose to CM-SJS/TEN with SOC. The findings reported here increase our understanding of pathogenic pathways involved in the development of CM-SJS/TEN with SOC.
References
Yetiv, J. Z., Bianchine, J. R. & Owen, J. A. Jr. Etiologic factors of the Stevens-Johnson syndrome. South Med. J. 73, 599–602 (1980).
Sotozono, C., Ueta, M., Nakatani, E., Kitami, A., Watanabe, H., Sueki, H. et al. Predictive factors associated with acute ocular involvement in Stevens-Johnson syndrome and toxic epidermal necrolysis. Am. J. Ophthalmol. 160, 228–237.e2 (2015).
Chan, H. L., Stern, R. S., Arndt, K. A., Langlois, J., Jick, S. S., Jick, H. et al. The incidence of erythema multiforme, Stevens-Johnson syndrome, and toxic epidermal necrolysis. A population-based study with particular reference to reactions caused by drugs among outpatients. Arch. Dermatol. 126, 43–47 (1990).
Power, W. J., Ghoraishi, M., Merayo-Lloves, J., Neves, R. A. & Foster, C. S. Analysis of the acute ophthalmic manifestations of the erythema multiforme/Stevens–Johnson syndrome/toxic epidermal necrolysis disease spectrum. Ophthalmology 102, 1669–1676 (1995).
Sotozono, C., Ang, L. P., Koizumi, N., Higashihara, H., Ueta, M., Inatomi, T. et al. New grading system for the evaluation of chronic ocular manifestations in patients with Stevens-Johnson syndrome. Ophthalmology. 114, 1294–1302 (2007).
Ueta, M., Sotozono, C., Nakano, M., Taniguchi, T., Yagi, T., Tokuda, Y. et al. Association between prostaglandin E receptor 3 polymorphisms and Stevens-Johnson syndrome identified by means of a genome-wide association study. J Allergy Clin Immunol. 126, 1218–1225 e10 (2010).
Ueta, M., Sawai, H., Sotozono, C., Hitomi, Y., Kaniwa, N., Kim, M. K. et al. IKZF1, a new susceptibility gene for cold medicine-related Stevens-Johnson syndrome/toxic epidermal necrolysis with severe mucosal involvement. TJ Allergy Clin Immunol. 135, 1538–1545 e17 (2015).
Ueta, M., Kannabiran, C., Wakamatsu, T. H., Kim, M. K., Yoon, K. C., Seo, K. Y. et al. Trans-ethnic study confirmed independent associations of HLA-A*02:06 and HLA-B*44:03 with cold medicine-related Stevens-Johnson syndrome with severe ocular surface complications. Sci Rep. 4, 5981 (2014).
Ueta, M., Kaniwa, N., Sotozono, C., Tokunaga, K., Saito, Y., Sawai, H. et al. Independent strong association of HLA-A*02:06 and HLA-B*44:03 with cold medicine-related Stevens-Johnson syndrome with severe mucosal involvement. Sci Rep. 4, 4862 (2014).
Ueta, M., Tamiya, G., Tokunaga, K., Sotozono, C., Ueki, M., Sawai, H. et al. Epistatic interaction between Toll-like receptor 3 (TLR3) and prostaglandin E receptor 3 (PTGER3) genes. J Allergy Clin Immunol 129, 1413–1416 e11 (2012).
Ueta, M., Tokunaga, K., Sotozono, C., Sawai, H., Yoon, K. C., Kim, M. K. et al. HLA-A*02:06 and PTGER3 polymorphism exerts additive effects in cold medicine-related Stevens-Johnson syndrome with severe ocular complications. Hum Genome Var. 2, 15023 (2015).
Kawai, Y., Mimori, T., Kojima, K., Nariai, N., Danjoh, I., Saito, R. et al. Japonica array: improved genotype imputation by designing a population-specific SNP array with 1070 Japanese individuals. J Hum Genet. 60, 581–587 (2015).
Nagasaki, M., Yasuda, J., Katsuoka, F., Nariai, N., Kojima, K., Kawai, Y. et al. Rare variant discovery by deep whole-genome sequencing of 1,070 Japanese individuals. Nat. Commun. 6, 8018 (2015).
Sotozono, C., Ueta, M., Koizumi, N., Inatomi, T., Shirakata, Y., Ikezawa, Z. et al. Diagnosis and treatment of Stevens-Johnson syndrome and toxic epidermal necrolysis with ocular complications. Ophthalmology. 116, 685–690 (2009).
Delaneau, O., Zagury, J. F. & Marchini, J. Improved whole-chromosome phasing for disease and population genetic studies. Nat. Methods 10, 5–6 (2013).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Ueta, M., Tokunaga, K., Sotozono, C., Inatomi, T., Yabe, T., Matsushita, M. et al. HLA class I and II gene polymorphisms in Stevens-Johnson syndrome with ocular complications in Japanese. Mol Vis. 14, 550–555 (2008).
Ueta, M., Sotozono, C., Tokunaga, K., Yabe, T. & Kinoshita, S. Strong association between HLA-A*0206 and Stevens–Johnson syndrome in the Japanese. Am. J. Ophthalmol. 143, 367–368 (2007).
Ueta, M. Ethnic differences in the association between human leukocyte antigen and Stevens–Johnson syndrome. Eur. Ophthalmic Rev. 3, 15–18 (2009).
Li, J., Hooker, G. W. & Roeder, G. S. Saccharomyces cerevisiae Mer2, Mei4 and Rec114 form a complex required for meiotic double-strand break formation. Genetics 173, 1969–1981 (2006).
Wilson, M. C., Kraus, M., Marzban, H., Sarna, J. R., Wang, Y., Hawkes, R. et al. The neuroplastin adhesion molecules are accessory proteins that chaperone the monocarboxylate transporter MCT2 to the neuronal cell surface. PloS One. 8, e78654 (2013).
Yi, K. H. & Chen, L. Fine tuning the immune response through B7-H3 and B7-H4. Immunol. Rev. 229, 145–151 (2009).
Ueta, M., Tokunaga, K., Sotozono, C., Sawai, H., Tamiya, G., Inatomi, T. et al. HLA-A*0206 with TLR3 polymorphisms exerts more than additive effects in Stevens-Johnson syndrome with severe ocular surface complications. PloS One. 7, e43650 (2012).
Acknowledgements
We thank the patients and volunteers who participated in our study. We also thank Hiromi Nishigaki, Natsumi Baba, Kayoko Yamada, Kayoko Kato for processing the blood samples and for genotyping, and Drs Sangchul Yoon, Hyo Seok Lee and Kyu-Yeon Hwang for their assistance in collecting blood and DNA samples and for providing clinical information. Computational resources for genome-wide imputation were provided by the ToMMo supercomputer system. This work was supported by a grant from Toshiba Corporation Healthcare Company and also supported, in part, by grants in aid from the Ministry of Education, Culture, Sports, Science and Technology of the Japanese government (BioBank Japan Project), and supported, in part, by JSPS Core-to-Core Program, A Advanced Research Networks. This work was also supported, in part, by the Tohoku Medical Megabank Project (Special account for reconstruction from the Great East Japan Earthquake). This work was also supported, in part, by the Promotion Project of Knowledge-Based Industrial Clustering of Okinawa Prefecture. The funding agencies had no role in the study design, data collection or analysis, the decision to publish or the manuscript preparation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
MN and KK hold concurrent positions at the Department of Cohort Genome Information Analysis endowed by Toshiba Corporation. KT, MU and MN received research funding from Toshiba Corporation.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Ueta, M., Sawai, H., Shingaki, R. et al. Genome-wide association study using the ethnicity-specific Japonica array: identification of new susceptibility loci for cold medicine-related Stevens–Johnson syndrome with severe ocular complications. J Hum Genet 62, 485–489 (2017). https://doi.org/10.1038/jhg.2016.160
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2016.160
This article is cited by
-
A genome-wide association study identifies distinct variants associated with pulmonary function among European and African ancestries from the UK Biobank
Communications Biology (2023)
-
Mapping of susceptible variants for cold medicine-related Stevens–Johnson syndrome by whole-genome resequencing
npj Genomic Medicine (2021)
-
Current Perspectives on Stevens-Johnson Syndrome and Toxic Epidermal Necrolysis
Clinical Reviews in Allergy & Immunology (2018)
-
Association of Human Leukocyte Antigen Class 1 genes with Stevens Johnson Syndrome with severe ocular complications in an Indian population
Scientific Reports (2017)