Abstract
In a loss-of-viability screen of small molecules against methicillin-resistant Staphylococcus aureus (MRSA) USA300, we found a small molecule, designated DNAC-2, which has an MIC of 8 μg ml−1. DNAC-2 is a quinolinol derivative that is bactericidal at 2X MIC. Macromolecular synthesis assays at 2 × MIC of DNAC-2 revealed inhibition of DNA, cell wall, RNA and protein synthesis within fifteen to thirty minutes of treatment when compared to the untreated control. Transmission electron microscopy of DNAC-2-treated cells revealed a significantly thicker cell wall and impaired daughter cell separation. Exposure of USA300 cells to 1 × MIC of DNAC-2 resulted in mislocalization of PBP2 away from the septum in an FtsZ-independent manner. In addition, membrane localization with FM4–64, as well as depolarization study with DiOC2 and lipophilic cation TPP+ displayed membrane irregularities and rapid membrane depolarization, respectively, in DNAC-2-treated cells vs -untreated control. However, DNAC-2 exhibited almost no toxicity toward eukaryotic membranes. Notably, DNAC-2 drives energy generation toward substrate level phosphorylation and the bacteria become more sensitive to DNAC-2 under anaerobic conditions. We propose that DNAC-2 affects USA300 by targeting the membrane, leading to partial membrane depolarization and subsequently affecting aerobic respiration and energy-dependent functional organization of macromolecular biosynthetic pathways. The multiple effects may have the desirable consequence of limiting the emergence of resistance to DNAC-2.
Similar content being viewed by others
Introduction
In the seemingly never-ending race between human host and fast evolving multidrug-resistant bacteria, the development of new efficient antibiotics is an important step in counteracting the emerging resistance. However this process is labor and capital intensive as well as time consuming.1 Due to a relative lack of financial incentive from government agencies, there are now fewer pharmaceutical companies in the business of antibiotic development. Even though several bacterial targets for antibiotic development have been identified by genomic sequencing,2, 3 high throughput screening of compound libraries using these targets has yet to produce a single drug candidate.4 As the antibiotic pipeline has dried up, drug repurposing has become a viable alternative, especially against multiple drug-resistant bacteria.5 This process involves repositioning already-approved antibiotics for secondary use, and then further developing it for that purpose. With this in mind, we and others6 have screened libraries of small molecules to identify candidate compounds that enhance activity of previously approved drugs and those facing patent expiry.
Here we describe a novel candidate compound called DNAC-2, a quinolinol derivative that is effective both in monotherapy and also in conjunction with oxacillin against methicillin-resistant Staphylococcus aureus (MRSA). We propose that DNAC-2 acts on the bacterial membrane to cause membrane depolarization and disrupt macromolecular synthesis, possibly interfering with aerobic respiration and thus driving a shift in the metabolic pathways of MRSA toward a fermentative mode which led to increased susceptibility to DNAC-2, resulting in reduced viability.
Results
DNAC-2 was one of the six compounds identified at the ICCB-Longwood/NSRB Screening Facility at Harvard Medical School in a loss-of-viability screen against MRSA USA300 as described previously.7 The structure of DNAC-2 (8-(trifluoromethoxy)-2-(trifluoromethyl)quinolin-4-ol) is shown in Figure 1a. DNAC-2 has a MIC of 8 μgml−1 against MRSA USA300 and of 32 μg ml−1 against the standard test strain Staphylococcus simulans 22. The MIC of DNAC-2 against a variety of CA-MRSA and HA-MRSA was in the range between 4 and 8 μg ml−1 (Supplementary Table 1).
Microbicidal action of DNAC-2 against MRSA USA300 with different concentrations of DNAC-2 with and without sub-MIC concentration of oxacillin. (a) Structure of DNAC-2, with a chemical formula of C11H5F6NO2 and a molecular weight of 297.1564, is delineated. For the growth curve, an overnight culture was diluted and grown in the presence of increasing concentrations of DNAC-2 with and without oxacillin, left untreated or treated only with sub-MIC oxacillin as a control. In (b), an overnight culture was diluted and grown in the presence of 0.5 × DNAC-2 (open circle), 1 × DNAC-2 (closed square), 2 × DNAC-2 (open square) and 4 × MIC of DNAC-2 (closed triangle) or left untreated (closed circle). In (c), cells were treated with 16 μgml−1 of oxacillin (¼ MIC) and increasing concentrations of DNAC-2 at 0.25 × MIC (open circle), 0.5 × MIC (closed diamond), 1 × MIC (open inverted triangle), 2 × MIC (closed triangle), 4 × MIC (closed square) or left untreated (closed circle). For the time-kill curve (d), samples were taken at 0, 2, 4, 6 and 24 h. and a viable count carried out. Asterisk denotes P-values of <0.05 as determined by Tukey’s multiple comparison test of two way analysis of variance. The lines show the mean with s.e.
A full colour version of this figure is available at the Journal of Antibiotics journal online.
Assessing toxicity of DNAC-2
We assessed the toxicity of DNAC-2 toward eukaryotic cells in several different ways: (1) measurement of lysis of 4% sheep erythrocytes with DNAC-2 at varying concentrations (0.5–16 × MIC) against USA300; (2) LDH release from exposed cultured human bronchial epithelial (CFBE) cells upon treatment with DNAC-2; (3) evaluating the in vitro inhibition of CYP using human liver microsomes and (4) exposure of yeast cells to 64 × MIC of DNAC-2. Using 2-fold dilutions ranging from ¼ MIC to 16 × MIC of DNAC-2, we quantified the RBCs lysis as well as LDH release from CFBEs. Red cell lysis and LDH release from CFBE attributable to 2% Triton X-100 was set to 100%. CFBE cells exposed to DNAC-2 disclosed ~10% LDH release at the 8 × MIC (64 μg ml−1) and 5% RBC lysis at 16 × MIC vs the Triton X-100 control. DNAC-2 did not affect a majority of the CYP isoforms tested with the exception of isoform 1A2, which showed a ~20% reduction in activity at 1 μm and 62% at 10 μm of DNAC-2. In addition, Candida albicans exposed to 64 × MIC of DNAC-2 grew normally and did not exhibit any sign of toxicity.
Pharmacokinetics of DNAC-2
The in vitro metabolic stability of DNAC-2 was determined using human, rat, dog and mouse liver microsomes. DNAC-2 was not significantly metabolized by human, rat and dog liver microsomes (~90% remaining after 30 min) while the control compound midazolam was (1.4% remaining after 30 min). Even at 90 min, ~70–80% of DNAC-2 remained in the three microsomes above. This contrasts with mouse liver microsome where the metabolism was faster, with 50% remaining at 30 min and 17.4% at 90 min. The in vitro half-life (t1/2) for 1 μm DNAC-2 was estimated to be ~630 min in human microsomes, ~495 min in rat microsomes and ~165 min in dog microsomes whereas the half-life in mouse liver microsomes was only 39 min. The intrinsic clearance (CLint) of 1 μm of DNAC-2 was 2.2, 2.8, 8.4 and 35.4 μl min−1 mg−1 for human, rat, dog and mouse liver microsomes, respectively. These data are consistent with a reasonable level of stability for DNAC-2 in human, rat and dog liver microsomes, but DNAC-2 is rapidly cleared in mouse liver microsomes. Collectively, these findings demonstrated the divergence in compound stability in mouse versus other animal species.
At both 1 and 10 μm, DNAC-2 was permeable in the Caco-2 cells in both directions, with mean recoveries ranging from 77.7 to 93.4% in the apical-basal direction and 92.8 to 98.1% in the basal-apical direction, indicating that the compound was stable and non-specific binding was not observed under the conditions used for the experiment. The presence of the P-gp inhibitor ketoconazole did not have any impact on the permeability or efflux ratio, further confirming that DNAC-2 is not a substrate of P-gp. At 1 μm concentration, DNAC-2 was ~89% bound to human, rat, dog and mouse plasma proteins at equilibrium.
Enhanced microbicidal activity of DNAC-2 on MRSA strain USA300 in sub-inhibitory concentrations of oxacillin
DNAC-2, with a MIC of 8 μg ml−1 against USA300, showed a concentration-dependent anti-MRSA activity (Figure 1b) and enhanced activity in the presence of a sub-inhibitory concentration of oxacillin at 16 μg ml−1 (Figure 1c)—MIC of oxacillin is 64 μg ml−1. This result corroborated with time-kill experiments where we found that DNAC-2 exhibited a concentration-dependent bactericidal activity (Figure 1d), resulting in a>99.9% reduction of the initial bacterial inoculum within 24 h when added at 4 × MIC (32 μg ml−1).
Effect of DNAC-2 on macromolecular synthesis in S. simulans
A vital part in the discovery and development of antibacterial drugs is an evaluation of mechanism of action (MOA). As the screen for DNAC-2 is a mechanistically unbiased screen against MRSA, a direct target was not immediately apparent. One way to determine the target is to obtain mechanistic fingerprints of the drug for different cellular targets using the macromolecular synthesis assays.
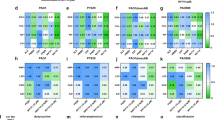
Using labeled precursors with different concentrations of DNAC-2 and control antibiotics (ciprofloxacin, rifampicin, vancomycin and tetracycline) as inhibitors of DNA, RNA, cell wall and protein synthesis, respectively, macromolecular synthesis inhibition assays were performed with samples taken at 5, 15, 30, 45 and 60 min after addition of antibiotics. As shown in Figure 2, S. simulans 22 cells treated with increasing concentrations of DNAC-2 (0.5, 1 and 2 × MIC) showed concentration-dependent inhibition of DNA, RNA, protein and cell wall synthesis. Such results are characteristic for compounds interfering with membrane-bound mechanisms of energy generation and indicate that either the uptake of the labeled metabolites or their incorporation into macromolecules is affected. Thus, DNAC-2 appears to have pleiotropic effects on biosynthetic pathways in Staphylococci.
Macromolecular synthesis assays with DNAC-2. Inhibition of macromolecular biosynthesis in Staphylococcus simulans 22 by DNAC-2 was measured by inhibition of incorporation of radiolabeled precursor into (a) nascent DNA, (b) RNA, (c) protein or (d) cell wall peptidoglycan. Bacteria were treated with different concentrations of DNAC [0.5 × MIC (open circle), 1 × MIC (open triangle) and 2 × MIC (close inverted triangle)] in the presence of appropriate radioactive precursors and samples were obtained at 0, 5, 15, 30, 45 and 60 min for analyses. A no drug negative control (close circle) was run in each experiment. The positive control antibiotics (denoted by open squares) used at 10 × MIC were ciprofloxacin to inhibit DNA synthesis, rifampicin to inhibit RNA synthesis, tetracycline to inhibit protein synthesis and vancomycin to inhibit cell wall synthesis.
Effect of DNAC-2 on the cellular morphology of MRSA strain USA300 by TEM
Given this multitude effect on cellular machinery, we wanted to assess the effect of DNAC-2 on cellular morphology by TEM. Accordingly, USA300 cells were exposed to 2 × MIC DNAC-2 for 90 min, after which they were processed for TEM as described in the Materials and Methods section. Evaluation of ~300 cells (Figures 3a and b) from both samples revealed that half of the DNAC-2-treated cells (~55%) displayed cross wall/septum formation with reduced or undetectable splitting compared to just 20% in untreated cells. Measurements of the cell wall thickness of both treated and untreated cells in ~30 different fields disclosed significantly thicker cell walls (30–42 nm) in treated cells compared to untreated control (25–30 nm; Figure 3c). The hemispherical daughter cells measured 0.2–0.5 μm3 in volume compared to completely divided daughters which are usually ~1 μm3. Furthermore, DNAC-2-treated cells displayed mesosome-like membrane inclusions predominantly at the cross wall in dividing cells but along the cell wall in non-dividing cells as seen in Figure 3d (arrows). In addition, in rare cases, branching and multiple cross walls without obvious cell separation were observed in the sample treated with DNAC-2.
Effect of DNAC-2 on cell wall by TEM. USA300 cells were grown to mid-log phase and treated with DNAC-2 for 90 min, or left untreated (control). Cells were then collected and processed for EM as described in Material and Methods section. (a) Control cells and DNAC-2-treated cells are shown at × 1200 magnification (b) DNAC-2-treated cells have significantly more septate cells than control; (c) DNAC-2-treated cells have a significantly thicker cell wall as demonstrated in the box and whiskers plot showing STD. The symbol (*) represents statistically significant difference, with P<0.05 comparing DNAC-2 treated to control cells; (d) Mesosome-like invaginations are seen both along the cell wall and along the septum in cells at × 15 000 magnification (arrows, right panel).
DNAC-2 causes mislocalization of PBP2 and FtsZ
As DNAC-2 inhibits cell wall synthesis, we examined the localization of PBP2, an enzyme involved in cell wall synthesis, with and without DNAC-2 (1 × MIC) for 30 min (Figure 4a, left panels). In untreated cells, PBP2 was localized at the septum in approximately one quarter of the population (23% of the cells, n=593). This number decreased three-fold to only 7% of the cells in the presence of 1 × MIC of DNAC-2 (n=524). Interestingly, PBP2 was localized to puntate areas in treated cells as opposed to distribution around the membrane as has been seen with interruption of cell wall synthesis.
(a) Localization of PBP2 and FtsZ in the presence of DNAC-2. Localization of the PBP2 and FtsZ was tracked using strains expressing GFP-PBP2 or FtsZ-CFP after receiving no treatment or treatment with 1 × MIC DNAC-2 for 30 min; (b) Localization of PBP2 and membrane in the presence of DNAC-2. Cells were grown until mid-exponential phase (O.D600nm=0.4) and incubated with DNAC-2 at 3 × the MIC (24 μg ml−1) or DMSO control for 30 min. Cells were then stained with membrane dye FM4–64 (Invitrogen) at a final concentration of 0.5 μgml−1 for 5 min with shaking, washed with PBS and mounted on microscopy slides covered with a thin layer of PBS agarose, before being imaged by Widefield microscopy.
FtsZ is known to play a role in the localization of PBP2 to the septum in dividing cells. To determine if DNAC-2 interferes with FtsZ and hence PPB2 localization, we exposed a S. aureus strain containing FtsZ tagged with CFP to 0.5–2 × MIC of DNAC-2 for 30 min (Figure 4a, right panels). Localization of FtsZ was not severely affected by DNAC-2, with septal localization in 57% of the cells following DNAC-2 treatment (n=477) compared to 61% of the cells (n=484) without treatment (Figure 4). The FtsZ division ring also appears to be normal in these cells. Thus, the mislocalization of PBP2 is not mediated by FtsZ.
DNAC-2 affects the cell membrane of MRSA USA300
The mesosome-like structures seen near the cross wall of the bacterial cell in Figure 3d of the EM studies prompted us to further examine the effect of DNAC-2 on the membrane. We examined this effect with several different techniques: (1) super resolution fluorescent microscopy of unfixed MRSA COL cells stained with membrane dye FM 4–64; (2) ascertaining change in membrane potential with the dye DiOC2 and lipophilic cation TPP+ and (3) measuring ATP and potassium leakage upon treatment with DNAC-2. To examine membrane integrity, we incubated MRSA COL with 1 × DNAC-2 for 5–30 min followed by staining with FM 4–64. To help localize the membrane, we also stained the cell wall with bodipy-vancomycin to visualize the cell wall. Membrane defects such as bulges in the membrane, as discerned by FM4–64 staining, can be seen in DNAC-2-treated cells vs -untreated control as early as 5 min, but more dramatic at 30 min (Figure 5a). We also found co-localization of the membrane defect with mislocalized PBP2 at 5 min (Figure 4b). In addition, we observed increased bodipy-vancomycin staining at the septum in treated cells. There was also a moderate decrease in membrane potential associated with treatment of USA300 with 1–4 × MIC of DNAC-2 (8–32 μg ml−1) as demonstrated by a loss in red fluorescence in cells exposed to DiOC2, with the effect being rapid and occurring within 5 min of exposure to DNAC-2 (Figure 5b). However, the reduction in membrane potential was less than the positive control, CCCP, a known proton translocator. Moderate reduction of the membrane potential was confirmed by means of the lipophilic cation TPP+ which moves rapidly across the cytoplasmic membrane in response to a trans-negative membrane potential and allows for quantification of the effect (Figure 5c). Remarkably, treatment of USA300 with DNAC-2 did not induce leakage of potassium (Supplementary Figure A) or ATP (not shown) from cells, different from what has been observed with daptomycin.8 Altogether, these data suggest that DNAC-2 may target the cell membrane, causing irregular cell membrane bulging, but DNAC-2 is certainly not a pore forming agent as has been described for cationic antimicrobial peptide and daptomycin which lead to rapid leakage of ATP and potassium upon contact with bacterial cells.
Disruption of cell membrane and depolarization by DNAC-2. (a) Cells were untreated (control) or treated with 1 × MIC DNAC-2 for 30 min and stained with FM4–64 to visualize the membrane, Van-FL to visualize the cell wall (middle panels) or Hoechst 33 342 to stain the DNA. (b) Actively growing USA300 cells were back diluted, resuspended in PBS with glucose and treated with DiOC2 in the dark for 30 min at 25 °C. Loss in fluorescence was monitored over time upon adding increasing concentrations of DNAC-1 at 1 × MIC (open square), 2 × MIC (closed square) or 4 × MIC (closed diamond). The untreated cells are shown as open circles and CCCP was used as a positive control (closed circle). Each line represents the means accompanied by s.e. of 3 independent readings. The asterisks represent statistical significance (P-values <0.001) from the control using the Kruskal–Wallis test with Dunn’s multiple comparisons. (c) Impact of DNAC-2 on the membrane potential (ΔΨ) of Staphylococcus simulans 22. ΔΨ was determined using [3H]TPP+ as described in Materials and Methods.8 Eight min (↓) after the addition of [3H]TPP+, the followings were added: 5 μM CCCP (closed triangles), 0.5 × MIC DNAC-2 (closed diamonds), 1 × MIC DNAC-2 (open diamonds), closed squares: untreated control.
DNAC-2 upregulates genes involved in fermentative metabolism and is more potent under anaerobic conditions against MRSA USA300
It has been shown that molecules activating heme synthesis can induce anaerobic toxicity in S. aureus.9 We thus measured the MICs of DNAC-2 alone or in combination with sub-MIC oxacillin against USA300, under both aerobic and anaerobic conditions (Table 1). For comparison, we also evaluated the MICs of well-known antibiotics that affect different biosynthetic pathways like vancomycin (cell wall), kanamycin (protein), erythromycin (protein) and levofloxacin (DNA). Anaerobic conditions had a minor effect (2-fold decrease) on the MIC of levofloxacin, but did not have any effect on the MIC of oxacillin, vancomycin, kanamycin or erythromycin. Surprisingly, DNAC-2 alone was more potent against the bacteria under anaerobic vs aerobic conditions (MIC of 1–2 vs 8 μgml−1). It has been previously shown that addition of hemin or menadione can alleviate the small colony variant phenotype of S. aureus10 that arises from the decoupling of the electron transport chain. To determine if membrane depolarization due to DNAC-2 causes a defect in the electron transport chain resulting in an induction of the fermentative pathway, we looked at the effect of DNAC-2 in the presence of menadione, hemin and nitrate (Table 1) under both aerobic and anaerobic conditions. We also used HQNO, which inhibits cytochrome B and causes uncoupling of the electron transport chain (ETC),11, 12 for comparison. Under both aerobic and anaerobic growth conditions, addition of menadione, hemin and nitrate did not have a significant effect on the MIC of DNAC-2 as compared to DNAC-2 alone. The inhibitory concentration of HQNO (at >32 μg ml−1), with and without oxacillin, also remained unchanged under anaerobic condition (Table 1). To ascertain the effect of DNAC-2 on genes active in fermentation, we used quantitative real-time PCR (qRT–PCR) to evaluate upregulation of representative genes in the fermentation pathway including those for formate (pflB, SAOUHSC_00187), lactate (ldh, SAOUHSC_00206), and ethanol metabolism (adh, SAOUHSC_00113). We observed a twofold increase in the transcript level of both pflB and ldh vs untreated control, but no significant change in adh upon treatment of actively growing USA300 with 1X MIC for just 10 min. Preliminary analysis of RNA-Seq data also confirmed the effect of DNAC-2 on selective induction of genes involved in the fermentation pathways while those involved in the ETC pathway were not significantly altered (data not shown). Thus, DNAC-2 appears to induce genes involved in fermentation impacting energy generation which appears to become even more relevant under anaerobic conditions.
Discussion
Antibiotic resistance in MRSA has long been recognized by CDC as a major public health threat. CDC estimates 80,461 invasive MRSA infections in the US and 11 285 related deaths occurring in 2011.13 An unknown but much higher incidence of less severe infections occurred in both community and healthcare settings. Concurrently, antibiotic resistance rates of MRSA are surpassing 50% in 5 of the 6 World Health Organization world regions. Despite multiple efforts, the success rate for discovery of new anti-MRSA therapeutics has been poor. Of the nine MRSA drugs that have been approved in the last decade, only one new drug class (the oxazolidinones) has emerged. Three of the nine new drugs belong to the same class as vancomycin,14 but as vancomycin resistance has crept up, these new derivatives could conceivably lose their effect.
One of the ways to combat increasing antibiotic resistance in MRSA is to repurpose already available compounds with known chemical structures. We thus screened the NERCE small molecule library for molecules that inhibit the viability of USA300. We have previously reported another molecule called DNAC-1 as a potent inhibitor of MRSA viability,7 especially in combination with cell wall active antibiotic such as oxacillin. DNAC-1 was shown to have a bimodal effect, acting mainly on the cell membrane in a concentration-dependent manner and, to a lesser extent, on the cell wall.
Our second candidate DNAC-2 is a quinolinol derivative,15 effective on its own against a variety MRSA strains. The activity of DNAC-2 is potentiated with β-lactams such as oxacillin against many MRSA strains. As DNAC-2 was identified by a mechanistically unbiased screen against MRSA, a direct target was not readily apparent. Drugs that have a specific target-driven mechanism typically exhibit signature-specific effects in macromolecular synthesis assays.16 In these assays, DNAC-2 was efficient in inhibiting macromolecular synthesis in a concentration-dependent manner, including DNA, cell wall, along with RNA and protein synthesis. Previous macromolecular synthesis assays16 have shown that a membrane active molecule such as the protonophore trifluorocarbonylcyanide phenylhydrazone (FCCP) can disrupt ATP synthesis by collapsing the proton gradient required for oxidative phosphorylation, thus causing a characteristic disruption of all pathways in the macromolecular synthesis assay and eventually potassium leakage at above MIC concentrations. In our hands, DNAC-2 also appears to target the membrane on staphylococcal cells as disclosed by fluorescence microscopy, macromolecular synthesis and membrane depolarization assays. Importantly, we did not observe any clear evidence of membrane pore formation as evidenced by the absence of significant ATP or potassium leakage into the extracellular milieu. In addition, there were no major differences in extracellular and total ATP levels between DNAC-2-treated and -untreated cells, thus indicating that the intracellular ATP level did not markedly change upon exposure to DNAC-2. DNAC-2 also differs from nisin (a cationic antimicrobial peptide) and daptomycin, another membrane active agent that disrupts DNA, RNA, protein and cell wall synthesis and rapidly depolarizes membrane potential, in that nisin and daptomycin are both membrane pore former, resulting in potassium leakage to the outside of the bacterial cell (24).
Our toxicity assays also clarify that the effect of DNAC-2 on bacterial membrane has little bearing on eukaryotic membranes. Even at high concentrations of DNAC-2 (16 × MIC), eukaryotic cell membranes were minimally disrupted. More specifically, we observed a mere 5% lysis in RBCs and ~10% LDH release from human bronchial epithelial cells at high concentrations of DNAC-2. Evaluation of the pharmacokinetic profiles of DNAC-2 shows that it does not inhibit cytochrome P450, consistent with a lack of major cytotoxicity toward eukaryotic cells. Finally, DNAC-2, even at high MIC, does not inhibit growth of Candida albicans. Altogether, these data suggest that DNAC-2 is a membrane active compound against bacteria, but has no major effect on eukaryotic membranes.
Using EM, we observed a significant thickening of the cell wall in DNAC-2-treated cells; this was also seen in the bodipy-vancomycin staining of the cell wall. Curiously, a significant portion of DNAC-2-treated cells (~55%) showed cross wall/septum formation and a reduced splitting system compared to just 20% of untreated cells. It has been shown previously17 that unsplit hemispherical cells with a complete septum have a cell volume ranging 0.2–0.5 μm3. Given that the hemispherical daughter cells in DNAC-2-treated sample have similar cell volume, it is likely there is impairment in cell division or a change in cell wall. These data were reminiscent of the lack of daughter cell separation seen upon treatment of vancomycin-sensitive cells with sub-MIC vancomycin.18 In the case of vancomycin-sensitive cells, the effect was traced back to an inability of the murein hydrolases to access the cell wall due to association of the cell wall subunits with vancomycin as shown by an inhibition in autolytic activity. DNAC-2, however, did not have a significant effect on autolysis as monitored by a Triton X-100 induced autolysis assay (data not shown). Mislocalization of the cell wall synthesis enzyme PBP2 upon DNAC-2 treatment confirmed disruption of cell wall synthesis as a result of treatment with DNAC-2. The appearance of the normal looking FtsZ rings ruled out mislocalization of FtsZ as a cause of the PBP2 defect in the cell wall. Normally, substrate binding directs PBP2 to the septum.19 In the absence of substrate binding or disruption in cell wall synthesis, PBP2 normally shows a homogenous signal around the entire cell. Amazingly, PBP2 appears to be localized to areas that show membrane bulging in cells exposed to DNAC-2. This was also confirmed by co-localization of the FM4–64-assicated membrane defect with misplaced PBP2-linked fluorescence marker at 5 min after exposure to DNAC-2 (Figure 4b). Taking into account that DNAC-2 has an immediate impact on the membrane potential (within 5 min—Figures 5b and c), we propose that DNAC-2 likely interacts with the membrane to increase its surface area, simultaneously reducing the membrane potential and impacting on the functionality of protein complexes that require the membrane as structural element for functional organization. Reduction of the membrane potential, for instance, is known to alter spatial organization of cell division proteins as well as those involved in the electron transport chain (ETC), ultimately disrupting cell wall, DNA, RNA and protein synthesis as has been described for daptomycin (24) (also unpublished data). This alteration in turn could lead to increased false septation and incomplete cell separation as seen in the EM studies.
The qRT–PCR data showing increased transcription of both ldh and pflB indicate that S. aureus shifts from aerobic respiration to fermentation in the presence of DNAC-2. This finding suggests a negative impact on energy generation from oxidative phosphorylation, which can be attributed to altered membrane potential. The increased cell wall thickness seen in the DNAC-2-treated cells is consistent with anaerobic growth of S. aureus. It has been suggested that targeting fermentation can be a viable therapeutic strategy.20 The decreased MIC for DNAC-2 under anaerobic conditions vs aerobic conditions suggests that fermentative growth may sensitize the bacteria to DNAC-2. One possible explanation is that increased fermentation in the presence of DNAC-2 results in intracellular acidification which ensues in cell death.21
Cui and colleagues22 have proposed that increased cell wall thickness can increase the burden on energy metabolism of the cells since murein production necessitates a large amount of ATP and key replication metabolite usage. Fuchs et al.23 have further propositioned that anaerobic growth in S. aureus could repress DNA replication and cell division. In consequence, this repression of DNA replication, in combination with multiple pleiotropic effects on the cell, may help explain why we have not been able to isolate spontaneous DNAC-2-resistant mutants so far despite multiple serial passages at sub-MIC or increasing MIC concentrations of DNAC-2 (data not shown). However, it is not clear if these multiple effects are simultaneous or sequential in the cause of bacterial cell death.
In conclusion, DNAC-2 is a quinolinol derivative that appears to have prokaryotic membrane- specific effects which result in membrane rearrangement, PBP2 accumulation in the area of perturbed membrane, disrupted membrane potential and eventually a myriad of altered metabolic processes including inhibition of cell wall, DNA, RNA and protein synthesis. These effects could conceivably make it harder for resistance to emerge against this compound. We have carried out limited SAR studies which will aid in the development of better quinolinol-based antimicrobials in the future. Further investigation into the mode of action of DNAC-2 is under way in our laboratory using improved derivatives of DNAC-2.
Material and methods
Bacterial strains, strain construction and media
The strain used for compound screening was MRSA USA300, a community-acquired MRSA strain.7 We also evaluated the MIC of clinical isolates of MRSA (Community-Acquired MRSA (20 strains), Hospital-Acquired MRSA (9 strains)) obtained from the Dartmouth Hitchcock Medical Centre, Lebanon NH. The bacterial cells were grown in Mueller Hinton Broth (Difco, Sparks, MD, USA) (MHB) supplemented with calcium sulfate (50 μg ml−1) and magnesium sulfate (25 μg ml−1) (MHC) for MIC studies and in Tryptone Soy broth (TSB) (Difco) for growth.
MRSA COL was used for assessing membrane integrity, permeability and cell wall integrity by fluorescence microscopy. Strains BCBPM073, a derivative of MRSA strain COL expressing fusion of sfGFP-PBP224 was used to evaluate defects in the localization of PBP2, an enzyme involved in cell wall synthesis. For FtsZ localization, MRSA strain BCBAJ020,24 a COL derivative expressing FtsZ-CFP ectopically from the spa locus under the control of the Pspac promoter, was induced with 0.5 mm IPTG.
Small molecule screening assay
We screened over 45 000 compounds7 from a pre-selected small molecule library at the ICCB-Longwood Screening Facility, a part of the New England Regional Centers of Excellence (NERCE), for inhibitory activity against MRSA USA300. We measured OD620 in a 384-well format both. We used 32 μg ml−1 cefoxitin as a positive control25 while cells grown in MHC alone were used as the negative control. Per CLSI protocol,26 the 384-well plates were grown without shaking at 37 °C for 24 h. Each well was scaled to respective positive and negative control to normalize the percent survival using the following equation:

Compounds yielding <50% survival with USA300 were considered as hits in the primary screen and further validated in the secondary screen where we looked for compounds with at least 80% inhibition in growth.
MIC by broth microdilution and macrodilution
The MICs of DNAC-2 was determined with USA300 and other MRSA clinical isolates according to CLSI guidelines.26 The MIC is defined as the lowest concentration of antibiotic inhibiting visible bacterial growth. To obtain the MIC of USA300 under relatively anaerobic (microaerophilic) conditions, cells were grown in 5 ml tubes without shaking for 15 h at 37 °C.20 MIC microdilution plates were set up as per CLSI protocol, incubated in the Bio-Bag environmental chamber (BD Diagnostics, Sparks, MD, USA) to promote anaerobic growth at 37 °C. To assess the effect of the compound on the electron transport chain, we supplemented the MHC broth with hemin or menadione (both at 2 μgml−1) and induced fermentative growth by adding sodium nitrate (5 mm) to the MHC medium (Sigma, St Louis, MO, USA).
Growth curves of USA300
USA300 cells were grown to exponential phase (OD620=0.5 using 18 mm borosilicate glass tube in a Spectronic 20 spectrophotometer) in MHB and back diluted 100-fold. Samples were treated with 4–32 μg ml−1 of DNAC-2 (diluted in DMSO), or with a combination of varying concentrations of DNAC-2 with ¼ × MIC of oxacillin (16 μg ml−1 or with DMSO alone as a control. The samples were set up in triplicate in a 96 well plate, incubated at 37 °C and the OD read at 650nm using the Tecan M1000 plate reader. The experiment was repeated 3 times. The results shown represent the mean and standard deviation from three replicates. Two way analysis of variance test (ANOVA) using Bonferroni’s multiple comparisons test in GraphPad Prism was applied to test significance, with a P-value of <0.05 considered significant.
Determination of time-kill curves using of DNAC-2
USA300 cells were grown to the exponential phase in MHB as described for the growth curve analysis and back diluted 100-fold. As described previously,7 samples were treated with 4–32 μg ml−1 of DNAC-2 (diluted in DMSO) or DMSO alone as the control. At specific time points (0, 2, 4, 6 and 24 h.), cells were washed free of compounds, diluted and plated on regular TSB agar with no antibiotic to enumerate the CFU, with each data point representing mean (CFU per ml) and standard deviation from three replicate experiments. Two way analysis of variance test (ANOVA) using Bonferroni’s multiple comparisons test in (GraphPad Prism GraphPad Software, La Jolla, CA, USA) was applied to test significance, with a P-value of <0.05 considered significant. Bactericidal activity is calculated by comparing the final CFU per ml of a particular time point to the initial CFU per ml at time zero.
Assessing the toxicity of DNAC-2: Three different methods were used to evaluate the cellular toxicity of DNAC-2. First, we exposed 4% sheep red blood cells (final concentrations) to increasing concentrations of DNAC-2 (ranging from ¼ –16 × MIC with twofold dilutions) and monitored for red cell lysis at OD540 in an in vitro hemolysis assay to assess the effect of DNAC-2 on membrane perturbation.27 Second, release of lactate dehydrogenase (LDH) from human bronchial epithelial cells (CFBE) exposed to DNAC-2 for 48 h was assayed. In brief, CFBE cells were maintained in RPMI 1640 medium (Sigma) with 10% fetal bovine serum and grown with 5% CO2 at 37 °C. Upon confluence, cells in the flask were released with trypsin, collected and counted. DNAC-2 at pre-determined concentrations (¼–16 × MIC) in serum free media was added to epithelial cells and incubated for 24 h at 37 °C. Triton X-100 at 2% final concentration was used as a positive control. Then 100 μl of the cytotoxicity reagents (Takara LDH cytotoxicity Detection kit Cat# MK401) was added to each well containing CFBE supernatant followed by incubation for 20 min in the dark and absorbance in each well read at 490nm. The LDH release was calculated as per manufacturer’s protocol. Third, inhibition of the human enzyme cytochrome P450 (CYP) by DNAC-2 in human liver microsomes was carried out by SRI Biosciences under the auspices of NIH Product Development Services. In brief, DNAC-2 was incubated with a cocktail of model CYP substrates specific for different CYP isoforms (phenactein for 1A2, bupropion for 2B6, diflofenac for 2C9, mepehenytoin for 2C19, bufuralol for 2D6, testosterone and midazolam for 3A4), human liver microsomes and co-factors for 20 min at 37 °C. Specific control inhibitors for different CYP isoforms (furafylline, thioTEPA. sulfaphenazole, nootkatone, quinide, ketoconazole) were also included. Formation of metabolites was measured by LC-MS/MS and compared to control incubations with no DNAC-2 or inhibitor. A decrease in metabolite formation in the presence of DNAC-2 indicated that the activity of the CYP isoform was inhibited under the condition used.
Pharmacokinetics of DNAC-2 in vitro
The in vitro metabolic stability of DNAC-2 was determined in human, rat, dog and mouse microsomes by SRI under NIH sponsorship. Briefly, DNAC-2 was incubated with active and heat inactivated human liver microsomes and co-factors at 37 °C. Aliquots were removed at 0, 15, 30, 60, 90 and 120 min and the amount of remaining DNAC-2 was determined using LC-MS/MS. The result was calculated as the percent of DNAC-2 remaining at a given time vs t=0 min. The in vitro half-life was calculated as t1/2=0.693/-k, where k is the slope of the linear regression of the natural log of the percent remaining vs time. Intrinsic clearance (CLint, μl min−1 mg−1) was calculated using the formula: CLint=(0.693/t1/2)/protein concentration in incubation.
Bidirectional permeability of DNAC-2 into Caco-2 cells was determined as follows: A CacoReady plate, consisting of Caco-2 cells already plated on a HTS Transwell 24-well plate, was purchased from ADMEcell, Inc. (Emeryville, CA, USA). DNAC-2 was prepared in transport buffer (pHs of 6.0 and 7.4) and added to the appropriate compartment (apical or basal) of the Transwell plate. At selected time points (0.5, 1, 1.5 and 2 h), aliquots were removed from the receiving compartment and analyzed by LC-MS/MS to determine the apparent permeability (Papp, × 10−6 cm s−1). Ketoconazole, a known inhibitor of the transporter protein P-gp, was added to control wells. Control incubations consisting of ganciclovir (not permeable), diazepam (permeable), and 3H-digoxin, a known substrate of P-gp, in the absence and presence of ketoconazole were also included and analyzed by liquid scintillation counting. The Efflux Ratio (ER) was calculated using the determined Papp (ER=Papp, B-A/Papp, A-B). An ER>2 in the absence of inhibitor indicates that the DNAC-2 is likely to be a substrate for P-gp. An ER<2 that is significantly reduced in the presence of inhibitor further confirms that the DNAC-2 is a substrate for P-gp. The % recovery of the test article was also determined. This was calculated by comparing the amount of DNAC-2 recovered during and at the end of the experiment with the amount of DNAC-2 added initially.
Plasma protein binding to DNAC-2 was determined by equilibrium dialysis for 4 h. at 37 °C using pooled human, Sprague Dawley rat, Beagle dog and BALB/C mouse plasma proteins.
Macromolecular analysis
The effect of DNAC-2 on macromolecular synthesis was studied by monitoring the incorporation of 3H- or 14C-labeled precursors (5-[3H] thymidine, [3H] glucosamine hydrochloride, [3H] uridine and l-[14C] isoleucine) as described previously.28 An overnight culture of Staphylococcus simulans 22 grown in MHC was diluted 50-fold into fresh medium and cultured at 37 °C to an OD600 of about 0.5. Cultures were aliquoted, diluted to an OD600 of 0.1 and allowed to regrow to an OD600 of 0.4. The respective labeled precursor was then added to each culture (final concentration 1 μCi ml−1); compound was added at 0.5, 1 or 2 × MIC, while another aliquot was run with 10X MIC of a control antibiotic and one without any antibiotic. Control antibiotics were vancomycin (3.1 μg ml−1) to inhibit cell wall synthesis, tetracycline (0.4 μg ml−1) to inhibit protein synthesis, ciprofloxacin (0.3 μg ml−1) to inhibit DNA synthesis and rifampicin (0.01 μg ml−1) to inhibit RNA synthesis. Incorporation of labeled precursors was monitored for up to 60 min and representative aliquots taken at 0, 5, 15, 30, 45 and 60 min. Macromolecules were precipitated with ice-cold TCA (10%) and incubated for at least 30 min on ice before being filtered through glass microfiber filters (Whatman). Filters were washed with 5 ml of TCA (2.5%) containing 10 mm unlabeled metabolite, dried, counted and the data expressed as mean of the counts incorporated from triplicate samples.
Transmission electron microscopy
USA300 cells grown in MHC at 37°C were treated with either DNAC-2 (2 × MIC) or DMSO for 90 min. The cells were washed twice with phosphate buffered saline (PBS) and processed for EM by fixing with 10 × volume of 2% glutaraldehyde-tannic acid (GTA)/1% paraformaldehyde in 0.1 m Na Cacodylate buffer (pH 7.4) followed by post-fixing in 1% OsO4 in sodium cacodylate buffer pH 7.4, embedded, serially dehydrated in ethanol, sectioned, stained with uranyl acetate and imaged using a JEOL TEM 1010 microscope at 100 kV at 1200 or × 15 000 magnification. For enumeration ~350 cells were evaluated for the presence or absence of the septum in the control and DNAC-2-treated conditions. Statistical significance was determined by Fisher’s exact test. Thirty fields of each strain with nearly equatorial cut surfaces were measured for cell wall thickness, and the results were expressed as a box and whiskers plot. Statistical significance was determined using the unpaired two tailed Student’s t-test, with a P-value of <0.05 considered significant.
Fluorescence microscopy
For fluorescence microscopy, strains were incubated overnight in TSB at 37 °C, supplemented with either erythromycin (10 μg ml−1) or kanamycin (200 μg ml−1) as needed, back diluted in fresh TSB and allowed to grow until mid-exponential phase (OD600~0.6). Each culture was then divided into five flasks with DNAC-2 (in DMSO) added to three at 0.5, 1 or 2 × MIC and the two remaining flasks kept as controls with DMSO or TSB alone. Cultures were incubated for 30 min, after which the cells were pelleted, washed in PBS buffer and mounted on microscope slides with pads of 1% agarose in PBS.
For staining of membrane, cell wall or DNA, cells were incubated with FM 4–64 (2 μg ml−1), BODIPY FL vancomycin (2 μg ml−1) or Hoechst 33342 (4 μg ml−1) (all from Molecular Probes) for 5 min at room temperature with shaking and washed before being imaged. Cells were imaged using a Zeiss Axio Observer microscope equipped with a Photometrics CoolSNAP HQ2 camera (Roper Scientific, Surrey, BC, Canada) and Metamorph 7.5 software (Molecular Devices, Sunnyvale, CA, USA) or by Structured Illumination Microscopy (SIM) or laser widefield microscopy in an ELYRA PS.1 Microscope (Zeiss, Oberkochen, Germany) with a sCMOS camera and 5 grating rotations for each channel. SIM images were reconstructed and analyzed with Zen Software (Zeiss). For quantification of the signal of fluorescent derivatives of PBP at the septa, five images were analyzed per condition using ImageJ (NIH, Bethesda MD USA). Only cells with a complete septum were analyzed.
Assay for membrane potential
The carbocyanine dye DiOC2 (3,3′-Diethyloxacarbocyanine iodide, Life Technologies, Carlsbad, Ca, USA) and lipophilic cationic tetraphenylphosphonium (TPP) were used in separate assays to assess membrane potenital as described previously.7, 28 For the DiOC2 study, S. aureus USA300 cultures were grown to the early exponential phase (OD620=0.3) in MHB, incubated with 10 μM of DiOC2 in PBS with 1% glucose at 24°C for 30 min in the dark and then transferred to a 384 well plate. Cells were analyzed at Ex485nm/Em680nm using the Tecan M1000 plate reader. After establishing a baseline reading, CCCP (positive control) or DNAC-2 was added to the wells and the drop in red fluorescence (Em680nm) was monitored over time. Significance was determined using the Kruskal–Wallis test with Dunn’s multiple comparisons in GraphPad Prism. For the determination of the membrane potential by means of the lipophilic cation TPP+ which diffuses rapidly across the bacterial membrane in response to a trans-negative Δψ, we use the S. simulans 22 strain following the procedure of Schneider et al.28 as described. Briefly, 1 μCi ml−1 of [3H]-tetraphenylphophonium bromide (TPP+; 26 Ci mMol−1) was added to the growth medium followed by the addition of DNAC-2. Samples were filtered through cellulose acetate filters (0.2 μm) and washed twice with phosphate buffer. Filters were dried and counted.28 For calculation of the membrane potential (Δψ), TPP+ concentrations were applied to the Nernst equation [(Δψ),=(2.3 × R × T/F) × log (TPP+in/TPP+out)]. Mean Δψ values were calculated from a minimum of two independent determinations.
RNA isolation and qRT–PCR
USA300 grown overnight (250 r.p.m., 37 °C) were inoculated into 10 ml of MHC adjusting to a starting OD650 of 0.05 and grown to an OD of~0.2 corresponding to ~1–3 × 108 CFU per ml. One hundred ml of the cells were treated with 1 × MIC of DNAC-2 for 10 min with shaking at 37 °C. The control flask had no drug. RNA was extracted from cells by resuspending the cell pellet in 1 ml Trizol and isolated with a reciprocating shaker with 0.1 mm glass beads as previously described (15). The extracted RNA was treated with DNase to remove any contaminating DNA and then repurified. cDNA was synthesized from total RNA using the Quanta qScript cDNA Supermix according to the manufacturer's protocol. qRT–PCR was performed in triplicate with S. aureus-specific primers designed for each gene of interest. For qRT–PCR, 3 different dilutions of the cDNA were used in duplicates for the reaction amplified for 40 cycles. The cDNA to each gene of interest was quantified based on cycle threshold (CT) compared to gyrB.
References
Spellberg, B., Bartlett, J., Wunderink, R. & Gilbert, D. N. Novel approaches are needed to develop tomorrow's antibacterial therapies. Am. J. Respir. Crit Care Med. 191, 135–140 (2015).
Haydon, D. J. et al. An inhibitor of FtsZ with potent and selective anti-staphylococcal activity. Science 321, 1673–1675 (2008).
Rasko, D. A. et al. Targeting QseC signaling and virulence for antibiotic development. Science 321, 1078–1080 (2008).
Payne, D. J. Microbiology. Desperately seeking new antibiotics. Science 321, 1644–1645 (2008).
Brown, D. Antibiotic resistance breakers: can repurposed drugs fill the antibiotic discovery void? Nat. Rev. Drug Discov. 14, 821–832 (2015).
Richter, S. G. et al. Small molecule inhibitor of lipoteichoic acid synthesis is an antibiotic for Gram-positive bacteria. Proc. Natl Acad. Sci. USA 110, 3531–3536 (2013).
Nair, D. R. et al. Characterization of a novel small molecule that potentiates beta-lactam activity against gram-positive and gram-negative pathogens. Antimicrob. Agents Chemother. 59, 1876–1885 (2015).
Silverman, J. A., Perlmutter, N. G. & Shapiro, H. M. Correlation of daptomycin bactericidal activity and membrane depolarization in Staphylococcus aureus. Antimicrob. Agents Chemother. 47, 2538–2544 (2003).
Dutter, B. F. et al. Decoupling Activation of Heme Biosynthesis from Anaerobic Toxicity in a Molecule Active in Staphylococcus aureus. ACS Chem. Biol. 11, 1354–1361 (2016).
Kahl, B. et al. Persistent infection with small colony variant strains of Staphylococcus aureus in patients with cystic fibrosis. J. Infect. Dis. 177, 1023–1029 (1998).
Magalon, A. et al. Inhibitor binding within the NarI subunit (cytochrome bnr) of Escherichia coli nitrate reductase A. J. Biol. Chem. 273, 10851–10856 (1998).
Toyofuku, M. et al. Influence of the Pseudomonas quinolone signal on denitrification in Pseudomonas aeruginosa. J. Bacteriol. 190, 7947–7956 (2008).
Centers for Disease Control and Prevention Active Bacterial Core Surveillance (ABCs): Emerging Infections Program Network Methicillin-Resistant Staphylococcus aureus, 2011 (2011).
Walsh, C. T. & Wencewicz, T. A. Prospects for new antibiotics: a molecule-centered perspective. J. Antibiot. (Tokyo) 67, 7–22 (2014).
Drlica, K. & Zhao, X. DNA gyrase, topoisomerase IV, and the 4-quinolones. Microbiol. Mol. Biol. Rev. 61, 377–392 (1997).
Cunningham, M. L., Kwan, B. P., Nelson, K. J., Bensen, D. C. & Shaw, K. J. Distinguishing on-target versus off-target activity in early antibacterial drug discovery using a macromolecular synthesis assay. J. Biomol. Screen. 18, 1018–1026 (2013).
Turner, R. D. et al. Peptidoglycan architecture can specify division planes in Staphylococcus aureus. Nat. Commun. 1, 26 (2010).
Sieradzki, K. & Tomasz, A. Inhibition of the autolytic system by vancomycin causes mimicry of vancomycin-intermediate Staphylococcus aureus-type resistance, cell concentration dependence of the MIC, and antibiotic tolerance in vancomycin-susceptible S. aureus. Antimicrob. Agents Chemother. 50, 527–533 (2006).
Pinho, M. G. & Errington, J. Recruitment of penicillin-binding protein PBP2 to the division site of Staphylococcus aureus is dependent on its transpeptidation substrates. Mol. Microbiol. 55, 799–807 (2005).
Mike, L. A. et al. Activation of heme biosynthesis by a small molecule that is toxic to fermenting Staphylococcus aureus. Proc. Natl Acad. Sci. USA 110, 8206–8211 (2013).
Thomas, V. C. et al. A central role for carbon-overflow pathways in the modulation of bacterial cell death. PLoS Pathog. 10, e1004205 (2014).
Cui, L. et al. Cell wall thickening is a common feature of vancomycin resistance in Staphylococcus aureus. J. Clin. Microbiol. 41, 5–14 (2003).
Fuchs, S., Pane-Farre, J., Kohler, C., Hecker, M. & Engelmann, S. Anaerobic gene expression in Staphylococcus aureus. J. Bacteriol. 189, 4275–4289 (2007).
Tan, C. M. et al. Restoring methicillin-resistant Staphylococcus aureus susceptibility to beta-lactam antibiotics. Sci. Transl. Med. 4, 126ra35 (2012).
Memmi, G., Filipe, S. R., Pinho, M. G., Fu, Z. & Cheung, A. L. Staphylococcus aureus PBP4 is essential for beta-lactam resistance in community acquired MRSA. Antimicrob. Agents Chemother. 52, 3955–3966 (2008).
Clinical and Laboratory Standards Institute Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria that Grow Aerobically; Approved Standard 9th ed. CLSI, Wayne, PA USA, (2012).
Saar, K. et al. Cell-penetrating peptides: a comparative membrane toxicity study. Anal. Biochem. 345, 55–65 (2005).
Schneider, T. et al. Plectasin, a fungal defensin, targets the bacterial cell wall precursor Lipid II. Science 328, 1168–1172 (2010).
Acknowledgements
We thank NERCE-BEID for facilitating this research, SRI Biosciences under the auspices of NIH Product Development Services for the pharmacokinetic and toxicity analyses, Louisa Howard for processing the TEM samples and imaging. HGS received support by the German Centre of Infection Research (DZIF). NERCE-BEID was funded by NIH grant U54AI057159. Work in the AC laboratory was partially funded by COBRE (NIH) # P30GM106394 and the Pfeiffer Foundation. MGP laboratory was funded by ERC-2012-StG-310987 grant from the European Research Council and JMM was supported by fellowship SFRH/BD/71993/2010.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on The Journal of Antibiotics website
Supplementary information
Rights and permissions
About this article
Cite this article
Nair, D., Chen, J., Monteiro, J. et al. A quinolinol-based small molecule with anti-MRSA activity that targets bacterial membrane and promotes fermentative metabolism. J Antibiot 70, 1009–1019 (2017). https://doi.org/10.1038/ja.2017.79
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2017.79