Abstract
During the course of screening for translocase I inhibitors, the new liposidomycin-related compounds, A-90289 A and B, were isolated from a culture broth of Streptomyces sp. SANK 60405. The structural elucidations were carried out by NMR and high-resolution mass spectral analyses, and they were classified as members of the liponucleoside antibiotics group with a sulfate group at the C-2′ position. A-90289 A and B inhibited bacterial translocase I with IC50 values of 36.5 ng ml−1 and 33.8 ng ml−1, respectively.
Similar content being viewed by others
Introduction
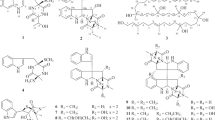
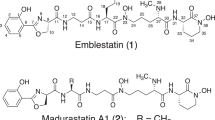
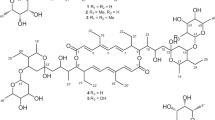
Translocase I has been intensely investigated as a prime target in the development of new antibacterial agents that possess this mechanism of action. In fact, a number of translocase I inhibitors,1 including mureidomycins,2 pacidamycins,3 napsamycins,4 liposidomycins,5, 6 tunicamycin,7 capuramycins,8, 9 muraymycins,10 caprazamycins,11, 12 A-102395,13 A-94964,14, 15 muraminomicins16 and A-97065s,17 have been reported and focused on. The structures of liposidomycin-related compounds are composed of a uridine moiety, a 5-amino ribose moiety, a diazepanone ring system, a 3-methyl glutaric acid moiety and a fatty acid moiety. Though many of them, which were produced with some homologs, simply differed from each other in the variety of fatty acid moieties, some antibiotics that have minor components with modified core structures such as liposidomycins (C-2′′ O-sulfated compounds), A-97065 (C-2′′ O-methylated compounds) and A-97065 B and D (C-3′′ O-phosphated compounds) are known. Herein, we report the isolation, structural elucidation and biological activity of the novel C-2′ O-sulfated liposidomycin-related compounds, A-90289 A (1) and B (2) (Figure 1).
Results
Taxonomy of the producing strain
Strain SANK 60405 formed an extensively branched substrate mycelium and aerial hypae that carried spiral spore chains with a spiny surface (Figure 2). The spore mass was gray to brownish grey on ISP-2 medium. The whole-cell hydrolysates of the strain contained L,L-diaminopimelic acid. An almost complete 16S rDNA sequence of the strain SANK 60405 was determined and deposited at the DNA Data Bank of Japan with the accession number AB448716. The resulting 16S rDNA sequence was certainly clustered into the genus Streptomyces by means of molecular phylogenetic analysis.14
On the basis of taxonomic properties and the 16S rDNA sequence, the strain was identified as Streptomyces sp. SANK 60405.
Physicochemical properties
The physicochemical properties of 1 and 2 are summarized in Table 1. These compounds, which are soluble in organic solvents such as EtOAc, acetonitrile, dimethylsulfoxide and acetone, were obtained as colorless powders. Both compounds showed identical UV spectra with an absorption maximum at 259 nm. The characteristic absorption bands of IR spectra were derived from the carbonyl group (around 1713 cm−1) and amide carbonyl groups in 1 (1639 cm−1) and 2 (1634 cm−1).
Structure elucidation of A-90289 A (1)
The 1H and 13C NMR assignments of 1 are listed in Table 2. The 1H NMR spectrum of 1 contained 79 proton resonances. The 13C NMR spectrum of 1 showed 53 carbon signals, which were classified into 8 methyl groups, 17 methylenes, 19 methines, 2 olefins and 7 carbonyl carbons by an analysis of DEPT spectra. The extensive connectivity of 1 was established through the DQF-COSY, HSQC-HOHAHA and HMBC experiments as shown in Figure 3. These analyses proved that 1 had a uridine moiety, a 5-amino-5-deoxypentofuranose moiety, a 3-methylglutaric acid moiety and a 6-deoxy-2,3,4-tri-O-methylhexopyranose moiety. These features of 1 were the same as those of caprazamycins. Neither HMBC correlation from H-2′′′ (δH 4.16) nor H-3′′′ (δH 5.33) to C-1′′′ carbonyl carbon was observed on the partial structure elucidated as the diazepanone ring system. However, the carbonyl carbon of δC 171.5 was assigned to the C-1′′′ position because the 13C chemical shift values of the signals from the diazepanone ring system of 1 corresponded well to those of caprazamycin12; the signal of δC 67.1 corresponded to C-2′′′ (δC 63.9), similarly it corresponded to δC 73.7 to C-3′′′ (δC 71.4), δC 58.3 to C-4′′′ (δC 57.1) and δC 171.5 to C-7′′′ (δC 170.8). In addition, there were no candidates for substituting the C-2′′′ position except for δC 171.5 carbonyl carbon. The structures regarding the 3-hydroxyhexadecanoic acid moiety were also elucidated by DQF-COSY and HMBC correlations between an oxymethine proton at H-3a (δH 5.12) and an ester carbon at C-1a (δC 171.1); methylene carbons at C-2a (δC 39.4) and C-5a (δC 26.3); DQF-COSY, HMBC and HSQC-HOHAHA correlations of C-13a through C-16a and the number of unassigned carbon signals.
The connectivities of these partial structures were revealed by the correlations in DQF-COSY and HMBC spectrum, as shown in Figure 3. The connectivity of the uridine moiety and 5-amino-5-deoxypentofuranose was established by the HMBC correlation of H-5′ (δH 4.17) with C-1′′ (δC 112.0) and that of H-1′′ (δH 5.07) with C-5′ (δC 77.4). The connection between the uridine moiety and the diazepanone ring was determined by the 1H-1H coupling between H-5′ (δH 4.17) and H-6′ (δH 3.73) in the DQF-COSY spectrum. The linkages between the diazepanone ring system, 3-hydroxyhexadecanoic acid, 3-methylglutaric acid and 6-deoxy-2,3,4-tri-O-methylhexopyranose moieties were elucidated by the HMBC correlations between H-3′′′ (δH 5.33) and C-1a (δC 171.1), H-3a (δH 5.12) and C-1b (δC 173.1), and H-1c (δH 4.41) and C-5b (δC 172.1).
Thus, the core structure of 1 was established as illustrated in Figure 3.
The residual substituent of 1 was mainly determined by ESI mass spectrometry. The molecular formula of 1 was estimated to be C53H87N5O25S or C53H88N5O25P by high-resolution ESI mass spectral analysis and by the above NMR results. Hence, the structure of 1 was expected to have a sulfate group or a phosphate group. The high-resolution ESI mass spectrum (Figure 4a) of 1 showed a fragment ion of m/z 1146.59082 due to the neutral loss of 79.95715 units from [M+H]+ at m/z 1226.54797. Considering each mass accuracy, the loss corresponded to the elimination of SO3 (79.95681 U, 1.3 p.p.m.) and not to the elimination of PO3H (79.96633 U, −117.7 p.p.m.), establishing the presence of a sulfate group.
Then, we determined the sulfate position as follows. In the study on 2′′-O-sulfated liposidomycin,6 the H-2′′methine proton (δH 4.67) was downfield shifted by 0.4–0.5 p.p.m. in comparison with the other typical oxymethine protons in the uridine and 5′′-amino-5′′-dehydroxypentofuranose moieties. In the 1H NMR spectrum of 1, the H-2′ (δH 4.70) signal was significantly downfield shifted by 0.5–0.7 p.p.m. This indicated that C-2′ of 1 was O-sulfated. The substituted position was finally confirmed by FAB mass spectral analysis as shown in Figure 5. In the negative ion mode FAB mass spectrum of 1, the characteristic fragment ion at m/z 592, which arose by the loss of CO2 and the lipid moiety, was observed. In the negative ion mode FAB/CID spectrum of uridine, a sugar fragment denominated S1 at m/z 153 corresponding to cleavages at C-1′-O and C-2′-C-3′ was reported by Crow et al.18 Then, we performed a linked scan experiment using m/z 592 as a precursor ion, and a product ion (3) at m/z 233 was observed. In our case, no S1 fragment ion at m/z 153 was observed, but a fragment ion at m/z 233 was observed. The difference of 80 U between m/z 153 and m/z 233 clearly indicated that it was because of the the substitution of a sulfate group at the C-2′ position. Furthermore, high-resolution analysis of this ion revealed its elemental composition as C6H5N2O6S (233.9868 U, 0 p.p.m.) and this agreed well with the composition of 3. Accordingly, the O-sulfated position was determined to be C-2′.
Thus, the structure of 1 was determined as shown in Figure 1.
The absolute configuration of 1 could not be determined because of the shortage of 1 for X-ray crystallography or chemical modification. However, the relative configuration of the uridine moiety of 1 was suggested to be the same as the known liposidomycin-related compounds because the observed NOE correlations between H-6 and H-2′, H-6 and H-3′ and H-6 and H-6′ agreed well with those of liposidomycin.6
Structure elucidation of A-90289 B (2)
The 1H and 13C NMR assignments of 2 are listed in Table 2. The 1H NMR spectrum contained 79 proton resonances, and 53 carbon signals were observed in the 13C NMR spectrum. These chemical shifts corresponded well to those of 1. In the high-resolution ESI mass spectrum as shown in Figure 4b, the protonated molecule (m/z 1226.54761) and fragment ion (m/z 1146.59131) were observed, suggesting that 2 also had a sulfate group. By comparison with their spectra, 2 would have had the same structure as 1, except for a fatty acid moiety. The results of the various 2D-NMR experiments of 2 are illustrated in Figure 6. These correlations proved that 2 had the same core structure as 1. In the 1H NMR of 2, two doublet methyl proton signals (δH 0.78 × 2) and a multiplet methine signal (δH 1.43) coupled with each other were characteristically observed. This suggested that the penultimate carbon of the fatty acid in 2 was methylated. From the DQF-COSY and HMBC correlations and the number of unassigned carbon signals, the structure of the fatty acid moiety in 2 was determined to be 3-hydroxy-14-methylpentadecanoic acid. As described above, the observation of a characteristic product ion at m/z 233 in the linked-scan experiment proved that 2 also had a sulfate group at the C-2′ position. Thus, the structure of 2 was determined as shown in Figure 1.
The stereochemistry of 2 was suggested to be the same as 1 because the NOE correlations of 2 corresponded to those of 1.
Biological properties of A-90289 A and B
The inhibitory activities of A-90289 A and B on bacterial translocase I were evaluated by comparing with that of muraminomicin F. As shown in Table 3, A-90289 A and B inhibited bacterial translocase I with IC50 values of 36.5 ng ml−1 and 33.8 ng ml−1, respectively, whereas muraminomicin F inhibited the enzyme with an IC50 value of 12.8 ng ml−1.16
The antimicrobial activities (MIC) of A-90289 A against Staphylococcus aureus ATCC 6538P, Streptococcus pyogenes and Enterococcus faecium were 8 μg ml−1, 4 μg ml−1 and 16 μg ml−1, respectively (Table 4). The antimicrobial activities of A-90289 B were not tested because of the depletion of sample.
Discussion
During a screening program of bacterial translocase I inhibitors, we found strong inhibitory activity in a culture broth of Streptomyces sp. SANK 60405 and isolated the new liposidomycin-related compounds, A-90289 A (1) and B (2) (Figure 1). Although a number of C-2′′ O-sulfated compounds have been reported as a member of the liposidomycin family, C-2′ O-sulfated liposidomycin-related compounds have not been reported yet. The translocase I inhibitory activities of A-90289 A and B are as potent as those of reported liposidomycin-related compounds, such as muraminomicins and A-97065s. It was indicated that the presence of a hydroxyl group at the C-2′ position was not crucial for translocase I inhibitory activity. A-90289 A exhibited antimicrobial activities against clinically significant Gram-positive bacteria. As A-90289 A shows nearly the same translocase I inhibitory activity as A-90289 B, it is expected that A-90289 B possesses an equal antimicrobial potential because the fatty acid analogs of liposidomycin and muraminomicin were reported to have equal antimicrobial activities,16, 19, 20 and the difference of the fatty acid moieties seemed to have no influence on the antibacterial activities. Therefore, not only muraminomicins and A-97065s, but also A-90289 A and B might be candidates for new antibacterial agents.
To date, all the reported liposidomycins have a sulfate group at their C-2′′ position. In the point of view of molecular diversity, the discovery of C-2′ O-sulfated liposidomycin-related compounds is noteworthy. The specific C-2′ O-sulfotransferase might lead to the generation of liposidomycin-related compounds, giving a chance to widen structural diversity besides the modifications of the core structure. Recently, the biosynthetic gene cluster of A-90289s has been cloned and reported.21 In vivo and in vitro studies were conducted to elucidate the LipB function, which was related to the incorporation of the sulfate group at C-2′. Genetically engineered LipB might allow us to create various O-sulfated compounds. They would be important for developing a structure–activity relationship. In the field of new drug discovery, one of the key points is to endow a compound with good pharmacokinetic features. Therefore, an introduction of the sulfate group catalyzed by sulfotransferase with broad substrate specificity would be useful in the case of improving some physicochemical properties such as solubility, permeability and so on, which is similar to the case of an O-sulfated echinocandin analog WF11899.22
Methods
Taxonomy of the producing organism
The producing organism, strain SANK 60405, was isolated from a soil sample collected in Tokyo, Japan. The morphological characterizations of the producing organism were performed on yeast extract-malt extract agar (ISP medium no. 2) incubated at 28 °C for 14 days. The cell walls and whole-cell hydrolysates were analyzed by the methods of Hasegawa et al.23 The 16S rDNA was PCR amplified from an isolated genome DNA and sequenced. The most related sequences were searched using the BLAST (basic local alignment search tool) algorithm in the National Center for Biotechnology Information.
Phylogenetic analysis of the 16S rDNA sequences was performed according to the method of Nakagawa and Kawasaki.24 A phylogenetic tree was constructed using the neighbor-joining methods25 in MEGA, version 4.0.26
General experimental procedures
HPLC analysis of A-90289 A and B was performed on an HP1100 system (Agilent, Tokyo, Japan) with a Unison UK-C18 column (4.6.φ × 75 mm, Imtakt Co. Ltd, Kyoto, Japan), at a flow rate of 1.0 ml min−1. The effluent was monitored at an absorbance of 260 nm. The chromatography was conducted with 30% acetonitrile in water containing 10 mM HCOONH4–HCOOH (pH 3.5) for 2 min, then with a linear gradient from 30 to 70% acetonitrile in water containing 10 mM HCOONH4–HCOOH (pH 3.5) for 10 min and finally maintained at 70% acetonitrile in water containing 10 mM HCOONH4–HCOOH (pH 3.5) for 2 min.
All the NMR spectra were recorded at 300 K on a DRX 500 spectrometer (Bruker BioSpin, Kanagawa, Japan) equipped with a cryogenic probe operating at 500 MHz for 1H and 125 MHz for 13C NMR spectra. The samples for NMR characterization were dissolved in methanol-d4/dimethylsulfoxide-d6/deuterium oxide (10:10:1). The optical rotation and IR spectra were measured with a DIP-370 (JASCO, Tokyo, Japan) and a FT-IR 8300 (Shimadzu, Kyoto, Japan), respectively. High-resolution ESI mass spectra were recorded on an LTQ-Orbitrap XL (Thermo Fisher Scientific, Kanagawa, Japan). High-resolution and linked-scan FAB mass spectra were determined by a JMS-700QQ mass spectrometer (JEOL, Tokyo, Japan). The UV spectra were measured by a UV-265FW spectrometer (Shimadzu).
Fermentation and isolation
To acquire 1 and 2, the producing organism Streptomyces sp. SANK 60405 grown on an ISP-2 plate was inoculated into 500-ml baffled flasks containing 100 ml of seed medium, which consisted of glucose, 0.3%; soybean meal, 0.3%; yeast extract, 0.03%; CaCO3, 0.04%; MgSO4 7H2O, 0.02%; and CB-442, 0.005%, and was cultured at 28 °C for 3 days. Aliquots (290 ml) of the seed culture were each transferred to individual 2-l baffled flasks, which contained 400 ml of production medium, which was the same composition as the seed medium, and the fermentation was carried out at 28 °C for 7 days.
Fermentation broth (4 l) was harvested, and an equal volume of acetone was added to extract the desired compounds. The mixture was filtered with an addition of Celite545 (KANTO CHEMICAL Co., Ltd, Tokyo, Japan) as an adjunct. After an addition of 76 g of NaCl, the filtrate (pH 3) was extracted with 3 l of EtOAc, and the recovered water layer was re-extracted twice by adding 2.4 l of mixed solvent, acetone/EtOAc (1:1). Afterward, 260 ml of 0.2 M phosphate buffer (pH 7) was added to the above combined extract (13 l), and the resulting whole emulsion was concentrated in vacuo to 2 l. The concentrate was adjusted to pH 6.8, and the water layer was separated from the organic layer. Resulting water layer (1.2 l) was concentrated in vacuo to 1.1 l and loaded onto a Diaion HP-20 column (200 ml, Mitsubishi Chemical, Tokyo, Japan). After the column was washed with water (600 ml), an active substance was eluted with 600 ml of 70% aqueous acetone. The eluate was concentrated to dryness and dissolved in 20 ml of 50 mM phosphate buffer (pH 6.8), and then applied to a Cosmosil 140 C18 column (200 ml, Nacalai Tesque Co. Ltd, Kyoto, Japan). After the column was washed with 50 mM phosphate buffer (pH 6.8), chromatography was performed by stepwise elution using 30, 35, 40, 45 and 50% acetonitrile in water containing 50 mM phosphate buffer (600 ml each), and was then fractionated into 100 ml portions. By HPLC analysis, 1 and 2 were detected in the 50% acetonitrile fraction from 2 to 4. The combined fraction was desalted with a Diaion HP-20 (Mitsubishi Chemical) as described above and concentrated to dryness to yield 46.0 mg of crude powder. This was dissolved in 55% acetonitrile in water containing 0.3% triethylamine-phosphate buffer (pH 3) and the solution was subjected to a Symmetry C18 column (19φ × 100 mm, Waters, Tokyo, Japan). The chromatography was performed with 55% acetonitrile in water containing 0.3% triethylamine-phosphate buffer (pH 3) at a flow rate of 10 ml min−1 with UV detection at 260 nm. As a result, 1 and 2 were eluted at the retention times of 26 and 23 min, respectively. Each fraction, desalted by a Diaion HP-20 column (Mitsubishi Chemical), was concentrated and freeze-dried to yield 8.2 mg of A-90289 A (1) and 1.5 mg of A-90289 B (2) in pure forms.
Measurement of translocase I inhibitory activity
Translocase I inhibitory activity was measured using a 96-well microtiter plate-based assay. Assay mixture composed of 100 mM Tris-HCl (pH 7.5), 50 mM KCl, 25 mM MgCl2, 0.8% Triton X-100, 166 μM undecaprenyl phosphate and 70 μM UDP-MurNAc-L-Ala-γ-D-Glu-m-DAP-[Nɛ-dansyl]-D-Ala-D-Ala in 100 μl solution was prepared in each well. The reaction was initiated by an addition of enzyme (0.625–2.5 μg protein). The enzyme activity was monitored by measuring an increase of fluorescence detected at 535 nm (excitation at 355 nm) after 30 min incubation at 30 °C
Antimicrobial activities
The MICs were determined by a standard broth microdilution method.27
Accession codes
References
Kimura, K. & Bugg, T. D. H. Recent advances in antimicrobial nucleoside antibiotics targeting cell wall biosynthesis. Nat. Prod. Rep. 20, 252–273 (2003).
Inukai, M. et al. Mureidomycin A-D, novel peptidylnucleoside antibiotics with spheroplast forming activity. I. Taxonomy, fermentation, isolation and physicochemical properties. J. Antibiot. 42, 662–666 (1989).
Karwowski, J. P. et al. Pacidamycins, a novel series of antibiotics with anti-Pseudomonas aeruginosa activity. I. Taxonomy of the producing organism and fermentation. J. Antibiot. 42, 506–511 (1989).
Chatterjee, S. et al. Napsamycins, new Pseudomonas active antibiotics of mureidomycin family from Streptomyces sp. HIL Y-82, 11372. J. Antibiot. 47, 595–598 (1994).
Ubukata, M. & Isono, K. The structure of liposidomycin B, an inhibitor of bacterial peptidoglycan synthesis. J. Am. Chem. Soc. 110, 4416–4417 (1988).
Ubukata, M. et al. Structure elucidation of liposidomycins, a class of complex lipid nucleoside antibiotics. J. Org. Chem. 57, 6392–6403 (1992).
Takatsuki, A., Arima, K. & Tamura, G. Tunicamycin, a new antibiotic. I. Isolation and characterization of tunicamycin. J. Antibiot. 24, 215–223 (1971).
Yamaguchi, H. et al. Capuramycin, a new nucleoside antibiotic. Taxonomy, fermentation, isolation and characterization. J. Antibiot. 39, 1047–1053 (1986).
Seto, H. et al. The structure of a new nucleoside antibiotic, capuramycin. Tetrahedron Lett. 29, 2343–2346 (1988).
McDonald, L. A. et al. Structures of the muraymycins, novel peptidoglycan biosynthesis inhibitors. J. Am. Chem. Soc. 124, 10260–10261 (2002).
Igarashi, M. et al. Caprazamycn B, a novel anti-tuberculosis antibiotic, from Streptomyces sp. J. Antibiot. 56, 580–583 (2003).
Igarashi, M. et al. Caprazamycns, novel lipo-nucleoside antibiotics, from Streptomyces sp. II. Structure elucidation of caprazamycins. J. Antibiot. 58, 327–337 (2005).
Murakami, R. et al. A-102395, a new inhibitor of bacterial translocase I, produced by Amycolatopsis sp. SANK 60206. J. Antibiot. 60, 690–695 (2007).
Murakami, R. et al. A-94964, novel inhibitor of bacterial translocase I, produced by Streptomyces sp. SANK 60404. I. Taxonomy, fermentation, isolation and biological activity. J. Antibiot. 61, 537–544 (2008).
Fujita, Y., Murakami, R., Muramatsu, Y., Miyakoshi, S. & Takatsu, T. A-94964, novel inhibitor of bacterial translocase I, produced by Streptomyces sp. SANK 60404. II. Physico-chemical properties and structure elucidation. J. Antibiot. 61, 545–549 (2008).
Muramatsu, Y. et al. New antibiotic muraminomicin. Jpn. Kokai Tokkyo Koho 2004-196780, 15 July (2004).
Fujita, Y., Kizuka, M. & Murakami, R. New compound A-97065. Jpn. Kokai Tokkyo Koho 2008-074710, 3 April (2008).
Crow, F. W., Tomer, K. B., Gross, M. L., McCloskey, J. A. & Bergstrom, D. E. Fast atom bombardment combined with tandem mass spectrometry for the determination of nucleosides. Anal. Biochem. 139, 243–262 (1984).
Isono, K. et al. Liposidomycins: novel nucleoside antibiotics which inhibit bacterial peptidoglycan synthesis. J. Antibiot. 38, 1617–1621 (1985).
Kimura, K. et al. New types of liposidomycins that inhibit bacterial peptidoglycan synthesis and are produced by Streptomyces. II Isolation and structure elucidation. J. Antibiot. 51, 647–654 (1998).
Funabashi, M. et al. The biosynthesis of liposidomycin-like A-90289 antibiotics featuring a new type of sulfotransferase. Chembiochem 11, 184–190 (2010).
Iwamoto, T. et al. WF11899A, B and C, novel antifungal lipopeptides I. Taxonomy, fermentation, isolation and physico-chemical properties. J. Antibiot. 47, 1084–1091 (1994).
Hasegawa, T., Takizawa, M. & Tanida, S. A rapid analysis for chemical grouping of aerobic actinomycetes. J. Gen. Appl. Microbiol. 29, 319–322 (1983).
Nakagawa, Y. & Kawasaki, H. Identification Manual of Actinomycetes. (eds The Society for Actinomycetes. Japan) 83–117 (Business Center for Academic Societies Japan, Tokyo, 2001).
Saitou, N. & Nei, M. The neighbor-joining method: a new method of constructing phylogenetic tree. Mol. Biol. Evol. 6, 514–525 (1987).
Tamura, K., Dudley, J., Nei, M. & Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0. Mol. Biol. Evol. 24, 1596–1599 (2007).
Clinical and Laboratory Standards Institute. Methods for dilution antimicrobial susceptibility test for bacteria that grow aerobically; approved standard, 7th ed. CLSI document M7-A7 (Clinical and Laboratory Standard Institute, Wayne, PA, 2006).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fujita, Y., Kizuka, M., Funabashi, M. et al. A-90289 A and B, new inhibitors of bacterial translocase I, produced by Streptomyces sp. SANK 60405. J Antibiot 64, 495–501 (2011). https://doi.org/10.1038/ja.2011.38
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2011.38
Keywords
This article is cited by
-
Peculiarities of promiscuous l-threonine transaldolases for enantioselective synthesis of β-hydroxy-α-amino acids
Applied Microbiology and Biotechnology (2021)
-
Pyridoxal-5′-phosphate-dependent alkyl transfer in nucleoside antibiotic biosynthesis
Nature Chemical Biology (2020)
-
Muraminomicins, novel ester derivatives: in vitro and in vivo antistaphylococcal activity
The Journal of Antibiotics (2019)
-
Liposidomycin, the first reported nucleoside antibiotic inhibitor of peptidoglycan biosynthesis translocase I: The discovery of liposidomycin and related compounds with a perspective on their application to new antibiotics
The Journal of Antibiotics (2019)
-
Muraminomicins, new lipo-nucleoside antibiotics from Streptosporangium sp. SANK 60501-structure elucidations of muraminomicins and supply of the core component for derivatization
The Journal of Antibiotics (2019)