Abstract
Disease risk prediction (DRP) is one of the most important challenges in personal genome research. Although many direct-to-consumer genetic test (DTC) companies have begun to offer personal genome services for DRP, there is still no consensus on what constitutes a gold-standard service. Here, we systematically evaluated the distributions of DRPs from three DTC companies, that is, 23andMe, Navigenics and deCODEme, for 22 diseases using three Japanese samples. We systematically quantified and analyzed the differences between each DTC company’s DRPs. Our independency test showed that the overall prediction results were correlated with each other, but not perfectly matched; less than onethird mismatching of the opposite direction occurred in eight diseases. Moreover, we found that the differences could mainly be attributed to four factors: (1) single nucleotide polymorphism (SNP) selection, (2) average risk estimation, (3) the disease risk calculation algorithm and (4) ethnicity adjustment. In particular, only 7.1% of SNPs over 22 diseases were reviewed by all three companies. Therefore, development of a universal core SNPs list for non-Caucasian samples will be important for achieving better prediction capacity for Japanese samples. This systematic methodology provides useful insights for improving the capacity of DRPs in future personal genome services.
Similar content being viewed by others
Introduction
Disease risk prediction (DRP) using an individual’s genome has attracted much attention because of the availability of low-cost high-throughput genotyping and whole-genome sequencing technologies.1, 2, 3, 4, 5, 6, 7 Several companies offer such services to evaluate an individual’s disease risks.8, 9, 10, 11 However, a study demonstrated that only 50% or fewer of the predictions of two direct-to-consumer genetic test (DTC) companies agreed across five individuals for seven diseases.12 Assessing DRPs of personal genome services is controversial and little is known about their real predictive capacities.13 Therefore, it is important to examine and understand the basis for such differences, especially as more people use such services. Most DRPs by DTC companies are based on genome-wide association study (GWAS) research in the Caucasian population; clearly, these predictions need to be adapted and/or modified when interpreting genotyping or sequence data from individuals from other ethnicities. It is also important to evaluate the predictive capacity of such services in non-Caucasians and to understand the basic mechanisms that explain general properties of DRPs.
Here, we report our observations and systematic interpretations of disease risk distributions by three DTC companies for three Japanese individuals. This is perhaps the first attempt at such analysis for Japanese individuals. We introduced methods for concordance rate (CR) analyses, independency tests and risk distribution analyses to quantify the concordances and discrepancies of DRPs. Through these methods, we quantified the degree of mismatches of the predictions and characterized the distribution of relative risk (RR) for each disease. We then investigated why discrepancies occurred for each of four factors, that is, single nucleotide polymorphism (SNP) selection, average risk, risk prediction algorithms and ethnicity adjustment. We hypothesized that utilizing a universal core SNPs list for non-Caucasians may allow for better predictions. In addition, we suggested the current capacity of DRPs of personal genome services for Japanese individuals and provide recommendations for future improvement of DRPs.
Materials and methods
Study subjects and samples
We investigated the distributions of DRPs for three Japanese samples using three DTC companies: 23andMe, Navigenics and deCODEme. For deCODEme, we used the Japanese customized service provided by Bioinfovision (Tokyo, Japan). First, we collected three saliva samples from healthy Japanese volunteers and tested them using the above-listed DTC companies. Then, we compared the relative disease risks reported by the DTC services over 22 diseases using the same criteria described by Ng et al.12 We explained our research purposes, protocols and potential risks of knowing one’s genetic risks to each volunteer, and received informed consent from them. Genetic counselors were provided for unforeseen situations. Our study protocol was approved by the Institutional Review Board of Rikengenesis.
Analysis of concordance and disease risk distribution
We attempted to validate the report by Ng et al.12 to understand the degree of inconsistency between the three reports. We used the methodology of Ng et al.12 for 22 diseases and three DTC companies, and to evaluate the consistency between the predictions from the three DTC companies. We applied the 5% threshold of RR to define increased/decreased risks (where 0.95⩽RR⩽1.05 was considered to be no risk). To mathematically quantify the consistency between the predictions, we calculated the CRs between all pairwise combinations among the three companies. The CR was calculated using the following formula:

where AD is the number of all test data, CD is the number of concordant test data and MD is the number of mismatched data (not concordant data).
We omitted NA (unavailable) data from AD (only ↑, ↓ and = were considered) and calculated the two types of CRs: counting ↑=, ↓= as mismatched data or counting them as concordant data (ignoring the mismatch with average risk [=]). We compared the CRs for all pairwise combinations of predictions from the three DTC companies. We also conducted 2 × 2 Fisher’s exact tests for each set of two DTC companies’ risk predictions for testing independence. We generated 2 × 2 contingency tables, where the rows represented company A’s test results (↑, ↓) and the columns represented company B’s results (↑, ↓). We counted the number of ↑↑, ↑↓, ↓↑ and ↓↓ combinations and conducted 2 × 2 Fisher’s exact tests using the R function ‘fisher.test()’. P-values were calculated for each set of two DTC companies’ DRPs. We compared similarities among the three DTC companies’ DRPs. We used the κ-statistic to analyze the agreement of DRPs. We conducted Cohen’s κ-test for each set of two DTC companies’ DRPs using the R function ‘Kappa.test()’. We also conducted Fleiss κ-tests for all three companies’ DRPs using the R function ‘Kappam.fleiss()’.
Next, we observed the distributions of RRs and examined the relationships between the number of increasing risks (higher risk than average, ↑) and decreasing risks (lower risk than average, ↓). We visualized DRPs by plotting minimum, maximum and average values of RRs for each of the three samples for 22 disease conditions. We calculated the average RRs of the three samples for each of the DTC companies. On the basis of our observations, we hypothesized that the number of individuals who had higher than average risk was larger than those who had lower than average risk. To validate this hypothesis, we calculated the average RRs where RR>1 and RR<1 for each DTC company’s prediction and compared their variation.
Causal analyses for discrepancy
To investigate the origin of the observed discrepancies, we followed methods reported in previous research by Swan10 and assumed that variance in multigenic risk interpretation could be mainly explained by differences in (1) the SNPs selected for the assessment, (2) the average risk and (3) the risk assignment methodologies. We added one additional factor, (4) the method for ethnicity adjustment, and evaluated each of the four factors for the three DTC companies for three volunteers. The reasons for their discrepancies were then classified.
SNP selections
We counted how many SNP markers were used for each of the 22 diseases and analyzed overlapping SNPs (hereafter named as ‘core SNPs’ as overlapping SNPs reviewed by more than three organizations) among 23andMe, Navigenics and deCODEme. We also counted overlaps and consistency of references for core SNPs among the three companies. We compared each company’s annotated information of core SNPs, such as P-values, odds ratios (ORs), number of samples and references. In addition, we examined the core SNPs reviewed by the NIH GWAS catalog and four companies, that is, deCODEme, Navigenics, 23andMe and Pathway Genomics, for type 2 diabetes (T2DM). We also examined the statistics of reference citations for core SNPs in T2DM.
Average risk
We compared the average risks provided by each of the companies for 22 diseases. Average risk was estimated from epidemiologic literature on average population disease risk. DTC companies may assign different average risks because they have selected different research studies as being the most representative and reliable.10 Publicly available information from each company’s website and Supplementary Information at technical conferences and informal interviews were gathered. 23andMe and deCODEme provided the average risks for East Asians in certain diseases, and Navigenics provided the average risks only for Caucasians. Although the data on East Asians were limited, we compared the influence of ethnicity differences between Caucasians and East Asians for available disease conditions.
Risk prediction algorithms
We reviewed the descriptions of the disease prediction algorithms provided in the white papers from the three companies and implemented each algorithm using R software to compare them. We defined the standard mathematical notations to represent each company’s algorithm for their systematic comparisons. Mathematical formulas of risk prediction algorithms for each DTC company are described in Supplementary Note 1. The disease prediction algorithms for each of the three companies were similar; however, they were not identical. The three companies reported absolute risk, that is, the probability that an individual will develop a disease. Absolute risk was derived from two parameters: RR and average risk. Each of the three companies required three critical parameters for risk prediction: risk allele frequency (RAF; p), average risk (q) and OR (r). Although two different ORs were used when available, most references did not describe two such ORs for a trait concerning a locus. The effects of a risk allele were interpreted to be additive on the log-transformed OR, and only a single OR was often used. These parameters were determined by literature selected according to the policy of each company. The probability of developing the disease (absolute risk) of each of genotype AA, Aa and aa was represented by d1, d2 and d3, respectively, and absolute risks (d1, d2 and d3) were calculated by the parameters (p, q and r) obtained by solving the system of equations described in Supplementary Note 1. Even if they choose the same parameter sets for p, q and r, their DRPs (d1, d2 and d3) may differ slightly because of differences between prediction algorithms. We investigated the degree of differences by calculating the DRP for each company with the same parameter settings (p, q and r).
Ethnicity adjustment
We compared the differences in policies and methods for ethnicity adjustments among the three companies and examined how different parameter sets (p, q and r) for different ethnicities affected DRPs (d1, d2 and d3). In addition, we compared the RAFs (p) of core SNPs for East Asians with those of Caucasians. We speculated on the applicability of Caucasian GWAS data to non-Caucasian DRPs.
Results
Validation of previous reports in Japanese samples
First, we examined the inconsistencies in disease risk assessments. A comparison of the results for 22 diseases is shown in Table 1. We obtained results similar to those of Ng et al.,12 even in the three Japanese individuals. For six diseases (rheumatoid arthritis (RA), psoriasis, lupus, Crohn’s disease, melanoma and Alzheimer’s disease), there were no mismatches across the three individuals. Seven diseases (prostate cancer, heart attack, breast cancer, obesity, abdominal aortic aneurysm, brain aneurysm and colorectal cancer) almost fully agreed across the three individuals, with the exception of small mismatches on average risk interpretation (↑ and =, or ↓ and =). Eight diseases (T2DM, restless leg syndrome, multiple sclerosis, celiac disease, atrial fibrillation, lung cancer, stomach cancer and age-related macular degeneration) had onethird or fewer mismatches of the opposite prediction (↑ and ↓).
CR analyses
Next, we calculated the CRs of DRPs between the three companies, comparing each company to one other company individually. We calculated the CR after omitting NA data. The formulae are described in the Materials and Methods section. For 23andMe and Navigenics, the CR was 0.692, and if we ignored the mismatch with average risk (=), the CR was 0.827. For 23andMe and deCODEme, the CR was 0.783, and if we ignored the mismatch with average risk (=), the CR was 0.913. For deCODEme and Navigenics, the CR was 0.722, and if we ignored the mismatch with average risk (=), the CR was 0.815. On the basis of these calculations, we found that 23andMe and deCODEme were the most concordant, whereas 23andMe and Navigenics were the least concordant in 22 DRPs.
Independency test
We conducted 2 × 2 Fisher’s exact tests for each set of two DTC companies’ DRPs for testing independence. For 23andMe and Navigenics, both predictions were correlated (P=5.3 × 10−4). For 23andMe and deCODEme, both prediction results were correlated (P=1.7 × 10−6). For Navigenics and deCODEme, both prediction results were correlated (P=2.8 × 10−4). Therefore, each set of two DTC companies’ DRPs was not independent; in other words, they correlated with each other. 23andMe and deCODEme were the most similar, whereas 23andMe and Navigenics were the least similar in their DRPs.
κ-statistics
Cohen’s κ-tests revealed that κ=0.54 between 23andMe and Navigenics (P=0.0006), κ=0.55 between deCODEme and Navigenics (P=0.0002) and κ=0.77 between 23andMe and deCODEme (P=7.3 × 10−6). The Fleiss’ κ-value for all three companies was 0.58 (P=3.0 × 10−10).
Increasing risk versus decreasing risk
Next, we counted the total number of increasing risks (higher risk than average, ↑) and decreasing risks (lower risk than average, ↓) in the three companies’ predictions for the three volunteers. We observed that the number of increasing risks was 54 and the number of decreasing risks was 105 (Table 1). Therefore, we observed that each company reported protective test results (decreasing risks) nearly two times more frequently than susceptible test results (increasing risks).
Distributions of RRs
Next, we visualized disease risk distributions to determine the characteristics of variations of the three DTC companies’ risk predictions. We compared these differences by plotting minimum, maximum and average values of RRs for each of the three samples for 22 disease conditions. The results are shown in Figure 1.
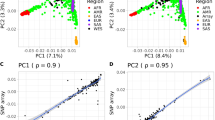
Relative disease risk distributions assigned by the three direct-to-consumer genetic test (DTC) services for a series of diseases. The relative risk (RR) (log-scale) distributions assigned by the three DTC services for the 22 disease conditions in the three Japanese samples are shown. The blue, red and green bars represent the disease risk variation (from minimum to maximum values in the three Japanese individuals’ log-scale RRs) for 23andMe, Navigenics and deCODEme, respectively. Values in parentheses indicate the number of single nucleotide polymorphisms (SNPs) analyzed.
The predictions by the three companies were related but not identical. Variations between minimum and maximum disease RRs varied widely from disease to disease, and the three DTC companies exhibited different variations. Navigenics had the largest variation of RRs in the three DTC companies’ predictions, followed by deCODEme and 23andMe. Celiac disease had the largest variation of all the 22 diseases, followed by macular degeneration and Alzheimer’s disease. Abdominal aortic aneurysm had the smallest variation of all the 22 diseases, followed by psoriasis, RA and obesity.
Increasing risk versus decreasing risk in RR distributions
Next, we calculated the average RRs of the three volunteers based on the algorithms of each of the companies. The average RRs calculated from the results of 23andMe, Navigenics and deCODEme were 0.95, 0.96 and 1.03, respectively, and the combined average RR of all three companies was 0.98. We also calculated the average RRs for RR>1 (susceptible test results) and the average of inverse of RRs for RR<1 (protective test results). In RR>1, the average RRs calculated from the results of 23andMe, Navigenics and deCODEme were 1.63, 1.82 and 1.76, respectively, and the combined average RR of all three companies was 1.74. In RR<1, the average of inverse of RRs calculated from the results of 23andMe, Navigenics and deCODEme were 2.20, 2.73 and 1.72, respectively, and the combined inverse of RR of all three companies was 2.22.
On the basis of above calculations, we observed that Navigenics had the largest variation in RR (2.73 × 1.82=4.97), whereas deCODEme had the smallest variation in RR (1.72 × 1.76=3.03). We also observed that the average RR where RR<1 had the larger deviation (2.22) compared with that of the average RR where RR>1 (1.74).
Root cause analyses of discrepancy
We then investigated the origin of these discrepancies. These mismatches could be attributed mainly to differences in the SNPs used in the calculation, the reference population used, different risk assignment methodologies and adjustment for ethnicities, as discussed below.
SNP selection
The first reason for the different risk assessments was the evaluation of different SNPs (Figure 2). For the 22 conditions covered by the three companies, only 18 out of 254 (7.1%) SNPs were reviewed by all companies, whereas 177 out of 254 (69.7%) SNPs were reviewed by only a single company (Figure 2). These 18 core SNPs shared by the three companies are listed in Table 3. Only 12 out of 22 diseases had core SNPs shared by the three companies. 23andMe and Navigenics had several overlaps, sharing 39 out of 192 SNPs (20.3%); deCODEme and Navigenics shared 39 out of 207 SNPs (18.8%); and 23andMe and deCODEme also shared 39 out of 208 SNPs (18.8%). Table 2 shows the number of SNPs for each of the 22 diseases covered by the three companies. Each company used a different number of SNPs in 19 out of 22 diseases except for 3 diseases: Alzeheimer’s disease, atrial fibrillation and psoriasis. We also showed expanded surveys of the differences in core SNPs listed in the example of T2DM (Supplementary Tables 1 and 2).
A list of 132 total SNPs was created from SNPs reviewed by the NIH GWAS catalog and four consumer genomic service providers, that is, deCODEme, Navigenics, 23andMe and Pathway Genomics. Supplementary Table 1 lists the top 27 SNPs, that is, those listed by more than one organization. Only 2% (3 of 132) of SNPs were listed by all five organizations, 3% (4) were listed by four entities, 8% (10) were listed by three organizations and an additional 8% (10) were listed by two organizations. The top seven SNPs were listed by four or five of the organizations. These SNPs were located in several genes associated with underlying physiological aspects of T2DM, including HHEX, IGF2BP2, JAZF1, KCNQ1, PPARG, SLC30A8 and TCF7L2. We also examined the statistics of SNP selections based on reference citations (Supplementary Table 2). Of 74 total studies cited by the five organizations in their T2DM analyses, Supplementary Table 2 lists the 15 that were listed by more than one organization. Metrics similar to Supplementary Table 1 can be seen. No single study was cited by all five organizations, only one study was cited by four organizations,14 five studies were cited by three organizations and nine studies were cited by two organizations. Eighty percent of the studies (59) were cited only by one organization, in this case, mostly by 23andMe.
Ethnicity adjustment
Ethnicity is important for DRP. Twelve out of the 22 disease predictions were customized for East Asian reports (Table 1). Only deCODEme provided different unique SNP marker sets for East Asians for 10 out of the 22 diseases. 23andMe customized the DRPs for East Asians in five out of the 22 diseases. Navigenics provided DRPs only for Caucasians. Table 3 also shows the differences in real allele frequencies between Japanese and Caucasian individuals for each of the core SNPs. RAFs for each of these diseases were very different in Japanese and Caucasian individuals. We also noted that the average RAFs of these core SNPs were <.5. This was consistent with NIH GWAS research database (http://www.genome.gov/GW/GWAStudies). RAF (p) is one of the most critical parameters for DRPs (absolute risk, d1, d2 and d3) in each DTC company’s prediction algorithm. Differences of RAFs between Caucasian and Japanese individuals in core SNPs for several diseases are important for understanding ethnic impact on DRP.
Average risks
DTC genomics companies did not always use the same values for the average risk of the overall population. Some of the biggest differences were for obesity (ranging from 34–64% in men and 32–59% in women), T2DM (ranging from 25–40% in men and 21–30% in women) and Alzheimer’s disease (ranging from 6–9% in men and 7–17% in women; Supplementary Table 3). Differences in average risks between European and Asian individuals in the report from 23andMe for four disease conditions are also shown in Supplementary Table 3. For example, the difference in prostate cancer was large (17.8% for European individuals versus 11.2% for Asian individuals), whereas the difference in T2DM was small (25.7% for European men versus 27.8% for Asian men and 20.7% versus 21.9% for European and Asian women, respectively).
Risk prediction algorithms
The final reason for the different risk interpretations was their differences in risk assessment technologies. 23andMe and Navigenics provided a composite risk score for each condition using different multiplicative approaches. 23andMe took the product of recentered ORs for all relevant SNPs and multiplied this value by the average population risk to estimate an individual’s absolute risk. Navigenics used a proprietary analysis method that implemented genetic composite index numbers. Supplementary Table 4A–C shows examples of simulations using the differences in predictions (d1, d2 and d3: absolute risks for each genotype) between 23andMe and deCODEme given by the same parameter sets: RAF (p), average risk (q) and OR (r). For q=0.01, both predictions were almost the same. The d1 of deCODEme was slightly lower than that of 23andMe, and the d3 of deCODEme was slightly higher than that of 23andMe. When q=0.1, both predictions were considerably different. The difference between both predictions was larger when q or r was large.
Discussion
Here, we extended the methodology of Ng et al.12 and observed the distributions of disease risk assessments for three DTC companies using three Japanese samples.
Our observations and systematic analyses of the three DTC companies demonstrated the following:
-
Our independency test showed that the overall prediction results from the three DTC companies for three volunteers in 22 diseases were correlated with each other. κ-statistic analyses showed that the agreement between each set of two DTC companies’ DRPs and among all three companies’ DRPs was better than chance (all κ P-values<0.01). However, they were not perfectly matched; less than onethird mismatching of the opposite prediction occurred in eight diseases. The consistency of predictions among the three DTC companies varied depending on the disease.
-
Four factors may explain the different predictions: SNP selection, average risk, ethnicity adjustment and prediction algorithms. In particular, the core SNPs were important for producing a better prediction consensus. Only 18 out of 254 (7.1%) SNPs over 22 diseases were reviewed by all three companies.
-
The number of predictions for decreasing risks was less than two times greater than the number of predictions for increasing risks for the 22 diseases. The RRs for reporting increasing risks had larger deviations from average than those for reporting decreasing risks.
Thus, discrepancies in predictions also occurred in Japanese individuals, similar to the report by Ng et al.12 However, our results are slightly different from theirs; specifically, the CR was improved in our results. The 2-year period between our study and the study by Ng et al. may have allowed for improved predictions by the DTC companies and hence more consistent results. However, similar to the report by Ng et al.,12 certain diseases exhibited better prediction agreements than others. For example, predictions for RA completely agreed among the three companies for all individuals. In contrast, although Ng et al. reported that 50% or less of the predictions agreed between the two companies across individuals for seven diseases,12 we did not observe large mismatches in three of the seven diseases: lupus, heart attack and Crohn’s disease. We also observed mismatches in four of the seven diseases: T2DM, restless legs syndrome, psoriasis and prostate cancer, consistent with their report. Moreover, although Ng et al.12 reported a perfect consensus in multiple sclerosis and celiac disease, we observed some mismatches.
Imai et al.15 reported that CRs between the three DTC companies for SNP genotyping data were >99.6%. We also validated that genotyping results for shared SNPs for each set of two companies for the 22 disease predictions were perfectly matched. Risk assessment differences among three companies may be because different SNPs were used to calculate risk for the same disease.15 This suggestion was confirmed by the fact that, although low prediction consensus diseases, such as T2DM and prostate cancer, were predicted with very different numbers of SNPs for each of the three companies, high prediction consensus diseases, such as Alzheimer’s disease, shared the same SNP markers among the three companies. However, the number of markers in common did not necessarily correlate with a better prediction agreement, as Ng et al.12 suggested. For example, RA predicted by very different numbers of SNPs achieved perfect consensus predictions in this study. This perfect agreement may be because of the consensus of one strong-effect marker. Ng et al.12 suggested that when the DTC companies did not use the same strong-effect markers, large differences in predictions occurred. We confirmed this for three mismatched diseases: restless leg syndrome, obesity and lung cancer, where there were no core SNPs shared by the three companies. Moreover, we found that only 7.1% of SNPs over 22 diseases were shared by all three companies. This was probably the most important factor for mismatch prediction.
Expanded comparative analyses for SNP selections with the addition of the NIH GWAS catalog and four DTC services yielded similar results for all disease conditions, and only a very small percentage of total SNPs associated in different studies were listed by the majority of organizations reviewing the condition. These core SNPs might be important for DRPs.
Most common disease conditions are polygenic, involving multiple genes and SNPs. Although research has focused on finding one or a few SNP associations, a comprehensive view of the polygenic landscape of genes relating to a particular condition has not been taken. An overall view of the most relevant SNPs and a composite of their quantitative risk score has been a barrier in establishing validity and utility in personalized genomics.10 An important challenge is compiling a list of all SNPs related to a condition. This SNP list could then be sorted for SNP quality, with granular parameters such as higher weighting for more important SNPs as determined by ORs, P-values or other metrics, or by some other technique. Adjustments could be made for SNPs in linkage disequilibrium, where many SNPs in the same region provide essentially the same information. An understanding of the core SNPs involved in a condition could advance our understanding of how SNPs operate together systemically, which along with epigenetics, structural analysis and exome and whole-human genome sequencing could lead to a more comprehensive means of determining the causality of common disease.
Average risk is another important parameter for DRP, especially as absolute disease risk is derived from RR and average risk. We compared the average risks of the three DTC companies in the same disease conditions and found considerable differences in certain disease conditions, including T2DM and Alzheimer’s disease. Average risk varies depending on how one defines the population.12 For example, Navigenics and deCODEme apply one average risk value each for men and women, whereas 23andMe provides different risk assignments by age tier (for example, the incidence of T2DM increases with age). This ambiguity in the definition of a ‘population’ should be carefully considered when we interpret absolute disease risk.12 There are also differences in phenotype definitions, and application of the same phenotype definition may result in different risk estimates.10 Increasing consistency in selecting the most accurate references for average risk is therefore important. Currently, most underlying research studies have been carried out in Western European (Caucasian) populations. Thus, expanding GWAS research to other non-Caucasian populations is important.
Even when the same core SNPs and average risks were used for the predictions, differences in parameter settings and algorithms may affect consistency. The three companies investigated here used similar prediction algorithms, but required three critical parameters for risk prediction: RAF (p), average risk (q) and OR (r). We found that the parameter sets for q and r were not always the same among the three companies. Reliable standardized data sets for q and r were important for better prediction agreements. Although we chose the same parameter sets (p, q and r), we found that prediction results (absolute risks, d1, d2 and d3) were different in certain cases, based on differences among their prediction algorithms. Navigenics used the most complex mathematical formula for solving simultaneous mathematical equations, and deCODEme used the simplest formula, assuming that the OR was nearly equal to the risk ratio. The methods for generating composite risk scores for disease conditions from the multiple SNP markers were also similar, but slightly different among the three companies. Our computer simulations showed that when q or r was larger, the difference of prediction results (d1, d2 and d3) was larger; alternatively, it was small when q was small. Thus, development of reliable prediction algorithms and evaluation of the reliability of the DRP is critical for future research.
Ethnicity was also a crucial issue. As the RAF (p) varied among different ethnicities, the reference population also had an influence on relative DRPs. Actually, the RAF (p) was different for Japanese and Caucasian individuals in core SNP markers for each disease. Ethnicity differences affect DRPs in T2DM.16 Navigenics does not provide reports specifically for Asian individuals, whereas 23andMe provides three reports (for T2DM, RA and prostate cancer) specifically for Asian individuals. Thus, establishment of reliable risk assessment methods for all ethnicities is required.
Risk assessments for each ethnicity are important, but only limited data are currently available. When we consider cases in which Caucasians’ GWAS data are applied for Japanese individuals, RAF (p) and average risk (q) should be obtained from Japanese (or at least East Asian) data. RAF (p) can be currently obtained from the Hapmap database. The impact of p for DRP (absolute risk, d1, d2 and d3) was larger, and, in particular, the average risk (q) was larger. Although we currently do not have consensus for universal data sets for average risks (q), standardization of reliable epidemiological data for non-Caucasians is important. It is also imperative to understand the mathematical basis for how RAF (p) and average risk (q) affects DRPs (absolute risk, d1, d2 and d3). We will explore this in our future work, for understanding basic mechanisms of reliable gold-standard DRPs.
In conclusion, establishing a universal core list of disease-associated SNPs for non-Caucasians is critical for better predictions, for which more genome-wide studies in these populations are needed. We recommend that DTC companies and the research community continue to investigate and share reliable and useful data sets for non-Caucasian ethnicities.
References
Dewey, F. E., Chen, R., Cordero, S. P., Ormond, K. E., Caleshu, C., Karczewski, K. J. et al. Phased whole-genome genetic risk in a family quartet using a major allele reference sequence. PLoS Genet. 7, e1002280 (2011).
Hivert, M. F., Jablonski, K. A., Perreault, L., Saxena, R., McAteer, J. B., Franks, P. W. et al. Updated genetic score based on 34 confirmed type 2 diabetes Loci is associated with diabetes incidence and regression to normoglycemia in the diabetes prevention program. Diabetes 60, 1340–1348 (2011).
Kamatani, Y., Matsuda, K., Okada, Y., Kubo, M., Hosono, N., Daigo, Y. et al. Genome-wide association study of hematological and biochemical traits in a Japanese population. Nat. Genet. 42, 210–215 (2010).
Miyake, K., Yang, W., Hara, K., Yasuda, K., Horikawa, Y., Osawa, H. et al. Construction of a prediction model for type 2 diabetes mellitus in the Japanese population based on 11 genes with strong evidence of the association. J. Hum. Genet. 54, 236–241 (2009).
Morgan, A. A., Chen, R. & Butte, A. J. Likelihood ratios for genome medicine. Genome Med. 2, 30 (2010).
Roberts, N. J., Vogelstein, J. T., Parmigiani, G., Kinzler, K. W., Vogelstein, B. & Velculescu, V. E. The predictive capacity of personal genome sequencing. Sci. Transl. Med. 4, 133ra158 (2012).
Topol, E. J. Comment on ‘the predictive capacity of personal genome sequencing’. Sci. Transl. Med. 4, 135le135. author reply 135lr133 (2012).
Lyon, G. J. Personalized medicine: bring clinical standards to human-genetics research. Nature 482, 300–301 (2012).
Pierce, B. L. & Ahsan, H. Clinical assessment incorporating a personal genome. Lancet. 376, (869; author reply), 869–870 (2010.).
Swan, M. Multigenic condition risk assessment in direct-to-consumer genomic services. Genet. Med. 12, 279–288 (2010).
Zehnbauer, B. Direct-to-consumer genetics testing—fair comparisons? Clin. Chem. 57, 369–371 (2011).
Ng, P. C., Murray, S. S., Levy, S. & Venter, J. C. An agenda for personalized medicine. Nature 461, 724–726 (2009).
Bloss, C. S., Schork, N. J. & Topol, E. J. Effect of direct-to-consumer genomewide profiling to assess disease risk. N. Engl. J. Med. 364, 524–534 (2011).
Steinthorsdottir, V., Thorleifsson, G., Reynisdottir, I., Benediktsson, R., Jonsdottir, T., Walters, G. B. et al. A variant in CDKAL1 influences insulin response and risk of type 2 diabetes. Nat. Genet. 39, 770–775 (2007).
Imai, K., Kricka, L. J. & Fortina, P. Concordance study of 3 direct-to-consumer genetic-testing services. Clin. Chem. 57, 518–521 (2011).
Chen, R., Corona, E., Sikora, M., Dudley, J. T., Morgan, A. A., Moreno-Estrada, A. et al. Type 2 diabetes risk alleles demonstrate extreme directional differentiation among human populations, compared to other diseases. PLoS Genet. 8, e1002621 (2012).
Acknowledgements
This project was supported by a grant from the JST, PRESTO. We would like to thank Professor Gavin Sherlock, Dr Keiichi Kodama and Dr Chirag Patel from Stanford University, Professor Kaori Muto from Tokyo University, and Shin Takizawa, a research assistant in Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Dr Atul Butte is a founder and advisor to Personalis, a company providing clinical genome analysis and interpretation.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Kido, T., Kawashima, M., Nishino, S. et al. Systematic evaluation of personal genome services for Japanese individuals. J Hum Genet 58, 734–741 (2013). https://doi.org/10.1038/jhg.2013.96
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2013.96
Keywords
This article is cited by
-
Direct-to-Consumer Genetic Testing and Its Marketing: Emergent Ethical and Public Policy Implications
Journal of Business Ethics (2022)
-
Direct‐to‐consumer genetic testing in China and its role in GWAS discovery and replication
Quantitative Biology (2021)
-
Estimation of the risk of a qualitative phenotype: dependence on population risk
Journal of Human Genetics (2017)