Abstract
Drug-induced liver injury (DILI) is the most common adverse drug reaction in the treatment of tuberculosis (TB). Several studies showed that patients with TB and the slow-acetylator phenotype caused by NAT2 variants are highly susceptible to DILI caused by anti-TB drugs, hereafter designated AT-DILI. However, the role of NAT2 variants in AT-DILI has never been assessed for an Indonesian population. We recruited 50 patients with TB and AT-DILI and 191 patients with TB but without AT-DILI; we then used direct DNA sequencing to assess single-nucleotide polymorphisms in the coding region of NAT2. NAT2*6A was significantly associated with susceptibility to AT-DILI (P=7.7 × 10−4, odds ratio (OR)=4.75 (1.8–12.55)). Moreover, patients with TB and the NAT2-associated slow-acetylator phenotype showed higher risk of AT-DILI than patients with the rapid- or intermediate-acetylator phenotypes (P=1.7 × 10−4, OR=3.45 (1.79–6.67)). In conclusion, this study confirms the significance of the association between slow-acetylator NAT2 variants and susceptibility to AT-DILI in an Indonesian population.
Similar content being viewed by others
Introduction
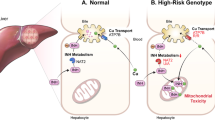
In 2013, Indonesia has the fifth highest incidence of tuberculosis (TB) in the world, with approximately 0.4–0.5 million people infected by pulmonary TB and a mortality rate of 25/100 000 people.1 The standard regimens for TB consist of isoniazid (INH), ethambutol, rifampicin and pyrazinamide.2 Despite their major contribution to the eradication of TB, first-line anti-tuberculosis (Anti-TB) drugs, particularly INH, rifampicin and pyrazinamide, are associated with hepatotoxicity and drug-induced liver injury (DILI) in patients undergoing treatment with these drugs.3 DILI is characterized by an increase in liver function, for example, aspartate aminotransferase, alanine transaminase and total bilirubin levels increase to more than twice the normal upper-limit levels.4 DILI caused by anti-TB drugs (hereafter called AT-DILI) is the most common adverse drug reaction in the treatment of TB, and AT-DILI frequently results in TB treatment failure because of treatment interruption.3 AT-DILI reactions are unpredictable and often unrelated to the treatment dose; therefore, it is reasonable to assume that genetic factors have a role in these adverse reactions.5 Although physiological, environmental and other non-genetic factors such as sex and age may also influence idiosyncratic DILI reactions, recent progress in the field and specifically a report on flucloxacillin (HLA-B*5701) indicate that genetic factors may have a bigger role in apparently idiosyncratic and non-dose-related DILI reactions.6 In the case of AT-DILI, genetic variations in genes that code for drug transporters and metabolizing enzymes are thought to cause irregular processing rate of toxic Anti-TB drug intermediates, such as hydrazine, which then increase the risk of AT-DILI.5
Several genes are associated with increased risk to AT-DILI; these include CYP2E1,7, 8 MnSOD,9 GSTM1,9, 10, 11, 12 GSTT1,13 HLA-DQA1,14 HLA-DQB114 and NAT2.14, 15 Each of these gene products, with the exception of the HLA gene products, is involved in the processing of anti-TB drugs in the liver. NAT2 has been previously studied in association with AT-DILI, and certain NAT2 variants are strongly associated with AT-DILI caused by INH.15, 16
NAT2 is located on chromosome 8p22 and encodes an acetyltransferase enzyme called arylamine N-acetyltransferase, which has an important role in acetylation of hydrazine-based drugs such as INH and in the deactivation of carcinogens in the liver. Several NAT2 variants affect NAT2-encoded enzymatic activity, specifically substrate acetylation rate.17 Based on this acetylation rate, NAT2 phenotypes can be categorized into two (bimodal distribution) or three (trimodal distribution) groups; the bimodal distribution comprises rapid and slow groups; the trimodal distribution comprises rapid, intermediate and slow groups. Slow acetylation of INH results in increased plasma levels of the drug, which may increase the toxicity of the drug.15, 18 Several studies have shown that the slow-acetylator NAT2 phenotype is strongly associated with a higher risk of AT-DILI consistently across different ethnic groups.15, 16, 19, 20, 21, 22
A previous study on NAT2 in individuals within the Javanese and Sundanese ethnic groups of Indonesia reveals that slow-acetylator NAT2 variants are frequently observed in the studied population.23 However, to our knowledge, no published study examines the association between NAT2 genotypes and AT-DILI in the population. Therefore, this study was designed to explore the relationship between NAT2 genotypes and susceptibility to AT-DILI in the Indonesian population. If there is a clear link between the NAT2 genotypes and AT-DILI incidence in the Indonesian population, NAT2 genotyping might be a useful tool for predicting patient susceptibility to AT-DILI before initiating anti-TB treatments.
Materials and Methods
Recruitment of Subjects
In this study, total of 50 unrelated pulmonary patients with TB and AT-DILI were recruited from Pasar Rebo Hospital, East Jakarta, Indonesia, and 191 unrelated pulmonary patients with TB and normal liver function were recruited from various TB clinics in Banten and West Java, Indonesia. Both sets of patients, those with and without AT-DILI, were of the same ethnicity. Whole-blood samples (5 ml each) were collected from each patient into an EDTA tube and stored in a −20 °C freezer until further processing. Patients included in the study were male and female, ranged in age from 15 to 70 years old, suffered from TB according to the World Health Organizatio standards, and were under monitored treatment with anti-TB drugs; patients with AT-DILI were classified as cases and those without AT-DILI were classified as controls. The ethnic backgrounds of patients in both groups were Javanese, Sundanese and Betawi. Patient exclusion criteria were as follows: (1) history of liver disease such as: hepatitis A B, or C; hepatoma; liver cirrhosis or cholelithiasis positive or (2) abnormal levels of any liver function test (alanine transaminase, aspartate aminotransferase or total bilirubin) before anti-TB treatment.
Single-nucleotide polymorphism (SNP) genotyping
QIAamp DNA blood mini kits (Qiagen, Hilden, Germany) were used according to the manufacturer’s instructions to extract genomic DNA from each blood sample. A Qubit 2.0 Fluorometer (Invitrogen, Carlsbad, CA, USA) was used to assess DNA quantity before SNP genotyping. PCR was performed to amplify the coding region of NAT2 on a Sensoquest Labcycler Thermocycler (Sensoquest, Gottingen, Germany); each 25-μl reaction mixture contained 20 ng of genomic DNA, 1 × of FastStart 10 × PCR Buffer with 20 mM MgCl2 (Roche Applied Science, Penzberg, Germany), 200 μM of dNTPs, 0.2 μM of each primers (Forward and Reverse) and 1 U of FastStart Taq DNA polymerase (Roche Diagnostics, Mannheim, Germany). PCR conditions were as follow: initial denaturation at 96 °C for 4 min; 40 cycles of denaturation at 96 °C for 30 s, annealing at 55 °C for 30 s and elongation at 72 °C for 30 s; and a final elongation step at 72 °C for 5 min. Forward and reverse primers used for PCR and DNA sequencing were based on a previously published report.23 Direct DNA sequencing was used to genotype each of six NAT2 SNPs. Cycle sequencing was performed using BigDYE terminator v3.1. (Applied Biosystems, Foster City, CA, USA) on a Sensoquest Labcycler Thermocycler (Sensoquest, Gottingen, Germany) according to the manufacturer’s instruction. Cycle sequencing products were run through a ABI 310 Genetic Analyzer (Applied Biosystems) to sequence the specific NAT2 fragments that contained the following SNPs: rs1041983 [282C>T], rs1801280 [341T>C], rs1799929 [481C>T], rs1799930 [590G>A], rs1208 [803A>G] and rs1799931 [857G>A].23
Data analysis
NAT2 SNP genotypes were extracted from sequencing trace files using NovoSNP.24 Haplotypes that each comprised a combination of alleles of six genotyped SNPs were determined for each sample. Haplotype reconstruction was performed using PHASE v2.1.1 software25, 26 and confirmed by Haploview software.27 NAT2 genotypes and acetylator phenotypes were inferred from the reconstructed haplotype by comparisons to the human NAT2 allele database at http://nat.mbg.duth.gr/HumanNAT2alleles_2013.htm. The χ2 test was used to compare between cases and controls with regard to each variable and to test for Hardy–Weinberg equilibrium. Odds ratios (ORs) and 95% confidential intervals were calculated using conditional logistic regression. Any two-tailed P-value of <0.05 was considered statistically significant.
Results
The distribution of SNP genotypes among the 241 TB patients was consistent with Hardy–Weinberg equilibrium (Table 1). Statistical analysis of the six SNPs assessed showed that the minor allele of rs1041983 was significantly more prevalent among cases, TB patients with AT-DILI, than among controls (P=0.0023). Moreover, the minor allele of rs1799930 also showed a modest association with AT-DILI (P=0.0222). Based on the NAT2 haplotype reconstruction, we identified eight NAT2 haplotype alleles each comprising the six SNPs (Table 2). The most frequent NAT2 haplotypes among all 241 patients were CTCGAG (NAT*4), TTCAAG (NAT2*6A), TTCGAA (NAT2*7B) and CCTGGG (NAT2*5B), with frequencies of 0.35, 0.35, 0.15 and 0.13, respectively. According to the official human NAT2 database (http://nat.mbg.duth.gr/HumanNAT2alleles_2013.htm), NAT2*4, NAT2*12A and NAT2*13 are classified as rapid-acetylator alleles, whereas NAT2*5B, NAT2*5C, NAT2*6A, NAT2*7A and NAT2*7B are classified as slow-acetylator alleles. Further analysis showed that the frequency of NAT2*6A was significantly higher among patients with TB and AT-DILI than among those with only TB (P=7.7 × 10−4, OR=4.8 (1.8–12.55); Table 2). Conversely, NAT2*4 was significantly higher among controls, patients with TB but without AT-DILI, than among cases (P=1.8 × 10−6, OR=0.2 (0.1–0.39)). Using a trimodal distribution model of NAT2 acetylator phenotype, the frequencies of the rapid-, intermediate- and slow-acetylator NAT2 phenotypes among patients with TB and AT-DILI (cases) versus those with only TB (controls) were 8% (cases) vs 15.2% (controls), 28% vs 50.8% and 64% vs 34%, respectively (Table 3). We further analyzed the association between NAT2 acetylator phenotypes and predisposition to AT-DILI and discovered that the slow-acetylator phenotype was more frequent among patients with TB and AT-DILI than among those with only TB; moreover, the slow-acetylator phenotype was a significant risk factor for susceptibility to AT-DILI among patients with TB (P=1.7 × 10−4, OR=3.45 (1.8–6.7); Table 4).
Discussion
Associations between NAT2 genetic variants and the risk of AT-DILI have been widely studied recently. Numerous studies have investigated the allele frequencies of NAT2 variants in major ethnic groups in the world;28, 29 however, none of these studies has assessed associations between NAT2 genotypes and AT-DILI in the Indonesian population. In this study, we investigated the distribution of NAT2 SNP genotypes among 241 patients with TB from Javanese, Sundanese and Betawi ethnic groups. All variants were in Hardy–Weinberg equilibrium (Table 1). We identified eight NAT2 haplotypes (Table 2) and analyzed associations between NAT2 acetylator phenotypes and predisposition to AT-DILI. We found that the frequency of NAT2*6A, which was categorized as a slow-acetylator haplotype, was significantly higher among patients with TB and AT-DILI than among those with only TB (P=7.7 × 10−4, OR=4.8 (1.8–12.55); Table 2). Conversely, NAT2*4 was significantly more common among patients with only TB than among those with both TB and AT-DILI (P=1.8 × 10−6, OR=0.2 (0.1–0.39); Table 2). Furthermore, we also analyzed the data based on the predicted NAT2 phenotype and found that the slow-acetylator phenotype was significantly more common among patients with both TB and AT-DILI (Tables 3 and 4).
The results of this study showed that NAT2 acetylator status, as determined based on SNP haplotype analysis, was a significant risk predictor of AT-DILI in this Indonesian population, which comprised the Javanese, Sundanese and Betawi ethnic groups. Moreover, to our knowledge, this study is the first to examine genetic associations with AT-DILI in an Indonesian population. To further confirm the results of our study, a replication of this study that employs larger case and control samples and that incorporates other ethnic groups of Indonesia (for example, Melayu (Malay), Batak, Bugis and so on) should be pursued to generate NAT2 data for this multi-ethnic country. We found two NAT2 haplotype alleles (NAT2*5C and NAT*7A) that are not identified and reported in the Javanese, Sundanese and Betawi ethnic.23 These alleles were very rare in this study of 241 patients with TB. In our previous study with the Javanese and Sundanese ethnic groups that involve only a healthy control sample, we only found six NAT2 haplotypes: NAT2*4, NAT2*5B, NAT2*6A, NAT2*7B, NAT2*12A and NAT2*13.23
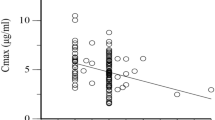
A visualization of the trend in the association between OR for AT-DILI and the four most frequent NAT2 haplotypes observed in this study is shown in Figure 1. This trend showed NAT2*6A and NAT2*4 on opposing ends of the trend line; NAT2*6A was associated with higher risk of AT-DILI and NAT2*4 was associated with protection from AT-DILI based on the respective ORs. Previous studies of in vitro NAT2 enzyme activity reveal that the degree of acetylation activity varies among NAT2 alleles, with NAT2*4 displaying the highest activity and NAT2*5 displaying the lowest; NAT2*6 and NAT2*7 show activities between NAT2*4 and NAT2*5.30 However, if lower NAT2 acetylator activity were associated with higher risk of AT-DILI, our results would be inconsistent with the in vitro activity data. The discrepancies between in vitro activity data and our population-genetics findings may stem from the fact that in our study both NAT2*5 and NAT2*7 were found in far lower frequencies than NAT2*6 and NAT2*4. Notably, The NAT2*6A allele significantly predicts predisposition to AT-DILI in Taiwanese, Japanese and Chinese.15, 31, 32 However, study of Caucasian populations showed a different result.33, 34 Therefore, a meta-analysis of these data would be worthwhile for determining whether NAT2 slow-acetylators alleles are associated with a higher risk of AT-DILI especially in Southeast Asian populations. Previous meta-analyses show that NAT2 slow-acetylator alleles are associated with a higher risk of AT-DILI across different populations.33, 34, 35 Significant results were also found for populations in East Asian, South Asian, Brazilians and Middle Easterns, whereas data on Caucasians did not show a similar result.33, 35 Unfortunately, the meta-analysis studies did not include data from Southeast Asian populations, including the Indonesian population that contributes to the fifth largest incidence of TB in the world. Thus, our study results will be important for addressing AT-DILI in this important Southeast Asian population.
The odds ratio (OR) between the most frequent NAT2 haplotypes. NAT2*6A (OR=4.75; 95% confidence interval (CI)=1.80–12.55) classified as slow-acetylator allele showed highest risk of susceptibility to AT-DILI, whereas NAT2*4 (OR=0.20; 95% CI=0.11–0.39) classified as rapid-acetylator allele showed strongest protection against AT-DILI. The trend line is shown in round dot line.
Functional characterization of NAT2 SNPs and NAT2 haplotypes by Zhang et al.36 shows that the SNPs in haplotype with slow-acetylator phenotypes reduce the sulfamethazine N-acetylation activity. The biochemical changes that reduce NAT2 enzyme activity vary between the SNPs, but the finding is consistent with the slow-acetylator phenotype being secondary to a reduction in NAT2 protein levels. The NAT2*6 haplotype, [590G>A], either alone or in combination with other SNPs, reduces the thermostability and the level of soluble NAT2 protein, whereas the tag SNP for alleles in NAT2*5, [341T>C], apparently causes different changes that result in the slow-acetylator phenotype. A proposed mechanism based on the Zhang et al.37 study involves enhancement of protein degradation as the primary mechanism for reduced NAT2 protein and activity.
Differences in the distribution of NAT2 genotypes among ethnic groups reflect the differences in the standard anti-TB drug dosages used among these groups; hence, our current study provides additional evidence to support the administration of individualized INH dosages in the Indonesian population. Individuals with a homozygous wild-type NAT2*4 genotype, which is predicted to result in a rapid-acetylator NAT2 phenotype, are reportedly relatively immune to AT-DILI with the standard dosage given to Indonesian (300 mg per day).31, 38 Moreover, due to the effectiveness of the acetylating property of the NAT2 enzyme, individuals with this genotype may benefit from increased anti-TB treatment dose.39 Conversely, the standard INH dosage may be excessive for individuals with a homozygous slow-acetylator genotype; therefore, a review of an individual’s NAT2 genotype may be useful when determining the appropriate INH dosage.
Complex diseases such as DILI are known to be affected by multiple genetic variants.34, 40 Our candidate-gene analysis showed results consistent with results from studies of other groups. Although NAT2 variants have consistently exhibited significant associations with AT-DILI, the notion that NAT2 is the only genetic factor that affects DILI underestimates the complexity of this adverse drug reaction. Therefore, further genome-wide association studies may assist in elucidating other genetic variants important to AT-DILI.41, 42
In conclusion, this study confirms the importance of NAT2 acetylator status in providing risk assessment of AT-DILI for patients with TB; notably, we found that the slow-acetylator type NAT2*6A haplotype was especially relevant in this Indonesian population. However, this encouraging study needs to be replicated with larger samples that include other major Indonesian ethnic groups to reflect the ethnic diversity of Indonesia. A positive replication of our findings could lead to routine implementation of NAT2 genotype-guided dosing in the Indonesian population. Furthermore, a meta-analysis should be conducted with data from Southeast Asian populations to clarify the relationship between NAT2 variants and AT-DILI risk in this group of closely related populations.
References
WHO Global Tuberculosis Report 2014, (World Health Organization, Geneva, Switzerland, 2014).
WHO. Treatment of Tuberculosis: Guidelines, (World Health Organization, Geneva, Switzerland, 2010).
Tostmann, A., Boeree, M. J., Aarnoutse, R. E., de Lange, W. C., van der Ven, A. J. & Dekhuijzen, R. Antituberculosis drug-induced hepatotoxicity: concise up-to-date review. J. Gastroen. Hepatol. 23, 192–202 (2008).
Bénichou, C. Criteria of drug-induced liver disorders. Report of an international consensus meeting. J. Hepatol. 11, 272–276 (1990).
Saukkonen, J. J., Cohn, D. L., Jasmer, R. M., Schenker, S., Jereb, J. A., Nolan, C. M. et al. An official ATS statement: hepatotoxicity of antituberculosis therapy. Am. J. Resp. Crit. Med. 174, 935–952 (2006).
Daly, A. K., Donaldson, P. T., Bhatnagar, P., Shen, Y., Pe'er, I., Floratos, A. et al. HLA-B* 5701 genotype is a major determinant of drug-induced liver injury due to flucloxacillin. Nat. Genet. 41, 816–819 (2009).
Huang, Y. S., Chern, H. D., Su, W. J., Wu, J. C., Chang, S. C., Chiang, C. H. et al. Cytochrome P450 2E1 genotype and the susceptibility to antituberculosis drug‐induced hepatitis. Hepatology 37, 924–930 (2003).
Vuilleumier, N., Rossier, M. F., Chiappe, A., Degoumois, F., Dayer, P., Mermillod, B. et al. CYP2E1 genotype and isoniazid-induced hepatotoxicity in patients treated for latent tuberculosis. Eur. J. Clin. Pharmacol. 62, 423–429 (2006).
Huang, Y.-S., Su, W.-J., Huang, Y.-H., Chen, C.-Y., Chang, F.-Y., Lin, H.-C. et al. Genetic polymorphisms of manganese superoxide dismutase, NAD (P) H: quinone oxidoreductase, glutathione S-transferase M1 and T1, and the susceptibility to drug-induced liver injury. J. Hepatol. 47, 128–134 (2007).
Li, C., Long, J., Hu, X. & Zhou, Y. GSTM1 and GSTT1 genetic polymorphisms and risk of anti-tuberculosis drug-induced hepatotoxicity: an updated meta-analysis. Eur. J. Clin. Microbiol. 32, 859–868 (2013).
Roy, B., Chowdhury, A., Kundu, S., Santra, A., Dey, B., Chakraborty, M. et al. Increased risk of antituberculosis drug-induced hepatotoxicity in individuals with glutathione S-transferase M1 'null' mutation. J. Gastroen. Hepatol. 16, 1033–1037 (2001).
Tang, N., Deng, R., Wang, Y., Lin, M., Li, H., Qiu, Y. et al. GSTM1 and GSTT1 null polymorphisms and susceptibility to anti-tuberculosis drug-induced liver injury: a meta-analysis. Int. J. Tuberc. Lung Dis. 17, 17–25 (2013).
Ambreen, K., Sharma, R., Singh, K. P., Abbas, M. & Kumar, S. Association of GSTM1, GSTT1 and CYP2E1 Gene Polymorphisms with Antituberculosis Drug Induced Hepatotoxicity in North Indian Population. IJHSR 4, 149–160 (2014).
Sharma, S. K., Balamurugan, A., Saha, P. K., Pandey, R. M. & Mehra, N. K. Evaluation of clinical and immunogenetic risk factors for the development of hepatotoxicity during antituberculosis treatment. Am. J. Respir. Crit. CAre Med. 166, 916–919 (2002).
Huang, Y.S., Chern, H. D., Su, W. J., Wu, J. C., Lai, S. L., Yang, S. Y. et al. Polymorphism of the N‐acetyltransferase 2 gene as a susceptibility risk factor for antituberculosis drug–induced hepatitis. Hepatology 35, 883–889 (2002).
Ohno, M., Yamaguchi, I., Yamamoto, I., Fukuda, T., Yokota, S., Maekura, R. et al. Slow N-acetyltransferase 2 genotype affects the incidence of isoniazid and rifampicin-induced hepatotoxicity. Int. J. Tuberc. Lung Dis. 4, 256–261 (2000).
Sim, E., Payton, M., Noble, M. & Minchin, R. An update on genetic, structural and functional studies of arylamine N-acetyltransferases in eucaryotes and procaryotes. Hum. Mol. Genet 9, 2435–2441 (2000).
Stanley, L. A. & Sim, E. Update on the pharmacogenetics of NATs: structural considerations. Pharmacogenomics 9, 1673–1693 (2008).
Cho, H.-J., Koh, W.-J., Ryu, Y.-J., Ki, C.-S., Nam, M.-H., Kim, J.-W. et al. Genetic polymorphisms of NAT2 and CYP2E1 associated with antituberculosis drug-induced hepatotoxicity in Korean patients with pulmonary tuberculosis. Tuberculosis 87, 551–556 (2007).
Khalili, H., Fouladdel, S., Sistanizad, M., Hajiabdolbaghi, M. & Azizi, E. Association of N-acetyltransferase-2 genotypes and anti-tuberculosis induced liver injury; first case-controlled study from Iran. Curr. Drug Saf. 6, 17–22 (2011).
Lee, S., Chung, L., Huang, H., Chuang, T., Liou, Y. & Wu, L. NAT2 and CYP2E1 polymorphisms and susceptibility to first-line anti-tuberculosis drug-induced hepatitis. Int. J. Tuberc. Lung Dis. 14, 622–626 (2010).
Rana, S. V., Ola, R. P., Sharma, S. K., Arora, S. K., Sinha, S. K., Pandhi, P. et al. Comparison between acetylator phenotype and genotype polymorphism of n-acetyltransferase-2 in tuberculosis patients. Hepatol. Int. 6, 397–402 (2012).
Yuliwulandari, R., Sachrowardi, Q., Nishida, N., Takasu, M., Batubara, L., Susmiarsih, T. P. et al. Polymorphisms of promoter and coding regions of the arylamine N-acetyltransferase 2 (NAT2) gene in the Indonesian population: proposal for a new nomenclature. J. Hum. Genet. 53, 201–209 (2008).
Weckx, S., Del-Favero, J., Rademakers, R., Claes, L., Cruts, M., De Jonghe, P. et al. novoSNP, a novel computational tool for sequence variation discovery. Genome Res. 15, 436–442 (2005).
Stephens, M. & Scheet, P. Accounting for decay of linkage disequilibrium in haplotype inference and missing-data imputation. Am. J. Hum. Genet. 76, 449–462 (2005).
Stephens, M., Smith, N. J. & Donnelly, P. A new statistical method for haplotype reconstruction from population data. Am. J. Hum. Genet. 68, 978–989 (2001).
Barrett, J. C., Fry, B., Maller, J. & Daly, M. J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Rabstein, S., Unfried, K., Ranft, U., Illig, T., Kolz, M., Rihs, H. P. et al. Variation of the N-acetyltransferase 2 gene in a Romanian and a Kyrgyz population. Cancer Epidemiol. Biomarkers Prev. 15, 138–141 (2006).
Sabbagh, A., Langaney, A., Darlu, P., Gérard, N., Krishnamoorthy, R. & Poloni, E. S. Worldwide distribution of NAT2 diversity: implications for NAT2 evolutionary history. BMC Genet. 9, 21 (2008).
Hein, D. W., Doll, M. A., Rustan, T. D. & Ferguson, R. J. Metabolic activation of N-hydroxyarylamines and N-hydroxyarylamides by 16 recombinant human NAT2 allozymes: effects of 7 specific NAT2 nucleic acid substitutions. Cancer Res. 55, 3531–3536 (1995).
Higuchi, N., Tahara, N., Yanagihara, K., Fukushima, K., Suyama, N., Inoue, Y. et al. NAT2 6A, a haplotype of the N-acetyltransferase 2 gene, is an important biomarker for risk of anti-tuberculosis drug-induced hepatotoxicity in Japanese patients with tuberculosis. World J. Gastroenterol. 13, 6003–6008 (2007).
An, H. R., Wu, X. Q., Wang, Z. Y., Zhang, J. X. & Liang, Y. NAT2 and CYP2E1 polymorphisms associated with antituberculosis drug-induced hepatotoxicity in Chinese patients. Clin. Exp. Pharmacol. Physiol. 39, 535–543 (2002).
Cai, Y., Yi, J. Y., Zhou, C. H. & Shen, X. Z. Pharmacogenetic study of drug metabolising enzyme polymorphisms on the risk of anti-tuberculosis drug-induced liver injury: a meta-analysis. PLoS ONE 7, e47769 (2012).
Wang, P. Y., Xie, S. Y., Hao, Q., Zhang, C & Jiang, B. F. NAT2 polymorphisms and susceptibility to anti-tuberculosis drug-induced liver injury: a meta-analysis. Int. J. Tuberc. Lung Dis. 16, 589–595 (2012).
Du, H., Chen, X., Fang, Y., Yan, O., Xu, H., Li, L. et al. Slow N-acetyltransferase 2 genotype contributes to anti-tuberculosis drug-induced hepatotoxicity: a meta-analysis. Mol. Biol. Rep. 40, 3591–3596 (2013).
Zhang, Y., Doll, M. A., Zhao, S., States, J.C. & Hein, D. W. Functional characterization of single-nucleotide polymorphism and haplotypes of human N-acetyltransferase 2. Carcinogenesis 28, 1665–1671 (2007).
Zhang, Y., Doll, M. A., Zhao, S., States, J. C. & Hein, D. W. Functional Characterization of the A411T (L137F) and G364A (D122N) genetic polymorphism in human N-acetyltransferase 2. Pharmacogent. Genomics 1, 37–45 (2007).
Kementrian Kesehatan Republik Indonesia: Direktorat Jendral Pengendalian Penyakit dan Penyehatan Lingkungan. Pedoman Pengendalian Tuberkulosis, (Kementrian Kesehatan Republik Indonesia, Jakarta, Indonesia, 1994).
Azuma, J., Ohno, M., Kubota, R., Yokota, S., Nagai, T., Tsuyuguchi, K. et al. NAT2 genotype guided regimen reduces isoniazid-induced liver injury and early treatment failure in the 6-month four-drug standard treatment of tuberculosis: a randomized controlled trial for pharmacogenetics-based therapy. Eur. J. Pharmacol. 69, 1091–1101 (2013).
Stephens, C., Lucena, M. I. & Andrade, R. J. Genetic variations in drug-induced liver injury (DILI): resolving the puzzle. Front. Genet. 3, 253 (2012).
Ma, Q. & Lu, A. Y. H. Pharmacogenetics, pharmacogenomics, and individualized medicine. Pharmacol. Rev. 63, 437–459 (2011).
Urban, T. J., Goldstein, D. B. & Watkins, P. B. Genetic basis of susceptibility to drug-induced liver injury: what have we learned and where do we go from here? Pharmacogenomics 13, 735–738 (2012).
Acknowledgements
This project was funded by a grant from the Indonesian Directorate General of Higher Education (DIKTI) of the Ministry of Higher Education, Research and Technology of the Republic of Indonesia. We are also thankful to the following institutions/organizations for their support in this project: YARSI Foundation, Pasar Rebo General Hospital, and YARSI Genomic Medicine Research Group. Last, special appreciation goes to our TB collaborators under JSPS (Japan Society for the Promotion of Science) core-to-core program and all TB patients who participated in this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Yuliwulandari, R., Susilowati, R., Wicaksono, B. et al. NAT2 variants are associated with drug-induced liver injury caused by anti-tuberculosis drugs in Indonesian patients with tuberculosis. J Hum Genet 61, 533–537 (2016). https://doi.org/10.1038/jhg.2016.10
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2016.10
This article is cited by
-
Risk adjustment model for tuberculosis compared to non-tuberculosis mycobacterium or latent tuberculosis infection: Center for Personalized Precision Medicine of Tuberculosis (cPMTb) cohort database
BMC Pulmonary Medicine (2023)
-
The rs1801280 SNP is associated with non-small cell lung carcinoma by exhibiting a highly deleterious effect on N-acetyltransferase 2
Journal of Cancer Research and Clinical Oncology (2023)
-
Genetic Counselling: the biomedical bridge between molecular diagnosis and precision treatment
Egyptian Journal of Medical Human Genetics (2022)
-
Isonicotinylation is a histone mark induced by the anti-tuberculosis first-line drug isoniazid
Nature Communications (2021)
-
The atorvastatin metabolic phenotype shift is influenced by interaction of drug-transporter polymorphisms in Mexican population: results of a randomized trial
Scientific Reports (2020)