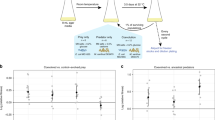
Abstract
Resource limitation and predation mortality are major determinants of microbial population dynamics, and optimization for either aspect is considered to imply a trade-off with respect to the other. Adaptation to these selective factors may, moreover, lead to disadvantages at rich growth conditions. We present an example of a concomitant evolutionary optimization to both, substrate limitation and predation in an aggregate-forming freshwater bacterial isolate, and we elucidate an underlying genomic mechanism. Bacteria were propagated in serial batch culture in a nutrient-restricted environment either with or without a bacterivorous flagellate. Strains isolated after 26 growth cycles of the predator–prey co-cultures formed as much total biomass as the ancestor at ancestral growth conditions, albeit largely reallocated to cell aggregates. A ~273 kbp genome fragment was lost in three strains that had independently evolved with predators. These strains had significantly higher growth yield on substrate-restricted media than others that were isolated from the same treatment before the excision event. Under predation pressure, the isolates with the deletion outcompeted both, the ancestor and the strains evolved without predators even at rich growth conditions. At the same time, genome reduction led to a growth disadvantage in the presence of benzoate due to the loss of the respective degradation pathway, suggesting that niche constriction might be the price for the bidirectional optimization.
Similar content being viewed by others
Introduction
Baas-Becking’s classic proposition about the ubiquitous dispersal but concomitant environmental filtering of microorganisms (Baas-Becking, 1934) implies that bacteria are incessantly transported between habitats or biomes. Such transitions are likely accompanied by a qualitative or quantitative change of selective factors, often more than one. Terrestrial microbes are continuously introduced into freshwaters (Ruiz-González et al., 2015), where they face both reduced substrate availability (Guildford and Hecky, 2000) and high mortality due to free-living protistan predators (Pernthaler, 2005). Adaptations to the former constraint include motility to track patchy substrates (Mitchell and Kogure, 2006), or the investment in high-affinity substrate uptake systems (Tang et al., 2012). Resistance to grazing can be realized by a variety of mechanisms, including the formation of protist-inedible morphotypes or of cell aggregates (Corno and Jürgens, 2008). Both optimizations may restrict the ability of planktonic bacteria to perform at rich growth conditions, for example, common freshwater genera such as Polynucleobacter and Limnohabitans need to be gradually acclimatized to higher substrate levels before isolation (Hahn et al., 2004; Kasalicky et al., 2010). In addition, these two strategies are widely considered to reflect an obligatory trade-off between high mortality rates and high defense costs and, therefore, to be mutually exclusive (Lenski and Levin, 1985). However, it has also been proposed that some bacteria may successfully overcome the dilemma of simultaneous ‘bottom up’ limitation and high predation risk (Thingstad et al., 2005). So far, this so-called ‘Winnie-the-Pooh’ strategy is, by and large, a conceptual model backed by a limited set of field observations. Here we show that, starting with inducible predation resistance (Blom et al., 2010b), it is possible to experimentally evolve bacteria that adopt such a strategy, that is, a simultaneously improvement in growth efficiency and predation resistance, even leading to superior competitive abilities at ancestral growth conditions. We subsequently elucidate and experimentally confirm an underlying genomic mechanism.
Materials and methods
Evolution experiment
Sphingobium sp. Z007 (16 S rRNA gene accession number: FN293045), originally isolated from Lake Zurich (Blom et al., 2010b) was evolved for 200 days (approximately 600 generations; 29 growth cycles) in sequential batch culture in Artificial Lake Water (ALW) medium (Zotina et al., 2003) containing 1 mg l−1 of glucose, yeast extract and soytone peptone (that is, 0.1% of the substrate concentrations on which the strain had been maintained before). Bacterial cultures were grown at 18 °C in triplicates of 100 ml, either with (P+) or without (P−) the bacterivorous flagellate Poterioochromonas sp. DS (Hahn and Höfle, 1999). Previous experiments with this predator–prey system revealed that the bacterial strain responds to a chemical cue from the flagellate with enhanced formation of cell aggregates (Blom et al., 2010a). Every 168 h, the cultures were 1:10 diluted into fresh ALW medium for the subsequent growth cycle. Five strains from all parallel treatments were isolated biweekly on solid DSMZ 7 medium and stored at −80 °C. Eight strains per treatment were isolated from the last time point (growth cycle 26).
Genome sequencing, assembly and annotation
Altogether six bacterial strains were genome sequenced: the ancestor and five evolved strains isolated from different experimental replicates (three from P+, two from P−) after 26 re-inoculation cycles. The ancestor strain was first sequenced on the Pacific Bioscience RS platform at the Functional Genomics Center Zurich, and all six strains were sequenced with the Illumina MySeq platform in the Genetic Diversity Centre at ETH Zurich. Library preparation was done with PacBio Template Prep Kit and the truseq DNA PCR-free kit (Illumina Inc., San Diego, CA, USA), respectively. To produce a high-quality reference genome of the ancestor strain, the corresponding Illumina reads served for error correction of the de novo assembly from the Pacific Bioscience sequencing using CLC Genomics Workbench 7.0 (CLC bio, Aarhus, Denmark). Gene calling and annotation of the reference genome was then performed with the RAST automated annotation pipeline (Aziz et al., 2008; Overbeek et al., 2014) and refined with the Artemis genome browser (Rutherford et al., 2000). In a second step, Illumina reads from the other strains were mapped to the ancestor reference genome and analysed with CLC Genomics Workbench 7.0. For the detection of single-nucleotide polymorphisms and of insertion and deletions (InDels) across all the evolved genome-sequenced strains, the variant caller tool of CLC Genomics Workbench 7 was used using default settings with a minimum coverage of five reads and a minimum frequency of 70%. Stand-alone versions of Blast (Altschul et al., 1990) and Dotter (http://www.ubuntu.com) were used for an in-depth analysis of differences between ancestor and evolved strains. Genomes of all sequenced isolates were submitted to European Nucleotide Archive under the accession no. PRJEB12244.
Verification of excision
To confirm the excision in strains of the P+ treatments, four primers were designed that target its flanking regions (Supplementary Table S1). DNA of the ancestor and evolved strains grown for 72 h in DSMZ 7 medium was extracted with the bacterial genome extraction Gen elute Kit (Sigma, Buchs, Switzerland). Three separate PCR reactions were performed for each strain (see Supplementary Figures S1A and B for details): two with the flanking primers for each side of the excision to verify its occurrence and one with the forward primer of the left side and with the reverse primer of the right side as an indicator of its absence. Colony PCR was performed on strains from growth cycle 26, to check whether the excision had spread through the whole population in the P+ treatment, and whether this excision was also present in any strains of the P− treatments. For this, eight additional isolates per biological replicate were tested for the presence and absence of the excision (N=40 per treatment) using the primers described above.
Growth experiments on substrate-restricted and benzoate media
Growth experiments were performed using strains isolated from each of the biological replicates of the P+ treatment. Three such sets of three independently evolved strains were tested that originated (i) from the early second half of the experiment (16 growth cycles), (ii) from immediately before the excision (isolated after 24 (PA), 20 (PB), 22 (PC) growth cycles) and (iii) from after the excision (26 growth cycles). Growth experiments with each individual isolate were carried out in technical triplicates. The strains were inoculated directly from frozen stock cultures into 50 ml of DSMZ 7 medium. After 72 h, the strains were transferred into ALW medium and grown for another 24 h. Thereafter, 5 × 105 bacterial cells ml−1 were inoculated into 50 ml of ALW medium either without flagellate predator or with 8 × 103 Poterioochromonas sp. DS cells ml−1. Bacterial cell densities were determined after 96 h of growth in the dark at 18 °C.
The ancestor and three independently evolved bacterial strains isolated after 26 growth cycles from each replicate of the evolution experiment (that is, the strains that had been genome sequenced) were first pre-cultivated from frozen stock cultures as described above. A total 5 × 105 bacteria ml−1 of each strain were then inoculated into triplicate 300 ml vessels with 50 ml ALW medium and grown in the dark at 18 °C either with or without benzoate (1 g l−1, Sigma). The cell densities were determined every 24 h for 72 h.
Competition assays on rich media
The ancestor and three strains isolated from separate vessels of the P+ and P− treatments after 26 growth cycles were pre-grown in DSMZ 7 medium for 72 h as described above. Thereafter, strains were grown in DSMZ 7 medium at an initial bacterial cell density of 2.5 × 105 cells ml−1, either without or with 104 predator cells ml−1. In the competition assays, each strain from P+ was either grown in triplicates with the ancestor or fully factorial with each strain from P− (n=9). Quantification of cell densities and DNA extraction (Bacterial DNA extraction kit, Sigma) was carried out at the beginning of the experiment, after 48 and 72 h, respectively.
The cell ratios of competing strains was determined by quantitative PCR. The primers for the 16 S rRNA region of Sphingobium sp. Z007 were designed to determine total cell numbers. To distinguish strains from different evolutionary background, a second primer pair was constructed for a single copy gene involved in the metabolism of aromatic compounds (nitriloacetate monoxygenase component A, ntaA). It is situated on the excision and is, therefore, absent in all P+ strains. The specificity of the primers was tested in pure cultures of the different strains. The quantitative PCR amplification was performed with the CFX96 real-time PCR detection system (Bio-Rad, Richmond, CA, USA) using the HOT FIREPol EvaGreen quantitative PCR Mix Plus (Solis BioDyne, Tartu, Estonia) and the protocol recommended by the manufacturer. Copy numbers of genes (corrected by copy numbers per cell) were used to determine frequencies of strains at the beginning and the end of the experiment. Changes in relative frequencies between the start and the end of the experiments was used to calculate relative fitness (Wr) according to the equation: Wr =(at(1−a0)/a0(1−at)), where a0 and at are the cell densities of the focal strain a at the beginning and the end, respectively (Ross-Gillespie et al., 2007).
Cell numbers and biomass determination
Bacterial cell densities were determined by flow cytometry. Subsamples of 1 ml were fixed with glutaraldehyde (v/v 2.5%) and stained for 20 min with DAPI (4',6-diamidino-2-phenylindole, 1 μg ml−1). The cell numbers were measured with an Influx V-Gs cell sorter (Becton Dickinson Inc., San Jose, CA, USA). DAPI fluorescence was excited with a UV laser (60 mW, 355 nm) and a blue laser (200 mW, 488 nm) was used for side scatter light and the autofluorescence of Poterioochromonas sp. DS. Bacterial cells were detected side scatter light vs DAPI fluorescence (431 nm) and flagellates were determined using side scatter light vs green fluorescence (531 nm).
As it was not possible to quantify cells within the aggregated subpopulations, the comparison of growth yield between the ancestor and the evolved strains at ancestral growth conditions was based on biomass rather than cell counts. The total protein biomass of the ancestor strain and of bacteria isolated after 26 re-inoculation cycles from P+ and P− was determined in pre-grown cultures after 72 h on DSMZ 7 medium. The subsamples were concentrated by centrifugation, the pellet was re-suspended in inorganic ALW, and the cells were disrupted by 4 × 30 s of sonication (Skan, Sonifier 250, Basel-Allschwil, Switzerland). Protein fluorescence was measured on a Qubit 2.0 (Invitrogen, Carlsbad, CA, USA) after staining with Qubit protein reagent according to the manufacturer’s specification. The biomass of free cells was determined separately by pre-filtration of subsamples through 3 μm pore size filters (Millipore, Bedfird, MA, USA).
Statistical analysis
The data sets were first tested for normalcy by the Shapiro–Wilk tests. Non-normally distributed data sets were analysed using nonparametric tests. To test for significant changes in the final cell densities between the treatments during the evolution experiment, a one-way analysis of variance (ANOVA) followed by a Tukey post hoc test was applied (N=5 per treatment). To examine the differences in biomass and biomass distribution between the ancestor strain and the evolved isolates, a Kruskal–Wallis ANOVA (N=3 per treatment) followed by a Dunn–Bonferroni post hoc test was used. For statistical analysis of the other phenotypic assays, that is, growth efficiency, competition and benzoate assay, the mean values of the technical replicates were compared. The difference in growth yield before and after the excision of the genetic element was tested for significance by ANOVA (N=3 per treatment) followed by a Tukey post hoc test. Significance of the change in relative fitness between the P+ and the ancestor or the P− strain was analysed with a one-sample t-test (N=3). The influence of benzoate on growth of the evolved and ancestor strains was tested by unpaired t-test.
Results
Consequences of substrate limitation and predation
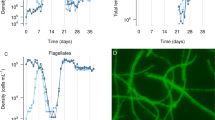
Cultures evolving without predation (P−) exhibited a gradual but moderate increase of final cell densities—as compared with the ancestor—over the course of the experiment (ANOVA; treatment without predator: F(2,12)=63.78, P<0.001,Tukey post hoc test for P-26 growth cycles vs ancestor, P<0.001; treatment with predator: F(2,12)=13.61, P=0.001, Tukey post hoc test for P+ 26 growth cycles vs ancestor P=0.001, Figure 1a). The cell yield of the P− cultures after 26 growth cycles was on average 1.7 times higher than at the beginning. The P+ cultures showed a more drastic adaptation, and their final cell densities were 15 times higher than their initial numbers. Strains isolated from the P+ treatment after 26 growth cycles formed slightly (but not significantly) higher total biomass than the P− and ancestor strains if grown in the rich DSMZ 7 medium that was used to maintain the ancestor (Figure 1b). In the P− strains, a significantly larger fraction of total biomass was present in free cells, in agreement with the notion of aggregate formation being a predation defence (Kruskal–Wallis test, χ2(2)=9.9, P=0.007, post hoc Dunn–Bonferroni test: P− vs ancestor, P=0.04; P− vs P+, P=0.021).
Increase of total growth yield during experimental evolution and reallocation of biomass between planktonic and aggregate associated cells. (a) Cell densities of the ancestor Sphingobium sp. Z007 strain (168 h of growth at oligotrophic conditions) and of three strains evolving without (P−) or with (P+) predators that were isolated after 16 and 26 growth cycles. (b) Biomass of planktonic cells (hatched) and aggregates (white) of the ancestor and of strains with different evolutionary background isolated after 26 growth cycles when grown on the original rich medium.
Genomic organization and characteristics of the ancestor
The genome of the Sphingobium sp. Z007 ancestor strain consists of two circular chromosomes and five plasmids with a final genome size of 4.9 Mbp. In all, 3276 from a total of 4998 coding sequences could be functionally annotated. A summary for each genetic element is listed in Table 1. The genome includes four rRNA operons and 60 tRNA genes that cover all amino acids. The closest relative of Sphingobium sp. Z007 is Sphingobium japonicum, with a 16 S rRNA gene identity of 98%. Chromosome 1 contains all genes essential to maintain a living cell (Gil et al., 2004). Chromosome 2 is only half the size of chromosome 1; it mainly harbours genes involved in substrate degradation, as well as nearly half of all transposons. We also found genes involved in quorum sensing, namely one autoinducer gene on plasmid 1 and three receptor genes on chromosomes 1, 2 and plasmid 1.
Low number of polymorphisms and plasmid overexpression
In general, all sequences of the evolved isolates showed low numbers of single-nucleotide polymorphisms (between 10 and 20 per isolate), and the majority of these variants were synonymous transitions with no effect on the amino acid composition. The non-synonymous substitutions were mainly found on chromosome 1 and plasmid 1. However, we could not assign a function to these substitutions because all except one site were annotated as hypothetical proteins. The annotated site with a non-synonymous substitution was found in a gene coding for a TonB-dependent transporter and present in all evolved isolates.
We examined the distribution of Illumina reads among all sequenced isolates to see whether there are treatment-dependent changes in copy numbers of genes or genomic elements. We found that the plasmid 1 had proportionally more reads in the strains evolved with predator then in the strains isolated from the P− treatment or the ancestor strain, which suggests that this plasmid might be present in a higher copy number in the P+ strains. Plasmid 1 harbours a quorum-sensing system with a receptor and autoinducer gene and also the complete operon for the type 4 secretion system.
Loss of a genome fragment
All three strains from different experimental replicates of the P+ treatment had a smaller genome size than both, the ancestor and the P− strains. In all three cases, the lost genomic fragment is located on chromosome 2 (Figure 2); it is nearly 274 kbp in size and contains 306 coding sequences, 164 of which could be sorted into different metabolic groups. The most abundant annotated coding sequences on the excision are mobile elements (insertion sequences from different families) and unique sets of genes involved in the metabolism of aromatic compounds. Consequently, the P+ strains have completely lost these degradation pathways. Mobile elements and large stretches of repeats are found on both flanking sites of the excision and might be responsible for its loss. In fact, the deleted 274 kbp fragment is flanked by direct repeats, which presumably have served as templates in the recombination event that led to the loss of the fragment. Screening of a collection of >50 isolates from different replicates and time points of the P+ treatment by the specific PCR assay (supplementary Figure 1A) revealed that the excision had occurred in the three isolates after 20, 22 and 24 re-inoculation cycles, respectively (corresponding to 136, 150 and 164 days). The same PCR assay also showed that the excision was not present in any of the 40 examined P− strains (eight isolates per biological replicate), whereas all 40 tested isolates evolved with the predator appeared to have lost the genetic element (Supplementary Figures 1B and C).
Growth efficiency at oligotrophic conditions after genome streamlining
The growth on oligotrophic medium of P+ strains from before (16 growth cycles) and after (26 growth cycles) the excision revealed beneficial effects of genome streamlining. Significantly higher growth yield of all strains with reduced genomes was observed, and this was equally pronounced in the absence (ANOVA, F(2,6)=54.07, P<0.001, Tukey post hoc test for reduced vs non-reduced genomes P=0.0003) and presence (ANOVA, F(2,6)=89.39, P<0.001, Tukey post hoc test for reduced vs non-reduced genomes P<0.0001) of predators (Figure 3). To verify that change in growth was mainly related to the loss of the genomic element and not to the prolonged cultivation in the medium, strains evolving for 16 growth cycles in the evolution experiment were compared with strains isolated directly before the genome reduction. No significant differences in growth between the two isolation time points was observed (Supplementary Figure 2).
Cell densities after 96 h of growth in nutrient-limited medium cultivated either without or with predator of three independently evolved P+ strains that were isolated in the middle of the experiment (16 growth cycles), directly before the excision (24, 20 and 22 growth cycles for strains A, B and C, respectively) and after genome reduction (26 growth cycles). Strains were cultivated either without or with predator.
Direct competition assays at ancestral growth conditions
To examine the respective competitiveness of the evolved strains and the competition between P+ strains and the ancestor at the substrate-rich ancestral growth conditions, we measured their fitness in co-culture as the change in relative frequency of marker genes. Prior tests confirmed that comparable abundance estimates of the ancestor strain were obtained using primers either for 16 S rRNA genes or for the excision region (data not shown). The quantitative PCR efficiencies, as determined from standard curves, were always between 98% and 102%. Significantly higher fitness of the P+ than the P− strains was observed, both, in the absence (one-sample t-test, t(8)=4.03, P=0.004) and the presence of predators (one-sample t-test, t(8)=3.14, P=0.014, Figure 4a). The P+ strain had a significant fitness advantage in the presence of the predator (one-sample t-test: treatment with predator, t(2)=4.98, P=0.038, Figure 4b), and a trend towards higher competitiveness (albeit not statistically significant) was also apparent in the absence of the predator.
Competitive performance of strains evolved with predators. (a) Differences in fitness between strains from the P+ and P− treatments (26 growth cycles) cultivated either without or with predators. (b) Differences in fitness between strains from the P+ and the ancestor (Anc) without or with predators.
Growth in the presence of benzoate
One consequence of the genome reduction in P+ strains was the loss of genes involved in the metabolism of aromatic compounds, including the complete pathway for benzoate degradation (Figure 3). The ancestor and the P− strains, both with an intact gene cluster for this metabolic function, grew to higher densities in media containing benzoate than in control treatments (unpaired t-test: Ancestor, t(4)=−4.8, P=0.008; evolved without predator, t(4)=−2.96, P=0.042, Figure 5), indicating that they utilized this compound as an additional substrate source. By contrast, benzoate negatively affected the growth of the P+ strains (unpaired t-test, t(4)=3.66, P=0.022, Figure 5).
Discussion
Sphingomonadaceae are distributed across a range of non-aquatic and aquatic habitats. Their genome evolution is characterized by a high extent of horizontal gene transfer, recombination and genome rearrangements (Aylward et al., 2013). Genome reduction within this phylogenetic lineage appears to reflect ecological transitions to less substrate-rich environments (Lauro et al., 2009), in agreement with a general trend of smaller genomes in bacteria from aquatic than from terrestrial habitats (Dini-Andreote et al., 2012).
Although the smallest, most eroded genomes are found in obligate endosymbionts (Andersson et al., 1998, Stephens et al., 1998), the hallmark examples of extreme genome reduction in free-living bacteria are from oligotrophic marine and freshwater environments (Dufresne et al., 2003; Giovannoni et al., 2005; Garcia et al., 2013; Ghylin et al., 2014). Genome streamlining of waterborne bacteria is considered to be a consequence of positive selection rather than of genetic drift (Giovannoni et al., 2014), to increase growth efficiency in a resource-limited environment. Both natural and experimentally introduced gene loss resulted in increased fitness of Salmonella enterica (Koskiniemi et al., 2012). Genome reduction in our strain was exclusively triggered under predation (supplementary Figure 2C), and only after several 100 generations. In the absence of predators, growth efficiency could be increased by re-tuning the resource allocation trade-off between growth and defence (Cressler et al., 2010), that is, by largely abandoning the aggregated phenotype (Figure 1b). This shift to the free-living phenotype at oligotrophic conditions, moreover, speaks for a growth disadvantage of the aggregated growth form, which at times may even be advantageous due to increased advective transport during sinking (Kiørboe et al., 2001). In any case, this evolutionary path was no option for the P+ strains, which maintained or even increased aggregate formation, likely due to the high mortality inflicted on the planktonic subpopulation. A preferential adaptation of bacteria to defence rather than to abiotic factors has been observed before in viral–bacterial co-cultures (Scanlan et al., 2015).
During the 1-week growth cycles, flagellates removed approximately 80% of newly formed free bacterial cells already after 24 h and subsequently reduced them to <25% of their maximal abundances. Consequently, the starter populations of Sphingobium sp. Z007 at each re-inoculation in P+ were two to three orders of magnitude smaller than in P−. Population size is a crucial factor to influence the total genetic diversity and fixation of mutations may be more likely if strong selection acts on a small population (Raynes et al., 2012). Such a growth scenario might also favour the success of a singular, more radical adaptation, as represented by the genomic excision. Moreover, 10 to 100 times more cells in the freshly re-inoculated P+ variant originated from aggregates rather than from planktonic bacteria, so that this phenotype constituted by far the largest effective subpopulation. It is, thus, conceivable that the excision event occurred in the aggregated phenotype, and was rapidly established by conveying a growth advantage to cells that were not subjected to predation mortality (Figure 3). This might also explain why it was never observed in the P− strains. Alternatively, the excised genome fragment might have harboured elements that were beneficial for planktonic growth. We previously observed that the growth performance of Sphingobium spp. strains in rich medium substantially improved after their transient residence at oligotrophic conditions (Baumgartner et al., 2016). Here we expand these results by showing that there may be additional beneficial effects in populations that simultaneously face both, oligotrophy and high mortality, which is a common fate of soilborne or plant associated bacteria transported into freshwaters (Ruiz-González et al., 2015).
Metabolic costs associated with larger genome size arise from regulation overhead and to a lesser extent from actual synthesis (Ranea et al., 2005). The superior growth of P+ strains after the excision on oligotrophic media irrespective of predation (Figure 3) suggests that the overall fitness benefit of the genome reduction for the P+ strains may have exceeded the 5% of saved nucleotides. Interestingly, it also bestowed an advantage on P+ strains at the original substrate-rich growth conditions, as they had a higher fitness than both, the ancestor and the P− strains under predation (Figure 4) and at least matched the fitness of the ancestor without predation. Streamlining might thus be interpreted as a compensatory adaptation to attenuate the metabolic cost of the formation of the defence trait (aggregation), which already developed during the initial growth cycles. Such compensatory evolution in trade-off situations or due to disproportional accumulation of deleterious mutations are common in pro- and eukaryotes (Maisnier-Patin and Andersson, 2004). In fact, the P+ strains after the excision event might be regarded as the first examples for an experimentally induced ‘Winnie-the-Pooh’ type of adaptation (Thingstad et al., 2005), that is, the simultaneous increase of growth and defence-related fitness aspects. However, the involved mechanism is unrelated to the originally proposed one.
Despite the drastic reduction of genome size in the strains evolved with predator, other genomic alterations in the evolved strains were rare. Single-nucleotide polymorphism calling revealed only a very small number of InDels, the majority of which did not affect the amino acid composition of the respective protein. In our experiments, all strains evolved towards higher growth yield at 1000-fold reduced substrate levels (Figure 1a). This adaptation to substrate restriction irrespective of predation might have been related to a non-synonymous substitution in a TonB-dependent transporter observed in all evolved isolates. These high-affinity transporters are specialized for siderophores or dissolved organic matter, and they are often found in bacterial strains that are adapted to oligotrophic environments (Tang et al., 2012). The Sphingomonadales show a particular high diversity of TonB transporters; our strain shares high similarities in some mobile elements with Sphingomonas wittichii, which features the highest number of these transporter of all so-far sequenced microorganisms (Miller et al., 2010). The high numbers of TonB transporters in the Sphingomonadacea hint at their importance for the interaction with their environment, and it is conceivable that the evolved strains improved in performance by mutation in a transporter involved in the uptake of scarce nutrients. However, the testing of this hypothesis was beyond the scope of the current study.
The excision clearly further mitigated the counteracting selection pressures of nutrient limitations and high mortality (Figures 1 and 3). However, it did not appear to be involved in the enhanced formation of aggregates, as the number of aggregates did not change substantially before and after the excision (Supplementary Figure 2). Instead, the analysis of sequence-read distributions indicated a higher copy number of plasmid 1 in strains evolved with predators. A positive effect of elevated plasmid copy number has been shown in bacteria evolving under antibiotic selection (San Millan et al., 2016): The increased gene dosage at higher copy numbers resulted in elevated resistance to the applied antibiotics. In analogy with these findings, a higher expression of plasmid 1, which harbours a complete quorum sensing and type IV secretion system, might have a positive effect on aggregate formation. Quorum sensing is important for collaborative behaviour in bacteria, such as the formation of biofilms, and quorum-sensing knockout mutants of Burkholderia thailandensis were not able to form cell aggregates (Chandler et al., 2009). In addition, the type IV secretion system, which is regulated by the quorum-sensing system (Delrue et al., 2005), has also been shown to be involved in the formation of biofilms and cell aggregates (Bieber et al., 1998). Therefore, it is feasible that enhancing quorum sensing and the resulting increased formation of cell–cell connections by the type IV secretion system might account for the elevated aggregate formation in the P+ strains.
Interestingly, the three independently evolved isolates from the P+ treatment lost the exact same genomic region. Moreover, additional PCR assays suggested that all 40 strains randomly isolated at the end of the experiment from the five biological replicates of P+ jettisoned this genomic region; this strongly suggests that the excision had swept through the whole population. The excised genome segment most likely represents an eroded remnant of an integrated plasmid, as deduced from plasmid-related genes and the disproportionally high number of transposable elements. The fact that this segment is flanked by repetitive elements that probably served as recombination templates suggests that a re-integration of that genomic segment might also be possible if environmental conditions change and the corresponding plasmid is available. Indeed, this segment harboured a fully functional pathway for the degradation of aromatic compounds (Figure 2), as demonstrated by enhanced cell yield of the ancestor and P− strains in the presence of benzoate (Figure 5). By contrast, this compound, which is known for its antimicrobial activity but is also common in terrestrial habitats such as the rhizosphere (Whitehead et al., 1981), constituted a growth disadvantage for the P+ strains. Thus, the price of genome reduction in Sphingobium sp. strain Z007 seems to be a constriction of its potential niche space. It is conceivable that similar eco-evolutionary mechanisms might lead to the loss of other functions that are acquired by lateral gene transfer, for example, antibiotic resistance.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ . (1990). Basic local alignment search tool. J Mol Biol 215: 403–410.
Andersson SGE, Zomorodipour A, Andersson JO, Sicheritz-Ponten T, Alsmark UCM, Podowski RM et al. (1998). The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396: 133–140.
Aylward FO, McDonald BR, Adams SM, Valenzuela A, Schmidt RA, Goodwin LA et al. (2013). Comparison of 26 sphingomonad genomes reveals diverse environmental adaptations and biodegradative capabilities. Appl Environ Microbiol 79: 3724–3733.
Aziz R, Bartels D, Best A, DeJongh M, Disz T, Edwards R et al. (2008). The RAST server: rapid annotations using subsystems technology. BMC Genomics 9: 75.
Baas-Becking LGM . (1934) Geobiologie; of inleiding tot de milieukunde. WP Van Stockum & Zoon NV: The Hague, The Netherlands.
Baumgartner M, Neu TR, Blom JF, Pernthaler J . (2016). Protistan predation interferes with bacterial long-term adaptation to substrate restriction by selecting for defence morphotypes. J Evol Biol 29: 2297–2310.
Bieber D, Ramer SW, Wu C-Y, Murray WJ, Tobe T, Fernandez R et al. (1998). Type IV pili, transient bacterial aggregates, and virulence of enteropathogenic Escherichia coli. Science 280: 2114–2118.
Blom JF, Hornak K, Simek K, Pernthaler J . (2010a). Aggregate formation in a freshwater bacterial strain induced by growth state and conspecific chemical cues. Environ Microbiol 12: 2486–2495.
Blom JF, Zimmermann YS, Ammann T, Pernthaler J . (2010b). Scent of danger: floc formation by a freshwater bacterium is induced by supernatants from a predator-prey coculture. Appl Environ Microbiol 76: 6156–6163.
Chandler JR, Duerkop BA, Hinz A, West TE, Herman JP, Churchill ME et al. (2009). Mutational analysis of Burkholderia thailandensis quorum sensing and self-aggregation. J Bacteriol 191: 5901–5909.
Corno G, Jürgens K . (2008). Structural and functional patterns of bacterial communities in response to protist predation along an experimental productivity gradient. Environ Microbiol 10: 2857–2871.
Cressler C, King A, Werner E . (2010). Interactions between behavioral and life‐history trade‐offs in the evolution of integrated predator‐defense plasticity. Am Nat 176: 276–288.
Delrue R-M, Deschamps C, Léonard S, Nijskens C, Danese I, Schaus J-M et al. (2005). A quorum-sensing regulator controls expression of both the type IV secretion system and the flagellar apparatus of Brucella melitensis. Cell Microbiol 7: 1151–1161.
Dini-Andreote F, Andreote F, Araújo W, Trevors J, van Elsas J . (2012). Bacterial genomes: habitat specificity and uncharted organisms. Microb Ecol 64: 1–7.
Dufresne A, Salanoubat M, Partensky F, Artiguenave F, Axmann IM, Barbe V et al. (2003). Genome sequence of the cyanobacterium Prochlorococcus marinus SS120, a nearly minimal oxyphototrophic genome. Proc Natl Acad Sci USA 100: 10020–10025.
Garcia SL, McMahon KD, Martinez-Garcia M, Srivastava A, Sczyrba A, Stepanauskas R et al. (2013). Metabolic potential of a single cell belonging to one of the most abundant lineages in freshwater bacterioplankton. ISME J 7: 137–147.
Ghylin TW, Garcia SL, Moya F, Oyserman BO, Schwientek P, Forest KT et al. (2014). Comparative single-cell genomics reveals potential ecological niches for the freshwater acl Actinobacteria lineage. ISME J 8: 2503–2516.
Gil R, Silva FJ, Peretó J, Moya A . (2004). Determination of the core of a minimal bacterial gene set. Microbiol Mol Biol Rev 68: 518–537.
Giovannoni SJ, Tripp HJ, Givan S, Podar M, Vergin KL, Baptista D et al. (2005). Genome streamlining in a cosmopolitan oceanic bacterium. Science 309: 1242–1245.
Giovannoni SJ, Cameron Thrash J, Temperton B . (2014). Implications of streamlining theory for microbial ecology. ISME J 8: 1553–1565.
Guildford SJ, Hecky RE . (2000). Total nitrogen, total phosphorus, and nutrient limitation in lakes and oceans: Is there a common relationship? Limnol Oceanogr 45: 1213–1223.
Hahn MW, Höfle MG . (1999). Flagellate predation on a bacterial model community: interplay of size-selective grazing, specific bacterial cell size, and bacterial community composition. Appl Environ Microbiol 65: 4863–4872.
Hahn MW, Stadler P, Wu QL, Pockl M . (2004). The filtration-acclimatization method for isolation of an important fraction of the not readily cultivable bacteria. J Microbiol Methods 57: 379–390.
Kasalicky V, Jezbera J, Simek K, Hahn MW . (2010). Limnohabitans planktonicus sp nov and Limnohabitans parvus sp nov., planktonic betaproteobacteria isolated from a freshwater reservoir, and emended description of the genus Limnohabitans. Int J Syst Evol Microbiol 60: 2710–2714.
Kiørboe T, Ploug H, Thygesen UH . (2001). Fluid motion and solute distribution around sinking aggregates I: Small-scale fluxes and heterogeneity of nutrients in the pelagic environment. Mar Ecol Prog Ser 211: 1–13.
Koskiniemi S, Sun S, Berg OG, Andersson DI . (2012). Selection-driven gene loss in bacteria. PLoS Genet 8: e1002787.
Lauro FM, McDougald D, Thomas T, Williams TJ, Egan S, Rice S et al. (2009). The genomic basis of trophic strategy in marine bacteria. Proc Natl Acad Sci USA 106: 15527–15533.
Lenski RE, Levin BR . (1985). Constraints on the coevolution of bacteria and virulent phage: a model, some experiments, and predictions for natural communities. Am Nat 125: 585–602.
Maisnier-Patin S, Andersson DI . (2004). Adaptation to the deleterious effects of antimicrobial drug resistance mutations by compensatory evolution. Res Microbiol 155: 360–369.
Miller TR, Delcher AL, Salzberg SL, Saunders E, Detter JC, Halden RU . (2010). Genome sequence of the dioxin-mineralizing bacterium Sphingomonas wittichii RW1. J Bacteriol 192: 6101–6102.
Mitchell JG, Kogure K . (2006). Bacterial motility: links to the environment and a driving force for microbial physics. FEMS Microbiol Ecol 55: 3–16.
Overbeek R, Olson R, Pusch GD, Olsen GJ, Davis JJ, Disz T et al. (2014). The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res 42: D206–D214.
Pernthaler J . (2005). Predation on prokaryotes in the water column and its ecological implications. Nat Rev Microbiol 3: 537–546.
Ranea JAG, Grant A, Thornton JM, Orengo CA . (2005). Microeconomic principles explain an optimal genome size in bacteria. Trends Genet 21: 21–25.
Raynes Y, Gazzara MR, Sniegowski PD . (2012). Contrasting dynamics of a mutator allele in asexual populations of differing size. Evolution 66: 2329–2334.
Ross-Gillespie A, Gardner A, West SA, Griffin AS . (2007). Frequency dependence and cooperation: theory and a test with bacteria. Am Nat 170: 331–342.
Ruiz-González C, Niño-García JP, del Giorgio PA . (2015). Terrestrial origin of bacterial communities in complex boreal freshwater networks. Ecol Lett 18: 1198–1206.
Rutherford K, Parkhill J, Crook J, Horsnell T, Rice P, Rajandream M-A et al. (2000). Artemis: sequence visualization and annotation. Bioinformatics 16: 944–945.
San Millan A, Escudero JA, Gifford DR, Mazel D, MacLean RC . (2016). Multicopy plasmids potentiate the evolution of antibiotic resistance in bacteria. Nat Ecol Evol 1: 0010.
Scanlan PD, Hall AR, Blackshields G, Friman VP, Davis MR, Goldberg JB et al. (2015). Coevolution with bacteriophages drives genome-wide host evolution and constrains the acquisition of abiotic-beneficial mutations. Mol Biol Evol 32: 1425–1435.
Stephens RS, Kalman S, Lammel C, Fan J, Marathe R, Aravind L et al. (1998). Genome sequence of an obligate intracellular pathogen of humans: Chlamydia trachomatis. Science 282: 754–759.
Tang K, Jiao N, Liu K, Zhang Y, Li S . (2012). Distribution and functions of TonB-dependent transporters in marine bacteria and environments: implications for dissolved organic matter utilization. PLoS One 7: e41204.
Thingstad TF, Øvreås L, Egge JK, Løvdal T, Heldal M . (2005). Use of non-limiting substrates to increase size; a generic strategy to simultaneously optimize uptake and minimize predation in pelagic osmotrophs? Ecol Lett 8: 675–682.
Whitehead DC, Dibb H, Hartley RD . (1981). Extractant ph and the release of phenolic compounds from soils, plant roots and leaf litter. Soil Biol Biochem 13: 343–348.
Zotina T, Köster O, Jüttner F . (2003). Photoheterotrophy and light-dependent uptake of organic and organic nitrogenous compounds by Planktothrix rubescens under low irradiance. Freshw Biol 48: 1859–1872.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Baumgartner, M., Roffler, S., Wicker, T. et al. Letting go: bacterial genome reduction solves the dilemma of adapting to predation mortality in a substrate-restricted environment. ISME J 11, 2258–2266 (2017). https://doi.org/10.1038/ismej.2017.87
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.87