Abstract
The Jomon period of the Japanese Archipelago, characterized by cord-marked ‘jomon’ potteries, has yielded abundant human skeletal remains. However, the genetic origins of the Jomon people and their relationships with modern populations have not been clarified. We determined a total of 115 million base pair nuclear genome sequences from two Jomon individuals (male and female each) from the Sanganji Shell Mound (dated 3000 years before present) with the Jomon-characteristic mitochondrial DNA haplogroup N9b, and compared these nuclear genome sequences with those of worldwide populations. We found that the Jomon population lineage is best considered to have diverged before diversification of present-day East Eurasian populations, with no evidence of gene flow events between the Jomon and other continental populations. This suggests that the Sanganji Jomon people descended from an early phase of population dispersals in East Asia. We also estimated that the modern mainland Japanese inherited <20% of Jomon peoples’ genomes. Our findings, based on the first analysis of Jomon nuclear genome sequence data, firmly demonstrate that the modern mainland Japanese resulted from genetic admixture of the indigenous Jomon people and later migrants.
Similar content being viewed by others
Introduction
Genome-wide single-nucleotide polymorphism (SNP) data analyses1, 2 support the dual-structure model3, 4 for the formation of modern Japanese populations, in which indigenous Jomon people admixed with later migrants who brought rice agriculture. The Jomon culture geographically ranged from Hokkaido to the Okinawa islands, and the Jomon people inhabited the Japanese Archipelago from ~16000 to 2500 years before present (YBP).5, 6 The origins and phylogenetic relationships of the Jomon people, however, are still elusive.
Ancient DNA sequences of the Jomon people provide direct evidence of their genetic characteristics. Mitochondrial DNA (mtDNA) sequences of the Jomon people and their haplotypes have been determined for many individuals.7, 8, 9, 10, 11, 12 Haplogroup N9b, whose frequency is generally low in modern East Eurasians,13, 14 was found to be quite frequent in the mtDNA of the Jomons.9, 10, 11 This suggests a long-term isolation of the Jomons from continental populations. However, inferring human population history only from mtDNA data are insufficient because of their limited genetic information. Thanks to new technologies, it is now possible to analyze nuclear genome sequences of ancient human remains15, 16, 17, 18, 19, 20, 21, 22, 23 and those of modern human individuals.24 We therefore determined the nuclear genome sequences of two Jomon individuals and compared them with available data so as to infer the origin of modern Japanese.
Materials and methods
DNA extraction, library preparation, sequencing and sequence mapping
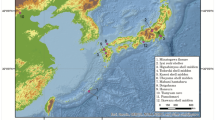
We extracted DNA from the teeth of Jomon individuals kept at the University Museum, the University of Tokyo. These samples were originally excavated at the Sanganji Shell Mound in the 1950s.25 This shell mound is located in the northern part of Fukushima Prefecture, Tohoku region, Japan (Supplementary Figure 1). The mtDNA haplotypes of four Sanganji Jomon individuals were previously genotyped.11 We designate two DNA extracts of the Sanganji 131421-3 individual, one extracted by Kanzawa-Kiriyama et al.,11 and the other newly extracted for this study, as A1 and A2, respectively. The DNA extract from sample 131464 is referred to as B. The new DNA extract (A2) was prepared using Adachi et al.’s12 protocol with some modifications (Supplementary Methods S1). Three DNA libraries for A1, A2 and B were prepared from the DNA solution extracted from their molars (Supplementary Methods S1). The GAIIx platform was used for library A1, and 120-bp paired-end sequence reads were generated following the manufacturer’s protocol. We used Hiseq2000 to generate 100-bp paired-end sequence reads for libraries of DNA samples A2 and B. Sequence mapping was then conducted (Supplementary Methods S2). PCR duplicates, which were produced during library preparations, were removed using MarkDuplicates from Picard program (https://broadinstitute.github.io/picard/) as well as PickingBases, newly developed by one of us (KK) (see Supplementary Perl Scripts S1).
DNA authenticity checking and error rate estimation
To clarify the authenticity of the mapped sequence reads, we checked the frequency of post-mortem misincorporation and depurination, which are characteristic of ancient DNA. As it is difficult to remove all modern human DNA contaminations that come from reagents and experimental rooms, it is also essential to estimate the frequency of endogenous DNA from Jomon individuals. To estimate the frequency of contamination, we focused on sequence reads mapped to mtDNA and estimated ratio of mismatched reads to each mtDNA haplogroup, which were previously classified into N9b11 (Supplementary Methods S3).
We estimated error rate using the method of Reich et al.16 Genome sequences of chimpanzee,26 14 present-day humans19, 24 and two ancient individuals19, 22 (see Supplementary Table S1 for their list) were used to estimate the degree of post-mortem changes in the Sanganji Jomon. The first and last few bases of sequence reads that harbor many post-mortem changes specific to ancient genomes were removed in the Denisovan,19 Altai Neanderthal22 as well as Sanganji Jomon sequences using bamUtil (http://genome.sph.umich.edu/wiki/BamUtil) to minimize their effect in downstream statistical analyses. For the Sanganji Jomon sequences, different numbers of bases were trimmed in each sample based on transition rates; in sample A1, we trimmed the first and last 10 bases because this sample had more transversion substitutions at the end of sequence reads compared with sample A2 and B, whereas we trimmed only five bases in sample A2 and B. For the archaic humans, we trimmed five bases. See Supplementary Methods S4 for details.
Principal component analysis (PCA)
SNPs observed in modern human populations were used to examine the genetic relationship between individuals through PCA. Four data sets from the 1000 Genomes Project Consortium,24 the Human Genome Diversity Project (HGDP),27 the HapMap Project28 and the Japanese Archipelago Human Population Genetics Consortium data,1 were used. First, three filters, Hardy–Weinberg equilibrium (HWE) p<1e-10, minor allele frequency<0.01, genotype call rate<0.95, were applied using PLINK.29 These filters were applied in all downstream analyses. For each individual, we randomly chose one nucleotide at each site and duplicated it to create homozygous diploid genotypes to compensate for the fact that no heterozygous/homozygous states could be ascertained in the Jomon genome data due to the low coverage.30 This step evens the data condition between modern samples and ancient samples. We then merged the modern human data sets with the Sanganji Jomon sequence. We used only sites where the Jomon genome sequences had coverage. Finally, we performed PCA based on the overlapping SNPs by using the smartpca program in the EIGENSOFT.31
Allele-sharing analysis
Because the number of individuals is limited in the Sanganji Jomon data set, it is not suitable to interpret population differentiation between Jomon and modern populations based on Fst values. We investigated the frequency of allele sharing between Sanganji Jomon individuals and modern populations to infer the degree of genetic similarity between them. We used array-based genome-wide SNP data, and the original genotype data were used for the analysis instead of the homozygous diploid data sets for PCA. For the Sanganji Jomon, we created homozygous diploid genotypes to compare those with modern populations by using PLINK-freq option. We counted the number of shared allele with Sanganji Jomon and calculated the proportion of shared allele in each individual. We then took the average and calculated the standard error in each population.
Phylogenetic analysis
We used TreeMix,32 neighbor-joining33 and Neighbor-Net34 to estimate phylogenetic relationships among ancient and modern humans (Supplementary Methods S5). All substitution sites data or transversion-only substitution sites data were used for the analysis, and we compared those results to check the effect of post-mortem changes. For HGDP data,27 we used the all substitution data only because of limited number of SNPs. In addition, we used modern human genome data from Papuan and Karitiana16 and ancient human genome from Mal’ta (MA1) dating to ~24 000 YBP,23 Ust’-Ishim dating to 45 000 YBP21 and Denisovans19 to the analyses.
D-statistics to detect gene flow
D-statistics analysis16 can infer gene flow without invoking demographic models, and is based on the number of shared alleles between three individuals of interest (H1, H2 and H3) and one outgroup (O). At any particular locus, the pattern ABBA is where H2 and H3 share the derived allele ‘B’, whereas H1 and O share the ancestral allele ‘A’. Conversely, BABA is where the derived allele is shared between H1 and H3 and the ancestral allele is shared between H2 and O. We basically used only transversion substitution sites to minimize errors in our data sets except for HGDP because of data limitation. The number of BABA and ABBA patterns, nBABA and nABBA, is used to calculate the D-value and Z-score with ADMIXtools.35 We added data of SGVP Malay population in Singapore36 to HGDP data sets for the D-statistics analysis.
Results
Dating, mapping and assignment of sequence reads
The Jomon pottery of the Sanganji Shell Mound site belong to the late to final Jomon period.25 The Jomon skeletal remains were dated by one of us (MY) using the radiocarbon method to be ~3000 YBP (Supplementary Table S2), corresponding to the final Jomon period.
The mapped sequence reads from three DNA libraries A1, A2 and B covered 21, 39 and 58 million nucleotides (0.46–2.18%) of the human genome build hg19, respectively (Supplementary Table S3). These low ratio of endogenous human DNA are typically observed in ancient samples.15, 20 In the DNA library treated with two restriction enzymes, the frequency of sequence reads mapped to hg19 was ~50% higher than the frequency obtained from the untreated library (1.30% and 0.87%, respectively). Many reads appear to be derived from bacteria, but ~50% of the reads were not assigned uniquely to specific species or groups in BLASTN (Supplementary Figure S2), consistent with previous studies.15, 20
We deposit all short read sequences produced for this study to DDBJ Sequence Read Archive (http://trace.ddbj.nig.ac.jp/dra/). Sequence data are available for download thorough SRA accession number DRA004001.
Damage estimates and DNA sequence authenticity
Average DNA lengths of mapped sequences reads were 94.5, 102.2 and 76.9 bp, respectively. Read length distribution for each library is shown in Supplementary Figure S3. From the ratio of sequence reads mapped to X and Y chromosomes, we concluded that individuals A and B were male and female, respectively. This identification is consistent with morphological observation of these individuals.25 We observed characteristic features of ancient DNA degradation, including C to T misincorporation at high frequencies in 3′ and 5′ ends and an increased fraction of purines close to sequence read termini (Supplementary Figures S4 and S5, respectively), indicating that those sequence reads most likely contain endogenous Jomon DNAs. Sample A1, however, has more transversion substitutions than samples A2 and B. We cannot explain the reason, but after mtDNA capturing and the sequencing with MiSeq, the pattern disappeared.
Although contamination frequency (3.03%; 0.0–7.17%, 95% confidence interval) of DNA sample A2 based on captured mtDNA sequence reads was low (Supplementary Results S1), the frequency (11.3%; 2.8–19.9%, 95% confidence interval) based on simple shotgun sequencing was high compared with the other two DNA samples. We thus used sequence data for only samples A1 and B in the downstream analyses to minimize the effect of the cross contaminations. After merging the data of A1 and B for downstream analyses, ~77.7 million nucleotide positions were available for analysis.
We estimated error rates in 16 modern/archaic human genomes16, 19, 22 as well as in Sanganji Jomon. Although the error rate of the Sanganji Jomon sequence was 0.828–0.848% over all sites, the error rate was only 0.011–0.015% in the transversion sites, which is comparable with the rate in modern humans (Supplementary Table S7). PickingBases (PB) removed more errors compared with MarkDuplicates (Supplementary Table S7). We found more transition substitutions (A/G or C/T) in Sanganji Jomon than in other modern humans, which appears to be caused by post-mortem changes characteristic of ancient DNA (Supplementary Tables S8-S9). We also would like to note that Jomon-specific transversion substitutions (0.0129%) and unreported transversions in dbSNP 141(0.0113%) were comparable (Supplementary Table S7). Similar values were also observed in modern human data, and this comparison strengthens the low error rate of the Sanganji Jomon nucleotide sequences. Interestingly, the number of SNPs reported in dbSNP was small in Sanganji Jomon compared with other non-Africans; for example, 4169 and 5270 in Sanganji Jomon and Han, respectively (Supplementary Table S7). This observation is consistent with our expectation that many Jomon SNPs are unreported in dbSNP. We confirmed the reliability of PB in PCA, and the result using PB are concordant with that of MarkDuplicates, and are not biased (Figure 1a and Supplementary Figure S7). Because of smaller sequence error rates of PB than that of MarkDuplicates, we used data sets treated with PB for further analyses. In total, 28 288 sites were masked as N with PB.
Genetic relationship between Sanganji Jomon and other modern and archaic humans
We investigated the genetic relationships between Sanganji Jomon and modern humans from different parts of the world, including Africans, West Eurasians, East Eurasians, Native Americans and Sahulians37 (descendants of people who migrated to Sahul land); see the map shown in Supplementary Figure S1.
PCA
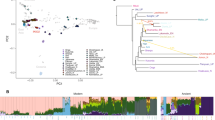
When worldwide populations are compared, PCA illustrates the genetic similarity of Sanganji Jomon and East Eurasians compared with African, European, Sahulian and Native American peoples (Figure 1a, Supplementary Figures S7-S11). However, Sanganji Jomon is located slightly closer to the center of the three major population groups in Figure 1a. This indicates the genetic uniqueness of Sanganji Jomon among East Eurasians, and/or the effect of post-mortem changes in ancient DNA, although the latter is unlikely because the result using only transversion sites showed very similar results (Supplementary Figures S8 and S10). Next, we investigated the genetic relationship between the Sanganji Jomon and East Eurasians. Comparison with 1000 Genomes Project24 East Asians (JPT (Japanese Tokyo), CHB (Han Chinese in Beijing), CHS (Southern Han Chinese), CDX (Chinese Dai in Xishuangbanna, China) and KHV (Kinh in Ho Chi Minh City, Vietnam)) based on 46 158 SNP sites show that the Sanganji Jomon is located quite apart from the other modern East Eurasians, and modern Japanese are situated between the Sanganji Jomon and continental East Eurasians (Figure 1b). Although the Sanganji Jomon data were merged for sequence data of A1 and B, the PCA plot location of the Sanganji Jomon was similar to Figure 1b when A1 and B were independently analyzed (Supplementary Figure S13). We therefore surmise that the merged data are reliable for further analyses. The uniqueness of the Sanganji Jomon was also observed when only transversion sites were used (Supplementary Figure S12), again indicating that the uniqueness was not the result of post-mortem changes.
The comparison with the genome-wide SNP data of HGDP populations27 also showed the unique status of the Sanganji Jomon, who was positioned far apart from all modern East Eurasians in PC2 and PC3, although only 6864 SNPs were used. (Figure 2a,Supplementary Figure S14). The uniqueness of the Sanganji Jomon within East Eurasians is consistent with the results including Europeans and Africans. When the Ainu, the mainland Japanese and the Ryukyuan from the Japanese Archipelago1 and CHB28 were compared with Sanganji Jomon, PC1 separated the Ainu and Sanganji Jomon from the other populations (Figure 2b). The population closest to the Sanganji Jomon was the Ainu, followed by the Ryukyuan and then the mainland Japanese. It appears that PC1 corresponds to the degree of genetic contribution from the Jomon people to the other Japanese Archipelago populations, whereas PC2 separated the Ainu from Sanganji Jomon. When the genomic data of 1000 Genomes Project East Asians24 were included in the PCA analysis (Figure 2b), PC3 separated the Ainu from the Sanganji Jomon (Supplementary Figure S15).
Principal component analysis with genome-wide SNP data. (a) Sanganji Jomon and HGDP East Eurasians based on 6864 SNPs with PB. (b) Sanganji Jomon, individuals of three populations inhabiting the Japanese Archipelago (Ainu, mainland Japanese and Ryukyuan), and Chinese Beijing (CHB) based on 5392 SNP sites with PB.
Allele sharing analysis
Allele sharing analysis using 5392 SNP sites (Figure 3a) showed that the Ainu had the highest percentage of allele sharing with the Sanganji Jomon, followed by the Ryukyuan, the mainland Japanese and CHB, similar to the projection of PC1 in Figure 2b. Using the HGDP East Eurasian data set with 7081 SNP sites (Figure 3b), the mainland Japanese had the highest allele sharing with the Sanganji Jomon. Interestingly, southern East Eurasians (green bars) had slightly higher allele-sharing percentages than northern East Eurasians (blue bars), although we have to be careful with the effect of post-mortem changes.
Allele sharing between Sanganji Jomon and modern humans. Vertical line indicates the frequency of having the same allele with the Sanganji Jomon. (a) Comparison with Japanese Archipelago populations and Chinese Beijing (CHB) based on 5392 SNP sites with PB, (b) comparison with HGDP East Eurasians, Sahulians and Native Americans based on 7081 SNP sites with PB. Colors blue, green, brown and violet indicate northern East Eurasians, southern East Eurasians, Sahulians and Native Americans, respectively.
TreeMix analysis
We constructed maximum-likelihood population trees (ML tree) using TreeMix to investigate the phylogenetic relationship and the presence of admixture events between Sanganji Jomon and other human populations. When three migration events were assumed, the gene flow from Sanganji Jomon to JPT (modern Japanese) appeared (Figure 4) (bootstrap probability=42%). The directionality of this gene flow event was as expected, however, the inferred gene flow from Karitiana to Mal’ta MA1 (Supplementary Results S2; Supplementary Figures S16b-j and S17c-j), was in the reverse direction to what was reported by Raghavan et al.23 who used a much larger sequence data. This result might have been caused by using a relatively small SNP data set. We therefore used only modern human data, and reran TreeMix. We used a much larger 0.7 million SNP loci data. However, the resulting tree showed some anomalous gene flow directions; from Papuans to Denisovans (bootstrap probability=98%) and CEU to Papuans (bootstrap probability=86%), whereas the tree topology was consistent with that of Figure 4 (see Supplementary Figure S19). The reason for this anomaly may be that some filtering steps between our analyses and Reich et al.16 and Meyer et al.19 are different, and/or homozygous diploid genotypes in individual genome (that is, Denisovan, Papuan and so on) were used instead of their original genotype, though gene flow from Mal’ta MA1 to Karitiana correctly appeared in both all sites and transversion only when we add Mal’ta MA1, Karitiana and Ust’-Ishim to the large SNP loci data (data not shown).
Phylogenetic analysis I. TreeMix tree with gene flows. A comparison of Sanganji Jomon, 1000 Genomes Project worldwide populations, Papuan, Karitiana, Mal’ta MA1, Ust’-Ishim and Denisovan based on 43 310 all sites. Denisovan was used as the outgroup, and three gene flow events were estimated. The tree was drawn by using MEGA6.38 Red colored values (only those higher than 90% are shown) are bootstrap probabilities (%) for their adjacent internal branch. Arrows were manually added to this tree, and colors of migration weight (ratio of gene flow) follow TreeMix outputs. Values inside arrows are the ratio of gene flow. Bootstrap probabilities (%) of the gene flow from Sanganji Jomon to JPT, Karitiana to the root of European and Mal’ta MA1, and LWK to European, estimated out of 1000 bootstrap replicate TreeMix outputs, are 42%, 86% and 0.4%, respectively.
Distance matrix-based analysis
We constructed neighbor-joining trees (Figure 5a and Supplementary Figure S20) from distance matrices, and the Native American diverged after the divergence of the Sanganji Jomon from the modern East Eurasians in these trees. This is consistent with Figure 4, although the bootstrap probability to support this branching pattern was only 64% in Figure 5a. We also drew a Neighbor-Net network using the same distance matrix (Figure 5b and Supplementary Figure S21). Major splits are consistent with the neighbor-joining tree (Figure 5a), and split X clustered the Sanganji Jomon and JPT. This split confirms the admixed nature of the mainland Japanese inferred from the PCA and TreeMix analyses. See Supplementary Results S3 for more detailed analysis using Neighbor-Net.
D-statistic analysis
In agreement with the PCA and TreeMix results, the D-statistic test also suggested genetic affinity of the Sanganji Jomon with the East Eurasians, especially with the mainland Japanese, compared with non-East Eurasians (Figure 6a and b). This reflects the genetic continuity between the Jomon and the mainland Japanese, and support the hypothesis of an early divergence of the Jomon with the ancestor of modern East Eurasians (Supplementary Figure S23). See also Supplementary Results S4 for additional analyses using D-statistics.
D-statistic tests of Sanganji Jomon and worldwide humans. (a) Comparison of Sanganji Jomon, 1000 Genomes Project worldwide populations, Mal’ta MA1 and Ust’-Ishim based on 15 549 transversion sites. (b) Comparison with HGDP East Eurasians and Native Americans and SGVP Malay population based on 7439 SNPs.
The genetic relationship between Sanganji Jomon and the archaic humans were then investigated. Genetic affinities between Neanderthal and non-African, and between Denisovan and Sahulian were previously reported.15, 16, 19, 22 The test of ((Sanganji Jomon, San), Altai Neanderthal) implies genetic affinity of the Jomon people with Neanderthals, but the degree was not much different from the other non-Africans (Supplementary Figure S31). Compared with the other East Eurasians, Sanganji Jomon did not show additional similarities with the Denisovans.
Discussion
This is the first study on the ancient nuclear genome of the Neolithic hunter-gatherers who lived on the Japanese Archipelago. We observed more post-mortem changes in the Sanganji Jomon sample than in two other archaic human examples (~0.8% in Sanganji Jomon, Supplementary Table S7). Environmental conditions (for example, temperature, humidity, pH value, geochemical properties of soil) or use of different sample treatments may affect the condition of the ancient DNA. We did not treat the ancient DNA with uracil DNA glycosylase and endonuclease VIII, which remove C to T misincorporation from the sequence reads. This is to reduce the number of experimental steps so as to minimize the risk of contamination in library preparations. We also chose to see the post-mortem damage specific to ancient DNA to more clearly evaluate the ancient origin of the extracted DNA. This may be the reason that the DNA damage was high in Sanganji Jomon sequences.
PCA and phylogenetic analyses revealed the genetic relationships between the Sanganji Jomon and the modern human populations that we compared. We found that Sanganji Jomon was genetically quite distinct from the other modern East Eurasians. Our Jomon genotypes are still mostly based on a single read per SNP site, whereas the 1000 Genomes Project data24 had at least 4–6 × coverage. We need to be careful about the effect of genotype error rate even if we use only transversion sites (error rate=0.0129%). However, PCA plots using all sites (Figure 1 and Supplementary Figures S7 and S9) and those using transversion only (Supplementary Figures S8) are more or less the same, and TreeMix trees using all sites (Figure 4 and Supplementary Figures S16 and S18) and those using transversion only (Supplementary Figure S17) also showed similar positions for the Sanganji Jomon. Therefore, we believe that the phylogenetic relationship of the Sanganji Jomon can still be reliably estimated with our data.
Some cranial metric and dental analyses suggested that the ancestral population of the Jomon people originated from Southeast Asia,3, 39, 40, 41, 42, 43 whereas other morphological analyses44, 45 and mtDNA sequence data9 suggested that their ancestors were of Northeast Asian origin. However, neither hypothesis was supported from our analysis using direct Jomon nuclear genome sequences. Our results suggest that the Jomon people were descendants of an ancestral East Eurasian population prior to population diversifications recognizable today.
The possibility that the Sanganji Jomon ancestors diverged before the Native Americans was also inferred through the maximum-likelihood tree, neighbor-joining tree, and the D-statistic tests, but this is still inconclusive because of the small number of SNPs in the analyses. However, all the results suggest a deep Jomon divergence within East Asia, close to the Native American split. The effect of genotype errors also need to be taken into account in interpreting the apparent deep divergence of the Jomon people. A relatively large amount of genotype errors in the Jomon data set would induce their long tree branch and/or genetic similarity between the Jomon and outgroups (for example, non-East Eurasians and archaic humans) as artifacts, placing the Jomon closer to the root of outgroup-East Eurasian splits and/or showing gene flow between Jomon and those outgroups. However, since we did not observe any of those artifacts, and the inferred genotype error rate in the transversion-only sites (0.0129%) is quite small, it is unlikely that the Jomon was erroneously located at the basal position of East Eurasians. The ages of the most recent common ancestors of mitochondrial DNA haplogroups N9b and M7a (considered indigenous to Jomon and modern Japanese, and called the Jomon haplotype), were estimated to be 22 000 YBP and 23 000 YBP, respectively.10 These estimates also support a deep divergence of the Jomon people.
We also examined whether the uniqueness of the Sanganji Jomon in East Eurasia was the result of gene flow from non-East Eurasians, but the evidence for this was not suggested (Supplementary Figures S22, S24 and S25). This indicates that the ancestor of the Sanganji Jomon had been genetically isolated from continental populations after their divergence. However, our results do not deny a possibility of small fraction of gene flow between the Jomon people and the non-East Eurasians, which might not be detected in the current study because of the small amount of the determined Jomon genome sequences. The TreeMix tree and the D-statistic tests also confirm the lack of non-East Eurasian contaminations in the Jomon data via reagents because if there were such contamination during experiments, the effect would be artificially detected as gene flow in those analyses. Genetic affinity between modern East Eurasians and Melanesian compared with Sanganji Jomon was inferred, but these results were also ambiguous because of the data limitation. Further genomic sequencing of Jomon genomes in future studies will clarify such questions.
Our results show that the populations genetically closest to Sanganji Jomon are those that live on the Japanese Archipelago. The current Japanese population is widely accepted to be a result of admixture of indigenous Jomon and later migrant people, whereby the latter were agricultural people that came from continental Asia into the Japanese Archipelago probably via the Korean Peninsula during and after the Yayoi era. Until now, this hybridization scenario has not been verified directly at the genomic level. Our result based on PCA, TreeMix and neighbor-joining trees, Neighbor-Net networks, and D-statistic tests clarified that the Jomon people genetically contributed to the modern Japanese Archipelago populations. The genetic similarity would not be explained by contamination from Japanese archipelago populations during experiments, because the direction of gene flow estimated in TreeMix tree was from Jomon to JPT, not from JPT to Jomon (Figure 4), and the D((KHV or CHS, CHB) Jomon) value in D-statistic tests (Figure 6a) was not negative with significant Z-score (JPT was genetically closer to CHB than to CHS or KHV). Furthermore, if there was contamination from modern Japanese, the Jomon would be genetically closer to CHB than to CHS and KHV, leading to negative D-values and probably significant Z-scores. Therefore, we can conclude that the modern Japanese Archipelago populations are the admixture of the Jomon people and other populations with genetic affinities with modern Northeast Asians. This result is consistent with the conclusion based on the genetic analyses of modern human populations.1 However, it is difficult to pin-point the geographic origins of the candidate populations because of lack of genetic data of relevant ancient populations.
The amount of genetic contribution from the Jomon people varied among the Japanese Archipelago populations (Figures 2b and 3a and Supplementary Figure S15). The Ainu and the Ryukyuan share more alleles with the Jomon than the mainland Japanese, suggesting smaller genetic contributions from continental populations in those two populations than for the mainland Japanese, in agreement with previous findings.1, 2, 4, 13, 46, 47, 48, 49, 50 This supports the dual-structure model of Hanihara,3, 50 which is a widely accepted theory that explains the history of the three current populations that inhabit the Japanese Archipelago, and which predicts that the mainland Japanese are more admixed with agricultural continental people than the Ainu and the Ryukyuan, with major admixture occurring in and after the Yayoi period.
PC1 values of some Ainu individuals were similar to that of Sanganji Jomon, whereas PC2 separated the Ainu people from Sanganji Jomon (Figure 2b). It is probable that the ancestors of the Ainu people experienced admixture with other population(s) (for example, continental population(s) not used in the current study) after the Jomon period, and they are thus genetically differentiated from the Jomon people.51 The candidate population is the Okhotsk people, who inhabited the Northeastern costal area of the Hokkaido region, Japan, from the 5th to 13th century. They were morphologically close to modern southern Siberians such as the Nivkhi and Ulchi people,52, 53, 54, 55 who were not included in the current study. Recent studies suggest that the descendants of the Hokkaido Jomon people had admixed with the Okhotsk people and became ancestors of the modern Ainu.56, 57, 58, 59
Jinam et al.2 recently estimated the proportion of the Jomon ancestry in the mainland Japanese using the f4-ratio test35 under a specified phylogenetic scenario. The estimated proportion of Jomon ancestry differs depending on the compared populations and were within the range of 13–21%. Jinam et al.2 also used ADMIXTURE60 and found that the mainland Japanese was best explained by two ancestry components. One of them has very high proportions in many Ainu individuals, and is considered to be Jomon ancestry if we assume that the Ainu people are largely descendants of the Jomon. The average proportion of this Jomon component in the mainland Japanese was 17%, which is within the range of f4-ratio test results.2 The same genome-wide SNP data were also used for ABC analysis, and the highest likelihood model estimated the admixture proportion from the Jomon ancestry to the mainland Japanese to be 36% (22–53%, 95% confidence interval).50 We used TreeMix for estimating Jomon ancestry proportions in this study, and the frequency was 12% as shown in Figure 4. We note that it is possible that the frequency was underestimated if the Jomon people had some population substructure and the northern Jomon including the Sanganji Jomon only indirectly contributed to the modern Japanese, and/or if there are contamination during experiments from the Japanese Archipelago populations. Previously, the proportion of the Jomon component in the modern mainland Japanese was estimated to be 35% based on partial mtDNA sequences of modern human populations46 and 23–40% based on application of a demographic model of population admixture.61 Although the Jomon admixture proportion is still debatable, we would like to conclude that the Jomon component in the mainland Japanese is probably lower than 20%.
One remaining question is whether or not the ancestors of the Jomon people experienced any gene flow with archaic humans. The ancestor of modern humans who moved out of Africa admixed with Neanderthals,15, 22 whereas Denisovan DNA sequences were shown to share some similarity with the modern East Eurasians and Sahulians, especially with Papuan, Melanesian, Australian Aboriginal and Southeast Asian islanders.16, 17, 19, 62 The result of D-statistic tests implies that there was no additional gene flow with archaic humans observed in the Sanganji Jomon compared with the other East Eurasians (Supplementary Figure S31).
In conclusion, this is the first report describing the genetic relationships between the prehistoric Jomon and modern and other archaic human populations based on nuclear genome sequences, and reveals new insights on population history in East Eurasia. This report is a starting point for future studies.
Accession codes
References
Japanese Archipelago Human Population Genetics Consortium, Jinam, T., Nishida, N., Hirai, M., Kawamura, S., Oota, H. et al. The history of human populations in the Japanese Archipelago inferred from genome-wide SNP data with a special reference to the Ainu and the Ryukyuan populations. J. Hum. Genet. 57, 787–795 (2012).
Jinam, A. T., Kanzawa-Kiriyama, H., Inoue, I., Tokunaga, K., Omoto, K. & Saitou, N. Unique characteristics of the Ainu population in Northern Japan. J. Hum. Genet. 60, 565–571 (2015).
Hanihara, K. Dual structure model for the population history of the Japanese. Jpn. Rev. 2, 1–33 (1991).
Omoto, K. & Saitou, N. Genetic origins of the Japanese: a partial support for the ‘dual structure hypothesis’. Amer. J. Phys. Anthropol. 102, 437–446 (1997).
Imamura, K. Prehistoric Japan: new perspectives on Insular East Asia, University of Hawaii Press, Honolulu (1996).
Habu, J. Ancient Jomon of Japan. (Cambridge University Press, Cambridge, UK, 2004).
Horai, S., Kondo, R., Murayama, K., Hayashi, S., Koike, H. & Nakai, N. Phylogenetic affiliation of ancient and contemporary humans inferred from mitochondrial DNA. Philos. Trans. Royal Soc. London B Biol. Sci 333, 409–416 (1991).
Shinoda, K. & Kanai, S. Intracemetery genetic analysis at the Nakazuma Jomon site in Japan by mitochondrial DNA sequencing. Anthropol. Sci. 107, 129–140 (1999).
Adachi, N., Shinoda, K., Umetsu, K. & Matsumura, H. Mitochondrial DNA analysis of Jomon skeletons from the Funadomari Site, Hokkaido, and its implication for the origins of Native American. Amer. J. Phys. Anthropol. 138, 255–265 (2009).
Adachi, N., Shinoda, K., Umetsu, K., Kitano, T., Matsumura, H., Fujiyama, R. et al. Mitochondrial DNA analysis of Hokkaido Jomon skeletons: remnants of archaic maternal lineages at the southwestern edge of former Beringia. Amer. J. Phys. Anthropol. 146, 346–360 (2011).
Kanzawa-Kiriyama, H., Saso, A., Suwa, G. & Saitou, N. Ancient mitochondrial DNA sequences of Jomon teeth samples from Sanganji, Tohoku district, Japan. Anthropol. Sci. 121, 89–103 (2013).
Adachi, N., Sawada, J., Yoneda, M., Kobayashi, K. & Itoh, S. Mitochondrial DNA analysis of the human skeleton of the initial Jomon phase excavated at the Yugura cave site, Nagano, Japan. Anthropol. Sci. 121, 137–143 (2013).
Tanaka, M., Cabrera, V. M., González, A. M., Larruga, J. M., Takeyasu, T., Fuku, N. et al (2004) Mitochondrial genome variation in Eastern Asia and the peopling of Japan. Genome Res. 14, 1832–1850 (2004).
Umetsu, K., Tanaka, M., Yuasa, I., Adachi, N., Miyoshi, A., Kashimura, S. et al. Multiplex amplified product-length polymorphism analysis of 36 mitochondrial single-nucleotide polymorphisms for haplogrouping of East Asian populations. Electrophoresis 26, 91–98 (2005).
Green, R. E., Krause, J., Briggs, A. W., Maricic, T., Stenzel, U., Kircher, M. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010).
Reich, D., Green, R. E., Kircher, M., Krause, J., Patterson, N., Durand, E. Y. et al. Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature 468, 1053–1060 (2010).
Skoglund, P. & Jakobsson, M. Archaic human ancestry in East Asia. Proc. Natl Acad. Sci. USA 108, 18301–18306 (2011).
Skoglund, P., Malmström, H., Raghavan, M., Storå, J., Hall, P., Willerslev, E. et al. Origins and genetic legacy of Neolithic farmers and hunter-gatherers in Europe. Science 336, 466–469 (2012).
Meyer, M., Kircher, M., Gansauge, M., Li, H., Racimo, F., Mallick, S. et al. A high-coverage genome sequence from an archaic Denisovan individual. Science 338, 222–226 (2012).
Fu, Q., Meyer, M., Gao, X., Stenzel, U., Burbano, H. A., Kelso, J. & Pääbo, S. DNA analysis of an early modern human from Tianyuan Cave, China. Proc. Natl Acad. Sci. USA 110, 2223–2227 (2013).
Fu, Q., Li, H., Moorjani, P., Jay, F., Slepchenko, S. M., Bondarev, A. A. et al. Genome sequence of a 45000-year-old modern human from western Siberia. Nature 514, 445–449 (2014).
Prüfer, K., Racimo, F., Patterson, N., Jay, F., Sankararaman, S., Sawyer, S. et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature 505, 43–49 (2014).
Raghavan, M., Skoglund, P., Graf, K. E., Metspalu, M., Albrechtsen, A., Moltke, I. et al. Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505, 87–91 (2014).
The 1000 Genomes Project Consortium An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Fukushima Prefectural Museum Sanganji Kaizuka (in Japanese). (Fukushima Prefectural Museum, Fukushima, Japan, 1988).
Pairwise alignment of panTro2 and human (hg19) http://hgdownload.cse.ucsc.edu/goldenPath/panTro2/vsHg19/.
Li, J. Z., Absher, D. M., Tang, H., Southwick, A. M., Casto, A. M., Ramachandran, S. et al. Worldwide human relationships inferred fro`m genome-wide patterns of variation. Science 319, 1100–1104 (2008).
The International HapMap Consortium A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Purcell, S., Neale, B., Todd-Brown, K., Thomas, L., Ferreira, M. A. R., Bender, D. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Amer. J. Hum. Genet. 81, 559–575 (2007).
Sánchez-Quinto, F., Schroeder, H., Ramirez, O., Ávila-Arcos, M. C., Pybus, M., Olalde, I. et al. Genomic Affinities of Two 7,000-Year-Old Iberian Hunter-Gatherers. Curr. Biol. 22, 1494–1499 (2012).
Patterson, N., Price, A. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Pickrell, J. K. & Pritchard, J. K. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet. 8, e1002967 (2012).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Bryant, D. & Moulton, V. Neighbor-net: an agglomerative method for the construction of phylogenetic networks. Mol. Biol. Evol. 21, 255–265 (2004).
Patterson, N., Moorjani, P., Luo, Y., Mallick, S., Rohland, N., Zhan, Y. et al. Ancient admixture in human history. Genetics 112, 1065–1093 (2012) ADMIXTOOLS 1.1 http://genetics.med.harvard.edu/reich/Reich_Lab/Software.html.
Teo, Y. Y., Sim, X., Ong, R. T., Tan, A. K., Chen, J., Tantoso, E. et al. Singapore Genome Variation Project: a haplotype map of three Southeast Asian populations. Genome. Res. 19, 2154–2162 (2009).
Saitou, N. A genetic affinity of human populations. Hum. Evol 10, 17–33 (1995).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Turner, C. G. II Late Pleistocene and Holocene population history of East Asia based on dental variation. Amer. J. Phys. Anthropol. 73, 305–321 (1987).
Turner, C. G. II Major features of sundadonty and sinodonty, including suggestions about East Asian microevolution, population history, and late Pleistocene relationships with Australian Aboriginals. Amer. J. Phys. Anthropol. 83, 295–317 (1990).
Yamaguchi, B. Establishment of Japanese (in Japanese), Misuzu Shobo, Tokyo, Japan, (1999).
Matsumura, H. Non-metric dental trait variation among local sites and regional groups of the Neolithic Jomon period, Japan. Anthropol. Sci. 115, 25–33 (2007).
Matsumura, H., Ishida, H., Amano, T., Ono, H. & Yoneda, M. Biological affinities of Okhotsk-culture people with East Siberians and Arctic people based on dental characteristics. Anthropol. Sci. 117, 121–132 (2009).
Hanihara, T. & Ishida, H. Regional difference in craniofacial diversity and the population history of Jomon Japan. Amer. J. Phys. Anthropol 139, 311–322 (2009).
Nakashima, A., Ishida, H., Shigematsu, M., Goto, M. & Hanihara, T. Nonmetric cranial variation of Jomon Japan: Implications for the evolution of Eastern Asian diversity. Amer. J. Hum. Biol. 22, 782–790 (2010).
Horai, S., Murayama, K., Hayasaka, K., Matsubayashi, S., Hattori, Y., Fucharoen, G. et al. mtDNA polymorphism in East Asian populations, with special reference to the peopling of Japan. Am. J. Hum. Genet. 59, 579–590 (1996).
Hammer, M. F. & Horai, S. Y Chromosomal DNA variation and the peopling of Japan. Am. J. Hum. Genet. 56, 951–962 (1995).
Tajima, A., Hayami, M., Tokunaga, K., Juji, T., Matsuo, M., Marzuki, S. et al. Genetic origins of the Ainu inferred from combined DNA analyses of maternal and paternal lineages. J. Hum. Genet. 49, 187–193 (2004).
Hammer, M. F., Karafet, T. M., Park, H., Omoto, K., Harihara, S., Stoneking, M. et al. Dual origins of the Japanese: common ground for hunter-gatherer and farmer Y chromosomes. J. Hum. Genet. 51, 47–58 (2006).
Nakagome, S., Sato, T., Ishida, H., Hanihara, T., Yamaguchi, T., Kimura, R. et al. The Asian DNA Repository Consortium. Model-based verification of hypotheses on the origin of modern Japanese revisited y Bayesian inference based on genome-wide SNP data. Mol. Biol. Evol. 32, 1533–1543 (2015).
Jinam, T. A., Kanzawa-Kiriyama, H. & Saitou, N. Human genetic diversity in the Japanese Archipelago: dual structure and beyond. Genes Genet. Syst. 60, 565–571 (2015).
Kozintsev, A. G. Ainu, Japanese, their ancestors and neighbors cranioscopic data. J. Anthropol. Soc. Nippon 98, 247–267 (1990).
Kozintsev, A. G. Prehistoric and recent populations of Japan: multivariate analysis of cranioscopic data. Arctic Anthropol. 29, 104–111 (1992).
Ishida, H. Metric and nonmetric cranial variation of the pre-historic Okhotsk people. Anthropol. Sci. 104, 233–258 (1996).
Komesu, A., Hanihara, T., Amano, T., Ono, H., Yoneda, M., Dodo, Y. et al. Nonmetric cranial variation in human skeleton remains associated with Okhotsk culture. Anthropol. Sci. 116, 33–47 (2008).
Sato, T., Amano, T., Ono, H., Ishida, H., Kodera, H., Matsumura, H. et al. Mitochondrial DNA haplogrouping of the Okhotsk people based on analysis of ancient DNA: an intermediate of gene flow from the continental Sakhalin people to the Ainu. Anthropol. Sci. 117, 171–180 (2009).
Kaburagi, M., Ushida, H., Goto, M. & Hanihara, T. Comparative studies of the Ainu, their ancestors, and neighbors: assessment based on metric and nonmetric dental data. Anthropol. Sci. 118, 95–106 (2010).
Kazuta, H., Sato, T., Dodo, Y., Matsumura, H., Amano, T., Ishisa, H. et al. Genotype frequencies of the ABCC11 gene in 2000–3000-year-old human bones from the Epi-Jomon and Jomon sites in Hokkaido, Japan. Anthropol. Sci. 119, 81–86 (2011).
Dodo, Y., Kawakubo, Y., Sawada, J. & Ishida, H. The Ainu and their neighbors as seen from the perspective of nonmetric cranial trait variation: I. The Hokkaido Ainu as a population among East and Northeast Asian peoples. (in Japanese) Anthropol. Sci. (Japanese series) 120, 1–13 (2012).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
He, Y., Wang, W. R., Xu, S. & Jin, L., Pan-Asia SNP Consortium Paleolithic contingent in modern Japanese: estimation and inference using genome-wide data. Sci. Rep 2, 355 (2012).
Reich, D., Patterson, N., Kircher, M., Delfin, F., Nandineni, M. R., Pugach, I. et al. Denisova admixture and the first modern human dispersals into Southeast Asia and Oceania. Am. J. Hum. Genet. 89, 516–528 (2011).
Acknowledgements
We thank Drs Y Shirakibara, T Baba and K Yanagihara of National Institute of Genetics for their kind permissions of using their experimental rooms, Mr T Kakuda of National Museum of Nature and Science for helping DNA extraction, Mrs M Mizuguchi for writing three perl scripts, and Mr I Babarinde and Ms Nilmini Hettiarachchi of Saitou Laboratory at National Institute of Genetics for helping computational analyses. We also thank Asian DNA Repository Consortium for use of the genome-wide SNP data for the Ainu population. This study was supported by a grant-in-aids of Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan to KS (25251043) and by SOKENDAI Strategic Interdisciplinary Research Project ‘Comprehensive study on creation of genetic and cultural diversity through modern human dispersal’ to NS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Kanzawa-Kiriyama, H., Kryukov, K., Jinam, T. et al. A partial nuclear genome of the Jomons who lived 3000 years ago in Fukushima, Japan. J Hum Genet 62, 213–221 (2017). https://doi.org/10.1038/jhg.2016.110
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2016.110
This article is cited by
-
Reconstructing the ancestral gene pool to uncover the origins and genetic links of Hmong–Mien speakers
BMC Biology (2024)
-
The Allen Ancient DNA Resource (AADR) a curated compendium of ancient human genomes
Scientific Data (2024)
-
Factors influencing cranial variation between prehistoric Japanese forager populations
Archaeological and Anthropological Sciences (2024)
-
Ancient Clostridium DNA and variants of tetanus neurotoxins associated with human archaeological remains
Nature Communications (2023)
-
Genomic analyses of 10,376 individuals in the Westlake BioBank for Chinese (WBBC) pilot project
Nature Communications (2022)