Abstract
Wood-inhabiting fungi have essential roles in the regulation of carbon stocks and nutrient cycling in forest ecosystems. However, knowledge pertaining to wood-inhabiting fungi is only fragmentary and controversial. Here we established a large-scale deadwood experiment with 11 tree species to investigate diversity and tree species preferences of wood-inhabiting fungi using next-generation sequencing. Our results contradict existing knowledge based on sporocarp surveys and challenge current views on their distribution and diversity in temperate forests. Analyzing α-, β- and γ-diversity, we show that diverse fungi colonize deadwood at different spatial scales. Specifically, coniferous species have higher α- and γ-diversity than the majority of analyzed broadleaf species, but two broadleaf species showed the highest β-diversity. Surprisingly, we found nonrandom co-occurrence (P<0.001) and strong tree species preferences of wood-inhabiting fungi, especially in broadleaf trees (P<0.01). Our results indicate that the saprotrophic fungal community is more specific to tree species than previously thought.
Similar content being viewed by others
Main
Wood-inhabiting fungi have an essential role in the decomposition of deadwood, an important carbon stock in global forest ecosystems (Pan et al., 2011; Rajala et al., 2012). Deadwood is a complex and poor quality (high C: N ratio) substrate composed of a heterogeneous assemblage of simple molecules combined with several types of complex biopolymers, creating a nutrient resource that is difficult to access and decompose for most organisms (Pan et al., 2011; Hoppe et al., 2016). Wood-inhabiting fungi secrete oxidoreductases and hydrolases (wood-decomposition enzymes) that mineralize or decompose most plant cell wall polymers into simple compounds that are accessible to other organisms (Floudas et al., 2012; Purahong et al., 2016a).
Diversity and distribution patterns of microbial community and their drivers are central issues in microbial ecology as this information is crucial for understanding and predicting the role played by microbes in maintaining ecosystem functions and stability (Kubartová et al., 2012) and can help when making decisions about land management. Recently, the conservation of microorganisms has become an issue of concern, especially for wood-inhabiting fungi (Seibold et al., 2015). However, our knowledge about wood-inhabiting fungi is only fragmentary and contested due to limitations in detection methods for fungal communities and a lack of well-designed field experiments with sufficient replicates (Kubartová et al., 2012; Seibold et al., 2015; Hoppe et al., 2016). Even the most fundamental questions about diversity and tree species preference of wood-inhabiting fungi have never been tackled using suitable approaches and experiments (Seibold et al., 2015). Based on existing knowledge pertaining to wood-inhabiting fungal ecology based on sporocarp surveys, wood-inhabiting fungal communities in temperate forests are thought to exhibit low α-diversity (average ∼2 species or less/deadwood log) (Blaser et al., 2013) and not be specific to tree species, leading to researchers differentiating only between softwood and hardwood degraders (Tuor et al., 1995), and these views have been confirmed recently (Baber et al., 2016). The lack of tree species preference of wood-inhabiting fungi in temperate forest has also not been questioned as it fits with the widely accepted view that saprotrophic fungi have weaker relationships to specific tree species than do symbiotic or parasitic fungi (Peay et al., 2013). However, there are few studies that showed some degrees of the selectivity of heart-rot fungi for trees species (Rayner and Boddy, 1988; Boddy, 2001; Boddy et al., 2017).
This study aimed to address two specific questions. (i) Are there differences in the diversity of wood-inhabiting fungi that colonize deadwood of different tree species? (ii) Do wood-inhabiting fungal taxa exhibit tree species preference? Therefore, we sampled deadwood, not fruiting bodies, and used pyrotag sequencing of the fungal internal transcribed spacer rRNA genes to investigate the diversity, composition and distribution patterns of wood-inhabiting fungal communities in the early phase of decomposition (3 years) in 11 tree species (7 broadleaf: birch (Betula pendula Roth, Betulaceae), hornbeam (Carpinus betulus L., Betulaceae), beech (Fagus sylvatica L., Fagaceae), ash (Fraxinus excelsior L., Oleaceae), aspen (Populus spp., Salicaceae), oak (Quercus spp., Fagaceae), and lime tree (Tilia spp., Malvaceae), and 4 coniferous species: larch, (Larix decidua Mill., Pinaceae), Norway spruce (Picea abies L., H.Karst., Pinaceae), pine (Pinus sylvestris L., Pinaceae), and Douglas fir (Pseudotsuga menziesii (Mirb.), Franco, Pinaceae) (Figure 1 and Supplementary Information) distributed across three geographical locations in Germany. To our knowledge, this study is among the largest experiment on deadwood investigating tree species preference using a molecular approach (11 tree species × 27 replicates (1 ha forest plot each)=297 deadwood logs). All details on set-up, location, sampling methodology and pyrotag sequencing are described elsewhere and in Supplementary Information (Baber et al., 2016; Purahong et al., 2016a, b). Sampling design, laboratory procedures, bioinformatics and statistical analysis are described in Supplementary Information. Briefly, the freshly cut logs from each tree species (∼4 m long and mean diameter of 31±5.9 cm (s.d.)) were randomly put in each forest plot beside each other with a distance of 1 m between logs in 2009 and allowed to decompose for 3 years before sampling (Kahl et al., 2015). For bioinformatics, we filtered for good-quality sequences and processed as described in Purahong et al. (2016b). The quality filtered reads were shortened to their first 300 bases and normalized to the smallest read number per sample (3011 reads). Potential chimeras were removed using UCHIME 4.2.40 (Edgar et al., 2011) as implemented in MOTHUR. Rare operational taxonomic units (OTUs; singletons to quadrupletons) could potentially have originated from sequencing errors (Kunin et al., 2010) and were therefore removed from the data set. The raw sequence data sets are available in the European Nucleotide Archive under the study number PRJEB21052 (http://www.ebi.ac.uk/ena/data/view/PRJEB21052). α-Diversity of wood-inhabiting fungi across different tree species and wood-inhabiting fungal tree species preference data sets were tested using Kruskal–Wallis test combined with Mann–Whitney U-test and analysis of similarity (ANOSIM) based on the presence–absence data and Jaccard distance measure.
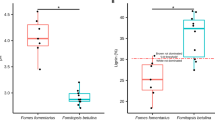
Tree species preference of wood-inhabiting fungal OTUs and diversity (α-, β-and γ-diversity) detected for different tree species. Tree species preference is indicated by the R statistics from analysis of similarity (ANOSIM) based on the presence–absence data and Jaccard distance measure (R=0–0.24, no separation to barely separated (green); R⩾0.25–0.75, separation with different degrees of overlap (yellow); R>0.75–1, well separated to complete separation (red); significant P-values (P<0.05) are given in bold and are based on 9999 permutations and Bonferroni corrections in all cases). Different letters indicate significant differences (P<0.05) according to Kruskal–Wallis test combined with Mann–Whitney U-test of average R statistics among different wood type combinations (broadleaf and broadleaf=blue, n=21; broadleaf and coniferous=gray, n=28; coniferous and coniferous=black, n=6) and average fungal richness (Ascomycota (Asco) richness per sample (mean±s.e., n=27); Basidiomycota (Basi) richness per sample (mean±s.e., n=27); total richness per sample (mean±s.e., n=27)). Other fungi=Zygomycota, Chytridiomycota and unidentified fungi.
Tree species preference of wood-inhabiting fungi can be explained by different ecological proxies (including tree community composition and the surrounding environmental conditions) and traits of the deadwood itself (Ferrer and Gilbert, 2003). Theoretically, when tree species diversity increases, opportunities for specialized wood-inhabiting fungi decrease as the probability of successful colonization drops when each specific tree species becomes rare (May, 1991). Therefore, in typical European temperate forests, which are characterized by a few abundant dominant tree species and some individuals of rare species, we expected wood-inhabiting fungi to exhibit tree species preferences for the dominant rather than the rare tree species. We determined dominant trees based on a percentage cover >10%, resulting in only four dominant species Picea abies, Pinus sylvestris, Fagus sylvatica and Quercus spp. (BMEL, 2014). The other seven species present (Betula pendula, Carpinus betulus, Fraxinus excelsior, Populus spp., Tilia spp., Larix decidua, Pseudotsuga menziesii were considered to be rare.
We detected an average of 22–42 wood-inhabiting fungal OTUs per log in the 11 tree species investigated, amounting to a total of 1254 OTUs, of which 677, 539 and 38 OTUs belonged to Ascomycota, Basidiomycota or other fungal groups (that is, Chytridiomycota, Zygomycota and unidentified fungi), respectively. Diversity and distributions of wood-inhabiting fungal OTUs in association with broadleaf and coniferous trees are shown in Figures 1 and 2 and described in detail in Supplementary Information (Supplementary Table S1). A recent study identified only 97 species based on sporocarp surveys in the same experimental plots (Baber et al., 2016), but we detected ∼12 times more wood-inhabiting fungal OTUs by sequencing DNA extracts from the logs (although in our study we had to exclude two broadleaf species not occurring in every plot but considered by Baber et al., 2016). The majority of wood-inhabiting fungi identified in the sporocarp study were also found in our molecular study (>70%), but many abundant wood-inhabiting fungal OTUs (that is, Amylostereum sp., Resinicium sp., Dacrymyces sp., Sistotrema sp., Phlebiopsis sp., and so on) were absent from the sporocarp survey. This discrepancy between the two approaches reflects the fact that the actively reproducing wood-inhabiting fungal community that can be seen in sporocarp surveys only poorly represents the whole wood-inhabiting fungal community and that a large portion of wood-inhabiting fungi reside in deadwood as vegetative mycelia or spores (Kubartová et al., 2012; Hoppe et al., 2016). Interestingly, we found a higher number of Ascomycota than Basidiomycota OTUs in all 11 tree species. Although Ascomycota are generally relatively poor at producing enzymes for deadwood decomposition, they may regulate wood decomposition rate by interacting and competing with Basidiomycota at least in the early stage of decomposition (van der Wal et al., 2014; Hoppe et al., 2016).
Specific and shared fungal OTUs detected in 11 tree species (a), 7 broadleaf tree species (b) and 4 coniferous tree species (c). The number next to each bar indicates the number of all detected fungal OTUs (Ascomycota, Basidiomycota, Zygomycota, Chytridiomycota and unidentified fungi). The overall architecture of tree species–fungal associations (d) illustrates how fungal OTUs that show preferences for particular tree species (detected in no more than two tree species) were distributed within a web of wood-inhabiting fungi. Different node sizes and colors represent different organismic and taxonomic groups: large nodes=plants (green=broadleaf tree and orange=conifer tree) and small nodes=fungi (red=Basidiomycota, navy blue=Ascomycota, sky blue=Zygomycota, purple=unidentified fungi).
We found differences in the wood-inhabiting fungi of different tree species at the levels of α-, β- and γ-diversity. In general, the coniferous species displayed higher α- and γ-diversities than the majority of the broadleaf species (Figure 1). However, among the broadleaf species, Quercus spp., Fraxinus excelsior and Populus spp. also had high α- and γ-diversities, similar to those of the conifers. In contrast, two broadleaf species (Fagus sylvatica and Carpinus betulus) with low α- and γ-diversities exhibited the highest β-diversity levels (regional-to-local diversity ratio). All these patterns remained when Ascomycota and Basidiomycota were considered separately (Figure 1).
It is noteworthy that we found a nonrandom co-occurrence pattern (C-score=46.028, P<0.001) and strong tree species preferences, especially in broadleaf species (Figures 1 and 2). Wood-inhabiting fungi colonizing coniferous wood showed less pronounced tree species preferences (Figures 1 and 2). Instead of detecting two separate clusters of wood-inhabiting fungal communities in coniferous vs broadleaf trees, we detected nine wood-inhabiting fungal communities (R=0.29–0.83, P<0.01), seven on the broadleaf species and two on the four conifers (Figure 1). In particular, although Quercus spp. and Fraxinus excelsior are broadleaf species with similar wood structure (ring porous) and C: N ratio (low), we found that the wood-inhabiting fungal communities associated with these two tree species were among the most different in all pairwise comparisons across all tree species (R=0.80, P<0.01) (Figures 1 and 2d). Of the conifers, Pinus sylvestris significantly separated from the other species (R=0.29–0.39, P<0.01) except Picea abies. Overall, we found only 16 generalist wood-inhabiting fungal OTUs (∼1%), while 331 (26%) were potential specialists (Figure 2). For broadleaf species, the proportion of potential specialists reached 41% (398 OTUs), whereas generalist wood-inhabiting fungal OTUs accounted for approximately 2% (18 OTUs). The proportion of the potential specialists in coniferous species was still high (35%, 288 OTUs) but there was high proportion of generalists as well (16%, 132 OTUs). The overall architecture of tree species–fungal associations illustrates how wood-inhabiting fungal OTUs that show preferences for particular tree species (that is, detected in a maximum of two tree species, Supplementary Table S2, Supplementary Information) were distributed within a web of wood-inhabiting fungi as shown in Figure 2d. We further discovered that the tree species preference of wood-inhabiting fungi was consistent in both Asco- and Basidiomycota and across the geographical locations (Figures 1 and 2 and Supplementary Information Figure S1, Supplementary Information). However, wood-inhabiting fungi of the Ascomycota (R=0.51, P<0.01) appeared more specific to tree species than those of the Basidiomycota (R=0.41, P<0.01) (Figure 1).
The high degree of tree species preference exhibited by wood-inhabiting fungi found in this study led us to ask ‘how do tree species influence the fungal saprotrophic community after their death?’ The results of the current study contrast sharply with the widely held belief that wood-inhabiting fungi in temperate forests are generalists and only separated into hardwood- and softwood-degrader communities (Tuor et al., 1995; Baber et al., 2016). Our results also further challenge the view that symbiotic fungi (that is, arbuscular mycorrhizal fungi or ectomycorrhizal fungi) have stronger and more specific relationships to their host plant (tree species preference) than saprotrophic fungi (Peay et al., 2013; Gao et al., 2013). On the contrary, we found very strong tree species preferences equal to or even stronger than those reported previously for symbiotic fungal communities. In our experiment, the logs from the 11 species were placed close to one another before being allowed to decompose for 3 years, which means that potential wood-inhabiting fungi from the surrounding environment had equal chance of reaching any of the logs and that cross colonization between logs was possible. Our finding of high specificity patterns despite this close arrangement of the logs led us to question the mechanisms behind this strong tree species preference of wood-inhabiting fungi. One might argue that, as we examined an early decomposition stage of the logs all of which originated from the same region, the detected wood-inhabiting fungi could correspond to endophytes and plant pathogens already present in the logs when they were cut. However, this is unlikely as some recent studies on the early stages of wood decay have shown that >70% of fungal endophytes in wood disappear after 140 days and the initial wood-inhabiting fungal community composition from freshly cut wood was completely different after 1 year of exposure (van der Wal et al., 2016; Song et al., 2017). Nevertheless, there are few studies indicating that latent endophytes and plant pathogens can survive in wood for longer time (Chapela and Boddy, 1988; Hiscox et al., 2015; Purahong et al., 2017), thus we checked all detected wood-inhabiting fungal OTUs. We found that, after 3 years of decomposition, <10% of the total wood-inhabiting fungal community could have originated from surviving initial endophytes and plant pathogens (see Supplementary Table S3, Supplementary Information). Removal of all these potential endophytes and plant pathogens from the data analyzed did not change our results relating to tree species preferences of wood-inhabiting fungi (Supplementary Figure S2, Supplementary Information).
The three important factors (traits of deadwood, the forest tree community composition and the surrounding environmental conditions) that are known to determine the tree species preference of wood-inhabiting fungi in tropical forests (Ferrer and Gilbert, 2003) could not explain our results completely. Strong tree species preferences for Fagus sylvatica and Quercus spp. can be explained based on the tree species abundances (May, 1991). However, for other rare broadleaf species, strong tree species preferences are unusual and difficult to understand. Wood-inhabiting fungi in temperate forests tend to be specific to tree species even when the trees are rare. If this is true, wood-inhabiting fungi should be prone to extinction as a result of monoculture forests. In addition, the wood traits described as shaping wood-inhabiting fungal communities (that is, wood types, wood structure, C: N ratio) failed to be predictors in our study, as we found that tree species with similar wood traits harbor different wood-inhabiting fungal communities (Seibold et al., 2015). In addition, our design of placing logs of all tree species close to each other but in random order across sites in three distinct regions enabled us to minimize the effects of surrounding environmental conditions in shaping wood-inhabiting fungal communities. We hypothesize that the tree species preferences of wood-inhabiting fungi may arise from (i) coevolution between tree species and wood-inhabiting fungi, as already envisaged for symbiotic fungi (Brundrett, 2002) and (ii) the intra- and inter-kingdom relationships among fungal, bacterial and invertebrate communities in deadwood (Müller et al., 2015; Hoppe et al., 2015; Song et al., 2017).
In conclusion, for the first time we investigated tree species preferences of wood-inhabiting fungi and quantified their diversity at different spatial scales (α-, β- and γ-diversity) using the next-generation sequencing approach. We were able to provide evidence of tree species preference exhibited by wood-inhabiting fungi in temperate forests in contrast to the widely accepted absence of tree species specificity. In deadwood, as well as other plant-derived substrates, the majority of microbes are unseen and much more diverse than those directly observable as fruiting bodies or revealed by isolation and culture techniques (Kubartová et al., 2012; Hoppe et al., 2016). High-resolution culture-independent molecular approaches (next-generation sequencing) should be applied to test and validate accepted knowledge and improve our understanding of the diversity and distribution patterns of microbial communities across wide ranges of habitats. Furthermore, such molecular approaches should be urgently incorporated and used to inform management and conservation strategies for microorganisms. Further studies on the wood traits, co-evolution of wood-inhabiting fungi and their tree species and the intra- and inter-kingdom relationships between fungi, bacteria and invertebrates in deadwood are needed for the mechanistic and functional understanding of tree species preferences of wood-inhabiting fungi in temperate forests.
References
Baber K, Otto P, Kahl T, Gossner MM, Wirth C, Gminder A et al. (2016). Disentangling the effects of forest-stand type and dead-wood origin of the early successional stage on the diversity of wood-inhabiting fungi. For Ecol Manag 377: 161–169.
Blaser S, Prati D, Senn-Irlet B, Fischer M . (2013). Effects of forest management on the diversity of deadwood-inhabiting fungi in Central European forests. For Ecol Manag 304: 42–48.
BMEL. (2014). The forests in Germany: selected results of the third national forest inventory. Federal Ministry of Food and Agriculture: Bonn, Germany.
Boddy L . (2001). Fungal community ecology and wood decomposition processes in angiosperms: from standing tree to complete decay of coarse woody debris. Ecol Bull, 43–56.
Boddy L, Hiscox J, Gilmartin EC, Johnston SR, Heilmann-Clausen J . (2017). Chapter 12: Wood decay communities in angiosperm wood. In: Mycology. The Fungal Community. CRC Press, pp 169–190.
Brundrett MC . (2002). Coevolution of roots and mycorrhizas of land plants. New Phytol 154: 275–304.
Chapela Ih, Boddy L . (1988). The fate of early fungal colonizers in beech branches decomposing on the forest floor. FEMS Microbiol Lett 53: 273–283.
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R . (2011). UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27: 2194–2200.
Ferrer A, Gilbert GS . (2003). Effect of tree host species on fungal community composition in a tropical rain forest in Panama: fungal community composition in Panama. Divers Distrib 9: 455–468.
Floudas D, Binder M, Riley R, Barry K, Blanchette RA, Henrissat B et al. (2012). The Paleozoic origin of enzymatic lignin decomposition reconstructed from 31 fungal genomes. Science 336: 1715–1719.
Gao C, Shi N-N, Liu Y-X, Peay KG, Zheng Y, Ding Q et al. (2013). Host plant genus-level diversity is the best predictor of ectomycorrhizal fungal diversity in a Chinese subtropical forest. Mol Ecol 22: 3403–3414.
Hiscox J, Savoury M, Müller CT, Lindahl BD, Rogers HJ, Boddy L . (2015). Priority effects during fungal community establishment in beech wood. ISME J 9: 2246–2260.
Hoppe B, Krüger D, Kahl T, Arnstadt T, Buscot F, Bauhus J et al. (2015). A pyrosequencing insight into sprawling bacterial diversity and community dynamics in decaying deadwood logs of Fagus sylvatica and Picea abies. Sci Rep 5: 9456.
Hoppe B, Purahong W, Wubet T, Kahl T, Bauhus J, Arnstadt T et al. (2016). Linking molecular deadwood-inhabiting fungal diversity and community dynamics to ecosystem functions and processes in Central European forests. Fungal Divers 77: 367–379.
Kahl T, Baber K, Otto P, Wirth C, Bauhus J . (2015). Drivers of CO2 emission rates from dead wood logs of 13 tree species in the initial decomposition phase. Forests 6: 2484–2504.
Kubartová A, Ottosson E, Dahlberg A, Stenlid J . (2012). Patterns of fungal communities among and within decaying logs, revealed by 454 sequencing. Mol Ecol 21: 4514–4532.
Kunin V, Engelbrektson A, Ochman H, Hugenholtz P. (2010). Wrinkles in the rare biosphere: pyrosequencing errors can lead to artificial inflation of diversity estimates. Environ Microbiol 12: 118–123.
May RM . (1991). A fondness for fungi. Nature 352: 475–476.
Müller J, Wende B, Strobl C, Eugster M, Gallenberger I, Floren A et al. (2015). Forest management and regional tree composition drive the host preference of saproxylic beetle communities. J Appl Ecol 52: 753–762.
Pan Y, Birdsey RA, Fang J, Houghton R, Kauppi PE, Kurz WA et al. (2011). A large and persistent carbon sink in the world’s forests. Science 333: 988–993.
Peay KG, Baraloto C, Fine PV . (2013). Strong coupling of plant and fungal community structure across western Amazonian rainforests. ISME J 7: 1852–1861.
Purahong W, Arnstadt T, Kahl T, Bauhus J, Kellner H, Hofrichter M et al. (2016a). Are correlations between deadwood fungal community structure, wood physico-chemical properties and lignin-modifying enzymes stable across different geographical regions? Fungal Ecol 22: 98–105.
Purahong W, Pietsch KA, Lentendu G, Schöps R, Bruelheide H, Wirth C et al. (2017). Characterization of unexplored deadwood mycobiome in highly diverse subtropical forests using culture-independent molecular technique. Front Microbiol 8: 574.
Purahong W, Wubet T, Lentendu G, Schloter M, Pecyna MJ, Kapturska D et al. (2016b). Life in leaf litter: novel insights into community dynamics of bacteria and fungi during litter decomposition. Mol Ecol 25: 4059–4074.
Rajala T, Peltoniemi M, Pennanen T, Mäkipää R . (2012). Fungal community dynamics in relation to substrate quality of decaying Norway spruce (Picea abies [L.] Karst.) logs in boreal forests. FEMS Microbiol Ecol 81: 494–505.
Rayner ADM, Boddy L . (1988) Fungal Decomposition of Wood: Its Biology and Ecology. Wiley.
Seibold S, Bässler C, Brandl R, Gossner MM, Thorn S, Ulyshen MD et al. (2015). Experimental studies of dead-wood biodiversity — a review identifying global gaps in knowledge. Biol Conserv 191: 139–149.
Song Z, Kennedy PG, Liew FJ, Schilling JS . (2017). Fungal endophytes as priority colonizers initiating wood decomposition. Funct Ecol 31: 407–418..
Tuor U, Winterhalter K, Fiechter A . (1995). Enzymes of white-rot fungi involved in lignin degradation and ecological determinants for wood decay. J Biotechnol 41: 1–17.
van der Wal A, Klein Gunnewiek PJA, Cornelissen JHC, Crowther TW, de Boer W. (2016). Patterns of natural fungal community assembly during initial decay of coniferous and broadleaf tree logs. Ecosphere 7: 1–17.
van der Wal A, Ottosson E, de Boer W . (2014). Neglected role of fungal community composition in explaining variation in wood decay rates. Ecology 96: 124–133.
Acknowledgements
The work has been (partly) funded by the DFG Priority Program 1374 ‘Infrastructure-Biodiversity-Exploratories’ (KR 3587/1-1, KR 3587/3-2, BU 941/17-1). We thank the managers of the three Exploratories—Kirsten Reichel-Jung, Swen Renner, Katrin Hartwich, Sonja Gockel, Kerstin Wiesner and Martin Gorke for their work in maintaining the plot and project infrastructure; Christiane Fischer and Simone Pfeiffer for giving support through the central office; Michael Owonibi for managing the central database; and Markus Fischer, Eduard Linsenmair, Dominik Hessenmöller, Jens Nieschulze, Daniel Prati, Ingo Schöning, Ernst-Detlef Schulze, Wolfgang W. Weisser and the late Elisabeth Kalko for their role in setting up the Biodiversity Exploratories project. The founders had no role in the study design, data collection and analysis, decision to publish or the preparation of the manuscript. We thank Peter Otto for his help during field sampling. We thank Katalee Jariyavidyanont for her help with DNA extraction and sequence library preparation. We thank Guillaume Lentendu for the bioinformatics support.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Purahong, W., Wubet, T., Krüger, D. et al. Molecular evidence strongly supports deadwood-inhabiting fungi exhibiting unexpected tree species preferences in temperate forests. ISME J 12, 289–295 (2018). https://doi.org/10.1038/ismej.2017.177
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.177
This article is cited by
-
Arbuscular mycorrhizal fungi and Streptomyces: brothers in arms to shape the structure and function of the hyphosphere microbiome in the early stage of interaction
Microbiome (2024)
-
Achieving structural heterogeneity and high multi-taxon biodiversity in managed forest ecosystems: a European review
Biodiversity and Conservation (2024)
-
Identifying environmental factors affecting the microbial community composition on outdoor structural timber
Applied Microbiology and Biotechnology (2024)
-
Long-term Dynamics of Fungal Communities Inhabiting Decaying Stumps of Quercus robur
Microbial Ecology (2024)
-
Applying molecular and genetic methods to trees and their fungal communities
Applied Microbiology and Biotechnology (2023)