Abstract
Marine alveolates (MALVs) are diverse and widespread early-branching dinoflagellates, but most knowledge of the group comes from a few cultured species that are generally not abundant in natural samples, or from diversity analyses of PCR-based environmental SSU rRNA gene sequences. To more broadly examine MALV genomes, we generated single cell genome sequences from seven individually isolated cells. Genes expected of heterotrophic eukaryotes were found, with interesting exceptions like presence of proteorhodopsin and vacuolar H+-pyrophosphatase. Phylogenetic analysis of concatenated SSU and LSU rRNA gene sequences provided strong support for the paraphyly of MALV lineages. Dinoflagellate viral nucleoproteins were found only in MALV groups that branched as sister to dinokaryotes. Our findings indicate that multiple independent origins of several characteristics early in dinoflagellate evolution, such as a parasitic life style, underlie the environmental diversity of MALVs, and suggest they have more varied trophic modes than previously thought.
Similar content being viewed by others
Introduction
The marine alveolate (MALV) lineages were discovered in the first large molecular surveys of SSU (18 S) rRNA from marine eukaryotes (López-García et al., 2001; Moon-van der Staay et al., 2001). Subsequent environmental SSU PCR-based surveys have consistently shown them to be diverse and widespread, often representing up to 50% of eukaryotic sequences (Chambouvet et al., 2008). SSU rRNA phylogeny has also shown that several ‘syndinian’ dinoflagellates branch with MALVs; these species have been investigated directly, and are all parasites infecting a broad range of host organisms such as cnidarians, fish, crustaceans, and protists (Guillou et al., 2010). Because of this, MALVs are generally assumed to be parasites, and by extension that their abundance in small size fractions may reflect large numbers of infectious cells (perhaps blooms following host blooms), which are thought to have ecological importance in controlling host populations. This is reinforced by recent surveys that show MALVs make up ~90% of putative parasites in the piconanoplankton globally (de Vargas et al., 2015). However, most MALVs are only known from these environmental SSU rRNA gene sequences, and those that have been characterized further are relatively rare and do not represent all MALV subgroups. As only SSU rRNA is generally known, the phylogenetic position of MALVs is also partially uncertain: they are typically basal to dinokaryotes (that is, dinoflagellates with permanently condensed chromosomes), but may be monophyletic or paraphyletic with dinokaryotes branching among them, and in either case the phylogenies lack support (Guillou et al., 2008; Skovgaard et al., 2009; Horiguchi, 2015). Overall, most MALV diversity is unexamined and how MALV groups are related to one another and to dinokaryotes remains poorly resolved.
Results and discussion
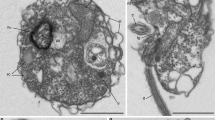
Using manual single cell isolation (or isolation of infected hosts) and fluorescence-activated cell sorting (FACS), we surveyed heterotrophic eukaryotes and identified cells corresponding to MALVs by SSU rRNA phylogeny, resulting in seven distantly related species from the Northeastern Subarctic Pacific Ocean (Line P) and from Monterey Bay out to the edge of the North Pacific Subtropical Gyre (Line 67; for collection and screening details, see Supplementary Table 1 and Supplementary Materials and methods). Two manual isolations were morphologically identifiable hosts (Figure 1); a copepod (L67-3) in the process of bursting to release spores identified as the dinoflagellate Chytriodinium, and a tintinnid ciliate (L67-6) that yielded two distinct MALV phylotypes (93.8% similarity, likely from two different MALV II species). Manually isolated cells L67-1 and L67-4 are inferred to be host-free and fit within size fractions that previously yielded MALV sequences: L67-1 is spherical-shaped with a diameter of 13.8 μm and with greenish inclusions, L67-4 is colourless, spindle-shaped, and 14.5 × 6.3 μm (Figure 1). One isolation (L67-5) was large and inferred to be a host, which could not be identified, and the remaining two cells (L67-2 and LP-1) were isolated by FACS.
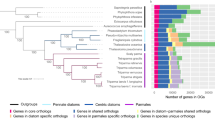
Morphology and phylogenetic relationships of MALVs. Tree topology is based on maximum likelihood analysis of concatenated SSU and LSU rRNA gene sequences (>3900 nucleotide positions; 7.1% gaps). Node support is shown by RAxML bootstraps (non-parametric) and MrBayes posterior probabilities. Black circles indicate a maximum support in both analyses. Sequences obtained in this study are printed in bold. Numbers in polygons refer to the number of grouped taxa, roman numerals signify the different MALV groups (compare with Guillou et al., 2008), and the arrows indicate parasitic clades. Removal/addition of the long-branching sequence (dotted line) did not change the tree topology. Micrographs show the organisms (including hosts) whose genomic DNA has been sequenced (not available for FACS-derived samples LP-1 and L67-2). L67-3 represents a disrupted copepod (identified by the extremities) with an attached cyst containing spores of the parasitic dinoflagellate Chytriodinium (Ch). The copepod was isolated after the cyst burst and released the spores, but it is not clear if the MALV is associated with the animal or the dinoflagellate. The micrograph of L67-6 shows a tintinnid ciliate. Since two non-related MALV II 18 S rRNA gene sequences (with 93.8% similarity; Supplementary Figure 2) and one 28 S rRNA gene sequence were obtained from three different contigs of this sample, its phylogenetic position is not shown in the SSU/LSU rRNA gene tree. L67-1 and L67-4 are assumed to represent host-free MALV cells. L67-4 is represented by two phylotypes with a SSU and a LSU sequence similarity of 97.9% and 98.6%, respectively. Length of the cleaned assembly, number of redundant/non-redundant KO homologs, number of hits to Tara Oceans V9 data (not available for L67-6, for which the V9 region was not sampled), and collection depth are presented beside each photo.
Genomic data from each isolation was generated, assembled, and screened for contamination (for example, from bacteria or host genomes) using a combination of BLAST-based and phylogenetic approaches (that is, phylogenetic trees were inferred from gene alignments of a broad range of taxa including the MALVs Hematodinium and Amoebophrya; for details see Supplementary Methods). Because dinoflagellate genomes are large and gene-sparse (Wisecaver and Hackett, 2011), we expected to recover partial genomes and focused only on regions encoding a recognizable gene, to avoid including non-coding regions from contaminants. This resulted in 0.5–3.1 Mb per cell confidently assigned to MALVs, encoding between 97 to 917 predicted genes, 30 to 60% of which corresponded to KEGG hits (Figure 1; Supplementary Tables 2 and 3; Supplementary Figure 1). Estimated coverage of eukaryotic conserved single copy genes using BUSCO showed 86 to 99% were missing. The genomic coverage was therefore low, as expected, but an improvement over only SSU rRNA.
The proteins that were identified matched the expectations for heterotrophic eukaryotes since no plastid-related protein was found, as observed in Hematodinium (Gornik et al., 2015), but mitochondrial proteins were detected. Interestingly, L67-5 encoded a photoactive proton pump, proteorhodopsin, most similar to homologues from Oxyrrhis and Alexandrium, and five samples encoded a vacuolar H+-pyrophosphatase (L67-1–L67-3, L67-5, LP-1). These two proteins have been hypothesized to function together in dinoflagellates to generate energy in the form of pyrophosphate (Slamovits et al., 2011). The presence of light-driven proton pumps in MALVs suggests an early origin of this system in the dinoflagellate lineage, and that some MALVs may be able to use light for energy (despite the absence of plastids), as this is the suspected function of these proteins in other non-photosynthetic dinoflagellates.
Assemblies from all seven cells include the rRNA operon, generally complete. Mapping Tara Oceans amplicon data to their SSU V9 region demonstrated that two cell types were moderately abundant (>27 000 and >77 000 reads for L67-2 and L67-4, respectively; Figure 1) and widespread in pico- and nano-plankton size fractions (Figures 2a and b). This distribution is beyond Tara’s sampling representation bias (that is, mostly pico- and nano-plankton were sampled; Figure 2c), so if MALVs are parasites, then they predominantly exist as free-living spores rather than host-associated (consistent with the image of L67-4; Figure 1). Four other phylotypes were rare (<5 reads; Figure 1) in the Tara Oceans data. Notably, however, North Pacific data from Tara are unavailable, so these four phylotypes may be endemic to the North Pacific (for example, parasites of North Pacific hosts).
Distribution of MALVs in SSU rRNA amplicon data. (a) Geographical distribution of the two most abundant cells, L67-4 and L67-2, based on Tara Oceans SSU V9 data using a sequence similarity cutoff of 98%. Dot sizes are proportional to the total number of reads in each location for the two phylotypes. (b) Size fraction, depth and temperature distributions of L67-4 and L67-2. The abundances are based on normalized numbers of Tara Oceans V9 reads. (c) Percentages of all Tara samples obtained from different size fractions, depths and water temperatures. N/A – information on size or temperature was not available; polar: <10 °C, temperate: 10–19 °C, tropical: >19 °C.
Originally, MALVs were divided into two major groups: MALV I including Ichthyodinium and Duboscquella (Harada et al., 2007; Skovgaard et al., 2009), and MALV II, including Syndinium, Hematodinium and Amoebophrya (Skovgaard et al., 2005). Both groups were subsequently subdivided further, but the relationships between subgroups remained uncertain (Guillou et al., 2008; Skovgaard et al., 2009; Horiguchi, 2015; see also Supplementary Figure 2). The concatenated SSU/LSU tree (Figure 1) recovers four strongly-supported subgroups: MALV Ia, Ib, II and IV (following the names of Guillou et al., 2008, but distinguishing two phylogenetically distant subgroups of MALV I). Most interestingly MALVs are paraphyletic, with MALV II/IV branching as sister to dinokaryotes (core dinoflagellates). Currently, no synapomorphy for a dinokaryote-MALV-II/IV group is known, but one possibility worth investigation is the presence of DVNPs (dinoflagellate viral nucleoproteins). Within MALVs, DVNPs are found in Hematodinium (MALV IV), Amoebophrya, L67-3, and LP-1 (MALV II; Gornik et al., 2012; Marinov and Lynch, 2015; Supplementary Table 3), but so far, no evidence for their presence in MALV I has been obtained.
The phylogeny raises interesting questions about the distribution of parasitism in early alveolate evolution. In particular, the paraphyletic relationship of seemingly aplastidal parasites at the base of dinoflagellates could suggest a parasitic ancestral state. However, this scenario leaves unexplained many specific shared similarities in the plastids of dinokaryotes, apicomplexans and chrompodellids (Janouškovec et al., 2015), and is inconsistent with the apparent ancestral function of the apical complex as a feeding apparatus (Okamoto and Keeling, 2014a). Alternatively, a mixotrophic ancestry that included both the plastid and the apical complex for feeding is possible; in this case, one must explain an apparently strong predilection to loosing photosynthesis and transitioning to parasitism by modifying the feeding apparatus. A recent analysis of metabolic redundancies within the apicomplexan/dinoflagellate clade also argued for a recent plastid gain, before the divergence of these groups but after they diverged from ciliates (Waller et al., 2015), which is also consistent with this view.
Despite their abundance, distribution, and inferred ecological significance, MALVs remain mysterious. They are assumed to be parasites, and the microscopic and genomic data described here are consistent with this, and in some cases identify new hosts. Their exact evolutionary relationship to dinokaryotes has also been controversial (Guillou et al., 2008; Massana et al., 2008; Horiguchi, 2015), and our data provide strong support for their paraphyly. These insights impact how we reconstruct the evolution of plastids and parasitism in MALVs and dinoflagellates. Finally, surprising aspects of gene content and potential endemism, reveal possible complexities in trophic modes and ecology that serve as a springboard for future investigations.
Data accessibility
Raw reads have been submitted to GenBank under accession numbers SRR5145189 and SRR5177625–SRR5177630, and the assembled contigs can be accessed via the Dryad Digital Repository http://doi.org/10.5061/dryad.hg56n.
References
Chambouvet A, Morin P, Marie D, Guillou L . (2008). Control of toxic marine dinoflagellate blooms by serial parasitic killers. Science 322: 1254–1257.
Gornik SG, Febrimarsa, Cassin AM, MacRae JI, Ramaprasad A, Rchiad Z et al. (2015). Endosymbiosis undone by stepwise elimination of the plastid in a parasitic dinoflagellate. Proc Natl Acad Sci USA 112: 5767–5772.
Gornik SG, Ford KL, Mulhern TD, Bacic A, McFadden GI, Waller RF . (2012). Loss of nucleosomal DNA condensation coincides with appearance of a novel nuclear protein in dinoflagellates. Curr Biol 22: 2303–2312.
Guillou L, Alves-de-Souza C, Siano R, González H . (2010). The ecological significance of small, eukaryotic parasites in marine ecosystems. Microbiol Today 37: 92–95.
Guillou L, Viprey M, Chambouvet A, Welsh RM, Kirkham AR, Massana R et al. (2008). Widespread occurrence and genetic diversity of marine parasitoids belonging to Syndiniales (Alveolata). Environ Microbiol 10: 3349–3365.
Harada A, Ohtsuka S, Horiguchi T . (2007). Species of the parasitic genus Duboscquella are members of the enigmatic Marine Alveolate Group I. Protist 158: 337–347.
Horiguchi T . (2015) Diversity and phylogeny of marine parasitic dinoflagellates. In: Ohtsuka S, Suzaki T, Horiguchi T, Suzuki N, Not F (eds). Marine Protists: Diversity and Dynamics. Springer Japan: Tokyo, pp 397–419.
Janouškovec J, Tikhonenkov DV, Burki F, Howe AT, Kolísko M, Mylnikov AP et al. (2015). Factors mediating plastid dependency and the origins of parasitism in apicomplexans and their close relatives. Proc Natl Acad Sci USA 112: 10200–10207.
López-García P, Rodriguez-Valera F, Pedrós-Alió C, Moreira D . (2001). Unexpected diversity of small eukaryotes in deep-sea Antarctic plankton. Nature 409: 603–607.
Marinov GK, Lynch M . (2015). Diversity and divergence of dinoflagellate histone proteins. G3 (Bethesda) 6: 397–422.
Massana R, Karniol B, Pommier T, Bodaker I, Beja O . (2008). Metagenomic retrieval of a ribosomal DNA repeat array from an uncultured marine alveolate. Environ Microbiol 10: 1335–1343.
Moon-van der Staay SY, de Wachter R, Vaulot D . (2001). Oceanic 18 S rDNA sequences from picoplankton reveal unsuspected eukaryotic diversity. Nature 409: 607–610.
Okamoto N, Keeling PJ . (2014a). The 3D structure of the apical complex and association with the flagellar apparatus revealed by serial TEM tomography in Psammosa pacifica, a distant relative of the apicomplexa. PLoS One 9: e84653.
Okamoto N, Keeling PJ . (2014b). A comparative overview of the flagellar apparatus of dinoflagellates, perkinsids, and colpodellids. Microorganisms 2: 73–91.
Skovgaard A, Massana R, Balagué V, Saiz E . (2005). Phylogenetic position of the copepod-infesting parasite Syndinium turbo (Dinoflagellata, Syndinea). Protist 156: 413–423.
Skovgaard A, Meneses I, Angelico MM . (2009). Identifying the lethal fish egg parasite Ichthyodinium chabelardi as a member of Marine Alveolate Group I. Environ Microbiol 11: 2030–2041.
Slamovits CH, Okamoto N, Burri L, James ER, Keeling PJ . (2011). A bacterial proteorhodopsin proton pump in marine eukaryotes. Nat Commun 2: 183.
de Vargas C, Audic S, Henry N, Decelle J, Mahe F, Logares R et al. (2015). Ocean plankton. Eukaryotic plankton diversity in the sunlit ocean. Science 348: 1261605.
Waller RF, Gornik SG, Koreny L, Pain A . (2015). Metabolic pathway redundancy within the apicomplexan-dinoflagellate radiation argues against an ancient chromalveolate plastid. Commun Integr Biol 9: e1116653.
Wisecaver JH, Hackett JD . (2011). Dinoflagellate genome evolution. Annu Rev Microbiol 65: 369–387.
Acknowledgements
We thank the captain and crew of the R/V Western Flyer, M Blum, F Chavez, V Jimenez, JT Pennington, JM Smith, S Sudek, J Swalwell, C Wahl and S Wilken, for logistical assistance before and during the cruises. This work was supported by a grant from the Gordon and Betty Moore Foundation (GBMF3307) to PJK, TAR, AZW and AES. Sequencing for L67-2 was performed through a JGI Technology Development Program grant to TAR, AZW, PJK and AES. Ship time and sorting methods were supported by a grant from the David and Lucile Packard Foundation through MBARI and GBMF3788 to AZW. JdC was supported by a Marie Curie International Outgoing Fellowship grant (FP7-PEOPLE-2012-IOF – 331450 CAARL), and AK, NO, MK and JdC were supported by a grant from the Tula Foundation to the Centre for Microbial Biodiversity and Evolution at UBC.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Strassert, J., Karnkowska, A., Hehenberger, E. et al. Single cell genomics of uncultured marine alveolates shows paraphyly of basal dinoflagellates. ISME J 12, 304–308 (2018). https://doi.org/10.1038/ismej.2017.167
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.167
This article is cited by
-
Water masses shape pico-nano eukaryotic communities of the Weddell Sea
Communications Biology (2023)
-
Multiple parallel origins of parasitic Marine Alveolates
Nature Communications (2023)
-
Beyond the limits of the unassigned protist microbiome: inferring large-scale spatio-temporal patterns of Syndiniales marine parasites
ISME Communications (2023)
-
Microscopic marine invertebrates are reservoirs for cryptic and diverse protists and fungi
Microbiome (2022)
-
Long rDNA amplicon sequencing of insect-infecting nephridiophagids reveals their affiliation to the Chytridiomycota and a potential to switch between hosts
Scientific Reports (2021)